Abstract
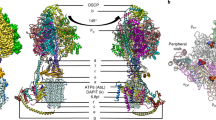
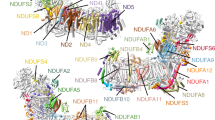
The nicotinamide nucleotide transhydrogenases (TH) of mitochondria and bacteria are membrane-intercalated proton pumps that transduce substrate binding energy and protonmotive force via protein conformational changes. In mitochondria, TH utilizes protonmotive force to promote direct hydride ion transfer from NADH to NADP, which are bound at the distinct extramembranous domains I and III, respectively. Domain II is the membrane-intercalated domain and contains the enzyme's proton channel. This paper describes the crystal structure of the NADP(H) binding domain III of bovine TH at 1.2 Å resolution. The structure reveals that NADP is bound in a manner inverted from that previously observed for nucleotide binding folds. The non-classical binding mode exposes the NADP(H) nicotinamide ring for direct contact with NAD(H) in domain I, in accord with biochemical data. The surface of domain III surrounding the exposed nicotinamide is comprised of conserved residues presumed to form the interface with domain I during hydride ion transfer. Further, an adjacent region contains a number of acidic residues, forming a surface with negative electrostatic potential which may interact with extramembranous loops of domain II. Together, the distinctive surface features allow mechanistic considerations regarding the NADP(H)-promoted conformation changes that are involved in the interactions of domain III with domains I and II for hydride ion transfer and proton translocation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hatefi, Y. & Yamaguchi, M. FASEB J. 10, 444–452 (1996).
Hatefi, Y. & Yamaguchi, M. in Molecular mechanisms in bioenergetics (ed. Ernster, L.) 265–281 (Elsevier Science Publisher, Amsterdam, 1992).
Jackson, J.B., et al. Biochim. Biophys. Acta 1365, 79– 86 (1998).
Rydström, J. et al. Biochim. Biophys. Acta 1365, 10– 16 (1998).
Clarke, D.M. & Bragg, P.D. Eur. J. Biochem. 149 , 517–523 (1985).
Yamaguchi, M. & Hatefi, Y. J. Biol. Chem. 268, 17871–17877 (1993).
Yamaguchi, M. & Hatefi, Y. J. Biol. Chem. 270, 28165–28168 (1995).
Yamaguchi, M. & Hatefi, Y. Biochim. Biophys. Acta 1318, 225–234 (1997).
Bellamacina, C.R. FASEB J. 10, 1257–1269 ( 1996).
Wakabayashi, S. & Hatefi, Y. Biochem. Int. 15, 915–924 (1987).
Yamaguchi, M. & Hatefi, Y. Biochemistry 28, 6050–6056 (1989).
Ahmad, S., Glavas, N.A. & Bragg, P.D. Eur. J. Biochem. 207, 733– 739 (1992).
Olausson, T., et al. Biochemistry 32, 13237– 13244 (1993).
Mueller, J., Hu, X., Bunthof, C. Olausson, T. & Rydström, J. Biochim. Biophys. Acta 1273, 191–194 (1996).
Bragg, P.D., Glavas, N.A. & Hou, C. Arch. Biochem. Biophys. 338, 57 –66 (1997).
Hu, X., Zhang, J., Fjellström, O., Bizouarn, T. & Rydström, J. Biochemistry 38 , 1652–1658 (1999).
Fjellström, O., et al. J. Biol. Chem. 274, 6350– 6359 (1999).
Johansson, C., Bergkvist, A., Fjellström, O., Rydström, J. & Karlsson, B.G. FEBS Lett. 458, 180–184 (1999).
Quirk, P.G., Jeeves, M., Cotton, N.P.J., Smith, J.K. & Jackson, B.J. FEBS Lett. 446, 127–132 (1999).
Leslie, A.G.W. Acta Crystallogr. D50, 760–763 (1994).
Otwinowski, Z. Acta Crystallogr. D50, 760–763 (1994).
Cowtan, K. NATO ASI Ser., Ser. C, 507, 329–337 (1998).
McRee, D.E. J. Struct. Biol. 125, 156–165 (1999).
Brünger, A.T., et al. Acta Cryst. D54, 905– 921 (1998).
Sheldrick, G.M. & Schneider, T.R. Methods Enzymol. 277, 319–343 ( 1997).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. J Appl. Crystallogr. 26, 283–291 (1993).
McRee, D.E. Molecular Images Software, San Diego, CA (1999).
Altschul, S.F., et al. Nucleic Acids Res. 25, 3389– 3402 (1997).
Bruns, C. M. http://www.scripps.edu/~bruns/sequoia.html (1999).
Kraulis, P.J. J. Appl. Cryst. 24, 946–950 (1991).
Nicholls, A., Sharp, K.A. & Honig, B. Proteins 11, 281– 296 (1991).
Acknowledgements
The authors thank N. Kresge for preparing several of the figures and the staff of the Stanford Synchrotron Radiation Laboratory for their generous assistance. The authors are indebted to U. Genick and E. Getzoff for their generous assistance in the use of their single crystal microspectrophotometer. This work was supported by a United States Public Health Service Grant to Y. Hatefi.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Prasad, G., Sridhar, V., Yamaguchi, M. et al. Crystal structure of transhydrogenase domain III at 1.2 Å resolution . Nat Struct Mol Biol 6, 1126–1131 (1999). https://doi.org/10.1038/70067
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/70067
This article is cited by
-
Energy transfer between the nicotinamide nucleotide transhydrogenase and ATP synthase of Escherichia coli
Scientific Reports (2021)
-
Structure and mechanism of mitochondrial proton-translocating transhydrogenase
Nature (2019)
-
Proton-translocating transhydrogenase: an update of unsolved and controversial issues
Journal of Bioenergetics and Biomembranes (2008)