Abstract
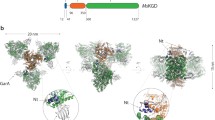
The family of giant multienzyme complexes metabolizing pyruvate, 2-oxoglutarate, branched-chain 2-oxo acids or acetoin contains several of the largest and most sophisticated protein assemblies known, with molecular masses between 4 and 10 million Da. The principal enzyme components, E1, E2 and E3, are present in numerous copies and utilize multiple cofactors to catalyze a directed sequence of reactions via substrate channeling. The crystal structure of a heterotetrameric (α2β2) E1, 2-oxoisovalerate dehydrogenase from Pseudomonas putida, reveals a tightly packed arrangement of the four subunits with the β2-dimer held between the jaws of a 'vise' formed by the α2-dimer. A long hydrophobic channel, suitable to accommodate the E2 lipoyl-lysine arm, leads to the active site, which contains the cofactor thiamin diphosphate (ThDP) and an inhibitor-derived covalent modification of a histidine side chain. The E1 structure, together with previous structural information on E2 and E3, completes the picture of the shared architectural features of these enormous macromolecular assemblies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Reed, L.J. Multienzyme complexes. Accounts Chem. Res. 7, 40–46 (1974).
Perham, R.N. Domains, motifs, and linkers in 2-oxo acid dehydrogenase multienzyme complexes: a paradigm in the design of a multifunctional protein. Biochemistry 30, 8501–8512 ( 1991).
Izard, T. et al. Principles of quasi-equivalence and Euclidean geometry govern the assembly of cubic and dodecahedral cores of pyruvate dehydrogenase complexes. Proc. Natl. Acad. Sci. USA 96, 1240– 1245 (1999).
Patel, M.S. & Roche, T.M. Molecular biology and biochemistry of pyruvate dehydrogenase complexes. FASEB J. 4, 3224–3233 (1990).
Wynn, R.M., Davie, J.R., Meng, M. & Chuang, D.T. In Alpha-keto acid dehydrogenase complexes (eds Patel, M.S., Roche, T.E. & Harris, R.A.) 101–117 (Birkhäuser Verlag, Basel; 1996).
Chuang, D.T. & Shih, V.E. In The metabolic and molecular bases of inherited disease (eds Scriver, C.R., Beudet, A.L., Sly, W.S. & Valle, D.) 1239–1227 (McGraw-Hill, New York; 1995).
Kerr, D.S., Wexler, I.D., Tripatara, A. & Patel, M.S. In Alpha-keto acid dehydrogenase complexes. (eds Patel, M.S., Roche, T.E. & Harris, R.A.) 249–267 (Birkhäuser Verlag, Basel; 1996).
Massey, L.K., Sokatch, J.R., & Conrad, R.S. Branched-chain amino acid catabolism in bacteria. Bacteriol. Rev. 40, 42– 54 (1976).
Hawkins, C.F., Borges, A. & Perham, R.N. A common structural motif in thiamin pyrophosphate-binding enzymes. FEBS Lett. 255, 77– 82 (1989).
Perham, R.N. & Packman, L.C. 2-Oxo acid dehydrogenase multienzyme complexes: domains, dynamics and design. Ann. N.Y. Acad. Sci. 573, 1–20 (1989).
Burns, G., Brown, T., Hatter, K., Idriss, J.M. & Sokatch, J.R. Similarity of the E1 subunits of branched-chain-oxoacid dehydrogenase from Pseudomonas putida to the corresponding subunits of mammalian branched-chain-oxoacid and pyruvate dehydrogenases. Eur. J. Biochem. 176, 311–317 (1988).
Nikkola, M., Lindqvist, Y. & Schneider, G. Refined structure of transketolase from Saccharomyces cerevisiae at 2.0 Å resolution. J. Mol. Biol. 238, 387–404 (1994).
Muller, Y.A. et al. A thiamin diphosphate binding fold revealed by comparison of the crystal structures of transketolase, pyruvate oxidase and pyruvate decarboxylase. Structure 1, 95– 103 (1993).
Hasson, M.S. et al. The crystal structure of benzoylformate decarboxylase at 1.6 Å resolution: diversity of catalytic residues in thiamin diphosphate-dependent enzymes. Biochemistry 37, 9918– 9930 (1998).
