Abstract
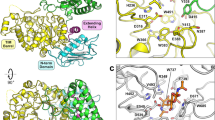
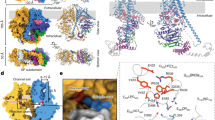
Chitin, the second most abundant polysaccharide on earth, is degraded by chitinases and chitobiases. The structure of Serratia marcescens chitobiase has been refined at 1.9 Å resolution. The mature protein is folded into four domains and its active site is situated at the C-terminal end of the central (βα)8-barrel. Based on the structure of the complex with the substrate disaccharide chitobiose, we propose an acid-base reaction mechanism, in which only one protein carboxylate acts as catalytic acid, while the nucleophile is the polar acetamido group of the sugar in a substrate-assisted reaction. The structural data lead to the hypothesis that the reaction proceeds with retention of anomeric configuration. The structure allows us to model the catalytic domain of the homologous hexosaminidases to give a structural rationale to pathogenic mutations that underlie Tay–Sachs and Sandhoff disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Henrissat, B. & Bairoch, A. New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem. J. 293, 781–788 (1993).
Davies, G.J. & Henrissat, B. Structures and mechanisms of glycosyl hydrolases. Structure 3, 853–859 (1995).
Chet, I., Barak, Z. & Oppenheim, A.B. Genetic engineering of microorganisms for improved biocontrol activity. in Biotechnology in Plant Disease Control (ed. I. Chet) 211–235 (Wiley Liss, Inc. 1993).
ZoBell, C.E. & Rittenberg, S.C. The occurence and characteristics of chitinoclastic bacteria in the sea. J. Bacteriol. 35, 275–287 (1937).
Perrakis, A. et al. Crystal structure of a bacterial chitinase at 2.3 Å resolution. Structure 2, 1169–1180 (1994).
Roey, P.V., Rao, V., Plummer, Jr., T.H. & Tarentino, A.L. Crystal Structure of Endo-β-N-acetylglucosaminidase F1, an α/β-Barrel Enzyme Adapted for a Complex Substrate. Biochem. 33, 13989–13996 (1994).
Rao, V., Guan, C. & Roey, P.V. Crystal structure of endo-β-N-acetylglucosaminidase H at 1.9 Å resolution: active-site geometry and substrate recognition. Structure 3, 449–457 (1995).
Terwisscha van Scheltinga, A.C., Kalk, K.H., Beintema, J.J. & Dijkstra, B.W. Crystal structure of hevamine, a plant defence protein with chitinase and lysozyme activity. Structure 2, 1181–1189 (1994).
Hart, P.J., Pfluger, H.D., Monzingo, A.F., Hollis, T. & Robertus, J.D. The Refined Crystal Structure of the Endochitinase from Hordeum vulgare L. Seeds at 1.8 Å Resolution. J. Mol. Biol. 248, 402–413 (1995).
Oppenheim, A.B. & Chet, I. Cloned chitinases in fungal plant-pathogen control strategies. Trends. Biotech. 10, 392–394 (1992).
Roberts, R.L. & Cabib, E. Serratia marcescens Chitinase: One-Step Purification and Use for the Determination of Chitin. Analyt. Biochem. 127, 402–412 (1982).
Sandhoff, K., Conzelmann, E., Neufeld, E.F., Kaback, M.M. & Suzuki, K. The G M2 ganglosidoses. in The Metabolic Base of Inherited Disease 6th ed. (ed. Scriver, C.R., Beaudet, A.L., Sly, W.S. & Valle, D.) 1807–1842 (McGraw-Hill, New York, 1988).
Mahuran, D.J. The biochemistry of HEXA and HEXB gene mutations causing GM2 gangliosidosis. Biochim. Biophys. Acta. 1096, 87–94 (1991).
Tews, I., Vincentelli, R. & Vorgias, C.E. N-acetylglucosaminidase (chitobiase) from Serratia marcescens: Gene sequence, and protein production and purification in Escherichia coli. Gene 170, 63–67 (1996).
