Abstract
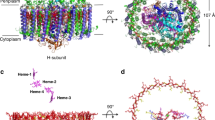
We report here the first three-dimensional structure of the D1 C-terminal processing protease (D1P), which is encoded by the ctpA gene. This enzyme removes the C-terminal extension of the D1 polypeptide of photosystem II of oxygenic photosynthesis. Proteolytic processing is necessary to allow the light driven assembly of the tetranuclear manganese cluster, which is responsible for photosynthetic water oxidation. The X-ray structure of the Scenedesmus obliquus enzyme has been determined at 1.8 Å resolution using the multiwavelength anomalous dispersion method. The enzyme is monomeric and is composed of three folding domains. The middle domain is topologically homologous to known PDZ motifs and is proposed to be the site at which the substrate C-terminus binds. The remainder of the substrate likely extends across the face of the enzyme, interacting at its scissile bond with the enzyme active site Ser 372 / Lys 397 catalytic dyad, which lies at the center of the protein at the interface of the three domains.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Takahashi, M., Shiraishi, T. & Asada, K. FEBS Lett. 240, 6–8 (1988).
Takahashi, Y., Nakane, H., Kojima, H. & Satoh, K. Plant Cell Physiol. 31, 273–280 (1990).
Nixon, P.J., Trost, J.T. & Diner, B.A. Biochemistry 31, 10859–10871 (1992).
Metz, J.G., Pakrasi, H., Arntzen, C.J. & Siebert, M. Prog. Photosynth. Res. Proc. Int. Congr. Photosynth. 7th 4, 679–682 (1987).
Taylor, M.A., Packer, J.C.L. & Bowyer, J.R. FEBS Lett. 237, 229–233 (1988).
Trost, J.T., Chisholm, D.A., Jordan, D.B. & Diner, B.A. J. Biol. Chem. 272, 20348–20356 (1997).
Inagaki, N., Yamamoto, Y., Mori, H. & Satoh, K. Plant Mol. Biol. 30, 39–50 (1996).
Shestakov, S.V. et al. J. Biol. Chem. 269, 19354–19359 (1994).
Hara, H., Yamamoto, Y., Higashitani, A., Suzuki, H. & Nishimura, Y. J. Bacteriol. 173, 4799–4813 (1991).
Silber, K.R., Keiler, K.C. & Sauer, R.T. Proc. Natl. Acad. Sci. USA 89, 295–299 (1992).
Tomb, J.F. et al. Nature 388, 539–547 (1997).
Keiler, K.C. & Sauer, R.T. J. Biol. Chem. 270, 28864–28868 (1995).
Paetzel, M. & Dalbey, R.E. Trends Biochem. Sci. 22, 28–31 (1997).
Tschantz, W.R., Sung, M., Delgado-Partin, V.M. & Dalbey, R.E. J. Biol. Chem. 268, 27349–27354 (1993).
Paetzel, M., Dalbey, R.E. & Strynadka, N.C.J. Nature 396, 186–190 (1998).
Peat, T.S. et al. Nature 380, 727–730 (1996).
Slilaty, S.N. & Little, J.W. Proc. Natl. Acad. Sci. USA 84, 3987–3991 (1987).
Strynadka, N.C.J. et al. Nature 359, 700–705 (1992).
Rawlings, N.D. & Barrett, A.J. In Proteolysis in cell functions (eds Hopsu-Havu,V.K., Järvinen, M., Kirschke, H.), 13–21 (IOS Press, Amsterdam; 1997).
Kim, J.L. et al. Cell 87, 343–355 (1996).
Slilaty, S.N. & Vu, H.K. Protein Eng. 4, 919–922 (1991).
Ponting, C.P. Protein Sci. 6, 464–468 (1997).
Beebe, K.D. et al. Biochemistry 39, 3149–3155 (2000).
Cabral, J.H.M. et al. Nature 382, 649–652. (1996).
Doyle, D.A. et al. Cell 85, 1067–1076 (1996).
Hillier, B.J., Christopherson, K.S., Prehoda, K.E., Bredt, D.S. & Lim, W.A. Science 284, 812–815.
Nixon, J.J., Rögner, M. & Diner, B.A. Plant Cell 3, 383–395 (1991).
Morden, C.W. & Golden, S.S. Nature 337, 382–385 (1989).
Otwinowski, Z & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Hendrickson, W.A., Horton, J.R. & LeMaster, D.M. EMBO J. 9, 1665–1672 (1990).
Sheldrick, G.M. In Direct methods for solving macromolecular structures (ed. Fortier, S.), 131–141 (Kluwer Academic Publishers, Dordrecht; 1998).
Furey, W. & Swaminathan, S. Methods Enzymol. 277, 590–619 (1997).
Cowtan, K. Joint CCP4 (1994) and ESF-EACBM Newsletter on Protein Crystallography 31, 34–38 (1994).
Collaborative Computational Project, Number 4. CCP4 Suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T., Kuriyan, J. & Karplus, M. Science 235, 458–460 (1987).
Tronrud, D.E. Acta Crystallogr. A 48, 912–916 (1991).
Navaza, J. Acta Crystallogr. A 50, 157–163 (1994).
Acknowledgements
We thank J. Wang for her contributions to the early stage of this project and M. Nelson for helpful discussions. This paper is Contribution No. 8074 of the Central Research and Development Department of E. I. du Pont de Nemours & Co.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Liao, DI., Qian, J., Chisholm, D. et al. Crystal structures of the photosystem II D1 C-terminal processing protease. Nat Struct Mol Biol 7, 749–753 (2000). https://doi.org/10.1038/78973
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/78973
This article is cited by
-
Antibiotic Interactions with Key Algal Proteins Responsible for Photosynthesis: Molecular Docking-Based Correlates
Proceedings of the National Academy of Sciences, India Section B: Biological Sciences (2021)
-
Structural basis of adaptor-mediated protein degradation by the tail-specific PDZ-protease Prc
Nature Communications (2017)
-
Structural basis of lantibiotic recognition by the nisin resistance protein from Streptococcus agalactiae
Scientific Reports (2016)
-
A novel carboxyl-terminal protease derived from Paenibacillus lautusCHN26 exhibiting high activities at multiple sites of substrates
BMC Biotechnology (2013)
-
A genome-wide study of two-component signal transduction systems in eight newly sequenced mutans streptococci strains
BMC Genomics (2012)