Chabrière, E., Charon, M.-H., Volbeda, A., Pieulle, L., Hatchikian, E.C. & Fontecilla-Camps, J.-C. Crystal structures of the key anaerobic enzyme pyruvate:ferredoxin oxidoreductase, free and in complex with pyruvate. Nature Struct. Biol. 6, 182– 190 (1999).
Sundström, M., Lindqvist, Y., Schneider, G., Hellman, U. & Ronne, H. Yeast TKL1 gene encodes a transketolase that is required for efficient glycolysis and biosynthesis of aromatic amino acids. J. Biol. Chem. 268, 24346– 24352 (1993).
Schneider, G. & Lindqvist, Y. Crystallography and mutagenesis of transketolase: mechanistic implications for enzymatic thiamin catalysis. Biochim. Biophys. Acta 1385, 387– 398 (1998).
Guo, F., Zhang, D., Kahyaoglu, A., Farid, R.S. & Jordan, F. Is a hydrophobic amino acid required to maintain the reactive V conformation of thiamin at the active center of thiamin diphosphate-requiring enzymes? Experimental and computational studies of isoleucine 415 of yeast pyruvate decarboxylase. Biochemistry 37, 13379–13391 (1998).
Schellenberger, A. Sixty years of thiamin diphosphate biochemistry. Biochim. Biophys. Acta 1385, 177–186 ( 1998).
Hübner, G. et al. Activation of thiamin diphosphate in enzymes. Biochim. Biophys. Acta 1385, 221–228 (1998).
Harris, R.A., Paxton, R. & DePaoli-Roach, A.A. Inhibition of branched chain α-ketoacid dehydrogenase kinase activity by α-chloroisocaproate. J. Biol. Chem. 257, 13915–13918 (1982).
Bürgi, H.B., Dunitz, J.D., Lehn, J.M. & Wipff, G. Stereochemistry of reaction paths at carbonyl centres. Tetrahedron 30, 1563–1572 (1974).
Hawes, J.W. et al. Roles of amino acid residues surrounding phosphorylation site 1 of branched-chain α-ketoacid dehydrogenase (BCKDH) in catalysis and phophorylation site recognition by BCKDH kinase. J. Biol. Chem. 270, 31071–31076 ( 1995).
Hester, K., Luo, J., Burns, G., Braswell, E.H. & Sokatch, J.R. Purification of active E1α2β 2 of Pseudomonas putida branched-chain-oxoacid dehydrogenase. Eur. J. Biochem. 233, 828– 836 (1995).
Korotchkina, L.G., Ali, M.S. & Patel, M.S. In Alpha-keto acid dehydrogenase complexes (eds Patel, M.S., Roche, T.E. & Harris, R.A.) 17–32 (Birkhäuser Verlag, Basel; 1996).
Berg, A. & de Kok, A. 2-Oxo acid dehydrogenase multienzyme complexes. The central role of the lipoyl domain. Biol. Chem. 378, 617–634 (1997).
Pan, K. & Jordan, F. D,L-S-Methyllipoic acid methyl ester, a kinetically viable model for S-protonated lipoic acid as the oxidizing agent in reductive acyl transfers catalyzed by the 2-oxoacid dehydrogenase multienzyme complexes. Biochemistry 37, 1357–1364 (1998).
Yang, Y.S. & Frey, P.A. Dihydrolipoyl transacetylase of Escherichia coli. Formation of 8-S-acetyldihydrolipoamide. Biochemistry 25, 8173–8178 (1986).
Perham, R.N. In Alpha-keto acid dehydrogenase complexes (eds Patel, M.S., Roche, T.E. & Harris, R.A.) 1–15 (Birkhäuser Verlag, Basel; 1996).
Wynn, R.M. et al. Cloning and expression in Escherichia coli of mature E1β subunit of bovine mitochondrial branched-chain α-keto acid dehydrogenase complex. J. Biol. Chem. 267, 1881–1887 (1992).
Mande, S.S., Sarfaty, S., Allen, M.D., Perham, R.N. & Hol, W.G.J. Protein–protein interactions in the pyruvate dehydrogenase multienzyme complex: dihydrolipoamide dehydrogenase complexed with the binding domain of dihydrolipoamide acetyltransferase. Structure 4, 277–286 (1996).
Mattevi, A. et al. Atomic structure of the cubic core of the pyruvate dehydrogenase multienzyme complex. Science 255, 1544– 1550 (1992).