Xu, G.-Y. et al. Solution structure of a cellulose-binding domain from Cellulomonas fimi by nuclear magnetic resonance spectroscopy. Biochem. 34, 6993–7009 (1995).
Baumann, U. Crystal Structure of the 50 kDa Metallo protease from Serratia marcescens. J. Mol. Biol. 242, 244–251 (1994).
Banner, D.W. et al. Structure of chicken muscle triose phosphate isomerase determined crystallographically at 2.5 Å resolution. Nature 255, 609–614 (1975).
De Vos, A.M., Ultsch, M. & Kossiakoff, A.A. Human growth hormone and extracellular domain of its receptor: crystal structure of the complex. Science 255, 306–312 (1992).
Koshland, D.E. Stereochemistry and the mechanism of enzymatic reactions. Biol. Rev. 28, 416–436 (1953).
Paulsen, H. Fortschritte bei der selektiven chemischen Synthese komplexer Oligosaccharide Angew. Chem. Int. Ed.. Engl. 21, 155–173 (1982).
Sinnot, M.L. Stereochemistry and the mechanisms of enzymatic glycosyl transfer. Chem Rev. 90, 1171–1202 (1990).
McCarter, J.D. & Withers, S. Mechanisms of enzymatic glycoside hydrolysis. Curr. Op. Struct. Biol. 4, 885–892 (1994).
Mo, F. & Jensen, L.H. The crystal structure of β-(1→4) linked disaccharide, α-N,N′-diacetylchitobiose monohydrate. Acta Cryst. B34, 1562–1569 (1978).
Lai, E.C.K. & Withers, S.G. Stereochemistry and kinetics of the hydration of 2-acetamido-D-glucal by β-N-acetylhexosaminidases. Biochem. 33, 14743–14749 (1994).
Thunissen, A.-M.W.H. et al. Doughnut-shaped structure of a bacterial muraminidase revealed by X-ray crystallography. Nature 367, 750–753 (1994).
Weaver, L.H., Grütter, M.G. & Matthews, B.W. The refined structures of goose lysozyme and its complex with a bound trisaccharide show that the “goose-type” lysozyme lack a catalyticaspartate residue. J. Mol. Biol. 245, 54–68 (1995).
Terwisscha van Scheltinga, A.C. et al. Stereochemistry of chitin hydrolysis by a plant chitinase/lysozyme and X-ray structure of a complex with allosamidin. Evidence for substrate assisted catalysis. Biochem. 34, 15619–115623 (1995).
Lowe, G. & Sheppard, G. Acetamido-group participation in lysozyme catalysis. J. Chem. Soc Chem. Commun. 529–530 (1968).
Soto-Gil, R.W. & Zyskind, J.W. N,N′-Diacetylchitobiase of Vibrio harveyi. J. Biol. Chem. 264, 14778–14783 (1989).
Somerville, C.C. & Colwell, R.R. Sequence analysis of the b-N-acetylhexosaminidase gene from Vibrio vulnificus: Evidence for a common evolutionary origin of hexosaminidases. Proc. Natl. Acad, Sci. USA 90, 6751–6755 (1993).
Beccari, T., Hoade, J., Orlacchio, A. & Stirling, J.L. Cloning and sequence analysis of a cDNA encoding the α-subunit of mouse β-N-acetylhexosaminidase and comparison with the human enzyme. Biochem. J. 285, 593–596 (1992).
Myerowitz, R., Piekarz, R., Neufeld, E.F., Shows, T.B. & Suzuki, K. human β-hexosaminidase a chain: Coding sequence and homology with the β chain. Proc. Natl. Acad. Sci. USA 82, 7830–7834 (1985).
Bapat, B., Ethier, M., Neote, K., Mahuran, D. & Gravel, R.A. Cloning and sequence analysis of the cDNA encoding the β-subunit of mouse β-hexosaminidase. FEBS Lett. 237, 191–195 (1988).
