Abstract
Missense mutations in p53 frequently occur at `hotspot' amino acids which are highly conserved and represent regions of structural or functional importance. Using the p53 mutation database and the p53 DNA sequences for 11 species, we more precisely defined the relationships among conservation, mutation frequency and protein structure. We aligned the p53 sequences codon-by-codon and determined the degree of substitution among them. As a whole, p53 is evolving at an average rate for a mammalian protein-coding gene. As expected, the DNA binding domain is evolving more slowly than the carboxy and amino termini. A detailed map of evolutionary conservation shows that within the DNA binding domain there are repeating peaks and valleys of higher and lower evolutionary constraint. Mutation hotspots were identified by comparing the observed distribution of mutations to the pattern expected from a random multinomial distribution. Seventy-three hotspots were identified; these 19% of codons account for 88% of all reported p53 mutations. Both high evolutionary constraint and mutation hotspots are noted at amino acids close to the protein-DNA interface and at others more distant from DNA, often buried within the core of the folded protein but sometimes on its surface. The results indicate that targeting highly conserved regions for mutational and functional analysis may be efficient strategies for the study of cancer-related genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Carson M. . 1991 J. Appl. Cryst. 24: 958–961.
Cho Y, Gorina S, Jeffrey P and Pavletich NP. . 1994 Science 265: 346–355.
Clore GM, Omichinski JG, Sakaguchi K, Zambrano N, Sakamoto H, Appella E and Gronenborn AN. . 1994 Science 265: 386–391.
Greenblatt MS, Hollstein M, Bennett WP and Harris CC. . 1994 Cancer Res. 54: 4855–4878.
Hollstein M, Rice K, Greenblatt MS, Soussi T, Fuchs R, Sorlie T, Hovig E, Smith-Sorensen B, Montesano R and Harris CC. . 1994 Nucleic Acids Res. 22: 3551–3555.
Hollstein M, Shomer B, Greenblatt M, Soussi T, Hovig E, Montesano R and Harris CC. . 1996 Nucleic Acids Res. 24: 141–146.
Hsu I-C, Metcalf RA, Sun T, Welsh J, Wang NJ and Harris CC. . 1991 Nature 350: 427–428.
Ina Y. . 1995 J. Mol. Evol. 40: 190–226.
Jeffrey PD, Gorina S and Pavletich NP. . 1995 Science 267: 1498–1502.
Jost Ca, Marin MC and Kaelin WG. . 1997 Nature 389: 191–194.
Kaghad M, Bonnet H, Yang A, Creacer L, Biscan J-C, Valent A, Minty A, Chalon P, Lelias J-M, Dumont X, Ferrara P, McKeon F and Caput D. . 1997 Cell 90: 809–819.
Kimura M. . 1980 J. Mol. Evol. 16: 111–120.
Kyte J. . 1995 Mechanism in protein chemistry. Garland Publishers: New York.
Li W-H and Grauer D. . 1991 Fundamentals of Molecular Evolution. Sunderland MA (ed.).. Sinauer Associates, Inc. pp. 69–70.
Li W-H. . 1997 Molecular Evolution. Sunderland MA (ed.).. Sinauer Associates, Inc.
Makalowski W and Boguski MS. . 1998 Proc. Natl. Acad. Sci. USA in press.
Miller RG. . 1981 Simultaneous Statistical Inference. Springer-Verlag: New York. pp. 6–8.
Rainwater R, Parks D, Anderson ME, Tegtmeyer P and Mann K. . 1995 Molec. Cell. Biol. 15: 3892–3903.
Rodin SN, Holmquist GP and Rodin AS. . 1998 Int. J. Molec. Med. 1: 191–199.
Schuler GD, Altschul SF and Lipman DJ. . 1991 Proteins 9: 180–190.
Soussi T, Caron de Fromental C and May P. . 1990 Oncogene 5: 945–952.
Sved J and Bird A. . 1990 Proc. Natl. Acad. Sci. USA 87: 4692–4696.
Ticher A and Graur D. . 1989 J. Mol. Evol. 28: 286–298.
Tornaletti S and Pfeifer GP. . 1994 Science 263: 1436–1439.
Walker DR and Koonin EV. . 1997 ISBM 5: 333–339.
Walker KK and Levine AJ. . 1996 Proc. Natl. Acad. Sci. USA 93: 15335–15340.
Wang XW, Vermeulen W, Coursen JD, Gibson M, Lupold SE, Forrester K, Xu G, Elmore L, Yeh H, Hoeijmakers JHJ and Harris CC. . 1996 Genes and Dev. 10: 1219–1232.
Acknowledgements
We appreciate the assistance of Gary J Nelson in preparation of the figures.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Walker, D., Bond, J., Tarone, R. et al. Evolutionary conservation and somatic mutation hotspot maps of p53: correlation with p53 protein structural and functional features. Oncogene 18, 211–218 (1999). https://doi.org/10.1038/sj.onc.1202298
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1202298
Keywords
This article is cited by
-
Novel MSX1 variants identified in families with nonsyndromic oligodontia
International Journal of Oral Science (2021)
-
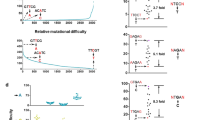
Defining relative mutational difficulty to understand cancer formation
Cell Discovery (2020)
-
Enhanced breast cancer progression by mutant p53 is inhibited by the circular RNA circ-Ccnb1
Cell Death & Differentiation (2018)
-
A novel mouse model of rhabdomyosarcoma underscores the dichotomy of MDM2-ALT1 function in vivo
Oncogene (2018)
-
Adaptive patterns in the p53 protein sequence of the hypoxia- and cancer-tolerant blind mole rat Spalax
BMC Evolutionary Biology (2016)