Mattevi, A., Obmolova, G., Sokatch, J.R., Betzel, C. & Hol, W.G.J. The refined crystal structure of Pseudomonas putida lipoamide dehydrogenase complexed with NAD+ at 2.45 Å resolution. Proteins 13, 336–351 (1992).
Kalia, Y.N. et al. The high-resolution structure of the peripheral subunit-binding domain of dihydrolipoamide acetyltransferase from the pyruvate dehydrogenase multienzyme complex of Bacillus stearothermophilus. J. Mol. Biol. 230, 323–341 ( 1993).
Dardel, F., Davis, A.L., Laue, E.D. & Perham, R.N. Three-dimensional structure of the lipoyl domain from Bacillus stearothermophilus pyruvate dehydrogenase multienzyme complex. J. Mol. Biol. 229 , 1037–1048 (1993).
Mattevi, A., Obmolova, G., Kalk, K.H., Teplyakov, A. & Hol, W.G.J. Crystallographic analysis of substrate binding and catalysis in dihydrolipoyl transacetylase. Biochemistry 32, 3887–3901 (1993).
Bagdasarian, M.M. et al. Specific-purpose plasmid cloning vectors. II. Broad host range, high copy number, RSF1010-derived vectors, and a host-vector system for gene cloning in Pseudomonas. Gene 16, 237–247 (1981).
Sykes, P.J., Menard, J., McCully, V. & Sokatch, J.R. Conjugative mapping of pyruvate, 2-ketoglutarate, and branched-chain keto acid dehydrogenase genes in Pseudomonas putida mutants. J. Bacteriol. 162, 203–208 (1985).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 277, 307–326 ( 1997).
Collaborative Computational Project Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 ( 1994).
Navazza, J. AMoRe, an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 ( 1994).
de La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement in the MIR and MAD methods. Methods Enzymol. 276, 472–494 (1997).
Abrahams, J.P. & Leslie, A.G.W. Methods used in the structure determination of bovine mitochondral F1 ATPase. Acta Crystallogr. D 52, 30– 42 (1996).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T., Krukowski, A. & Erickson, J.W. Slow-cooling protocols for crystallographic refinement by simulated annealing. Acta Crystallogr. A 46, 585–593 (1990).
Brünger, A.T., et al. Crystallography and NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946 –950 (1991).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. 15, 132–134 (1997).
Nicholls, A., Shar, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Gen. 11, 282–296 (1991).
Merritt, E.A. & Murphy, M.E.P. Raster3D Version 2.0—a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 ( 1994).
Acknowledgements
We thank the members of the Biomolecular Structure Center for assistance and support, in particular S.S. Antonysamy, F. Athappilly, E. Merritt, E. Pohl, M. Redinbo and S. Sarfaty. Access to synchrotron sources SSRL (7.1), CHESS (F1) and ESRF (BM14) is deeply appreciated, and we thank the staff for their assistance. Postdoctoral grant from The Swedish Foundation for International Cooperation in Research and Higher Education (STINT) to A.Æ. is gratefully acknowledged. W.G.J.H. acknowledges a major equipment grant from the Murdock Charitable Trust to the Biomolecular Structure Center. This research was supported by grants from NIH and Presbyterian Health Foundation to J.R.S.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ævarsson, A., Seger, K., Turley, S. et al. Crystal structure of 2-oxoisovalerate and dehydrogenase and the architecture of 2-oxo acid dehydrogenase multienzyme complexes. Nat Struct Mol Biol 6, 785–792 (1999). https://doi.org/10.1038/11563
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/11563
This article is cited by
-
Host-adaptive traits in the plant-colonizing Pseudomonas donghuensis P482 revealed by transcriptomic responses to exudates of tomato and maize
Scientific Reports (2023)
-
Branched-chain amino acid catabolism in muscle affects systemic BCAA levels but not insulin resistance
Nature Metabolism (2023)
-
Detailed kinetics and regulation of mammalian 2-oxoglutarate dehydrogenase
BMC Biochemistry (2011)
-
Structural basis of enzyme encapsulation into a bacterial nanocompartment
Nature Structural & Molecular Biology (2008)
-
Evolutionary Analysis of the TPP-Dependent Enzyme Family
Journal of Molecular Evolution (2008)