Muldoon, L.L., Neuwelt, E.A., Pagel, M.A. & Weiss, D.L. Characterization of the molecular defect in feline model for type II GM2 gangliosidoesis (Sandhoff disease). Am. J. Pathol. 144, 1109–1118 (1994).
Neote, K. et al. Characterization of the human HEXB gene encoding lysosomal β-hexosaminidase. Genomics 3, 279–286 (1988).
Korneluk, R.G. et al. Isolation of cDNA clones coding for the α-subunit of human β-hexosaminidase. J. Biol. Chem. 261, 8407–8413 (1986).
Higgins, D.G., Bleasby, A.J. & Fuchs, R. CLUSTAL V: improved software for multiple sequence alignment. CABIOS 8, 189–191 (1992).
Sander, C. & Schneider, R. Database of homogy-derived structures and the structurally meaning of sequence alignment. Proteins 9, 56–68 (1991).
Chothia, C. & Lesk, A.M. The relation between the divergence of sequence and structure in protein. EMBO J. 5, 823–826 (1986).
Rost, B. & Sander, C. Combining evolutionary information and neural networks to predict protein secondary structure. Proteins 19, 55–72 (1994).
Vriend, G. WHAT-IF: A molecular modelling and drug design program. J. Mol. Graph. 8, 52–56 (1990).
Chinea, G., Padron, G., Hooft, R.W.W., Sander, C. & Vriend, G. The use of protein-specific rotamers in model building by homology. Proteins 23, 415–421 (1995).
de Fillippis, V., Sander, C. & Vriend, G. Predicting local structural changes that result from point mutations. Prot. Engng. 7, 1203–1208 (1994).
Kytzia, H.-J. & Sandhoff, K. Evidence for two different active sites on human β-hexosaminidase A. J. Biol. Chem. 260, 7568–7572 (1985).
Tanaka, A. et al. GM2-gangliosidosis B1 variant: analysis of β-hexosaminidase a gene abnormalities in seven patients. Am. J. Hum. Genet. 46, 329–339 (1990).
Ohno, K. & Suzuki, K. Mutation in GM2 gangliosidosis B1 variant. J. Neurochem. 50, 316–318 (1988).
Triggs-Raine, B.L., Akerman, B.R., Clarke, J.T.R. & Gravel, R.A. Sequence of DNA flanking the exons of the HEXA gene, and identification of mutations in Tay–Sachs Disease. Am. J. Hum. Genet. 49, 1041–1054 (1991).
Akli, S., Chelly, J., Lacorte, J.-M., Poenaru, L. & Kahn, A. Seven novel Tay–Sachs mutations detected by chemical mismatch cleavage of PCR-amplified cDNA fragments. Genomics 11, 124–134 (1991).
Fernandes, M. et al. A new Tay–Sachs disease B1 allele in exon 7 in two compound heterozygotes each with a second novel mutation. Hum. Mol. Genet. 1, 759–761 (1992).
Tanaka, A., Punnett, H.H. & Suzuki, K. A new point mutation in the β-hexosaminidase a subunit gene responsible for infantile Tay–Sachs Disease in a non-Jewish Caucasian patient (a Kpn Mutant). Am. J. Hum. Genet. 47, 567–574 (1990).
Brown, C.A., Neote, K., Leung, A., Gravel, R.A. & Mahuran, D.J. Introduction of the a subunit mutation associated with the B1 variant of Tay–Sachs Disease into the β subunit produces a β-hexosaminidase B without catalytic activity. J. Biol. Chem. 264, 21705–21710 (1989).
Brown, C.A. & Mahuran, D.J. Active arginine residues in β-hexosaminidases. J. Biol. Chem. 266, 15855–15862 (1991).
Nakano, T. et al. A new point mutation within exon 5 of β-hexosaminidase a gene in a Japanese infant with Tay–Sachs Disease. Ann. Neurol. 27, 465–473 (1990).
Nakano, T., Muscillo, M., Ohno, K., Hoffman, A.J. & Suzuki, K. A point mutatation in the coding sequence of the β-hexosaminidase a gene results in the enzyme protein in an unusual GM2-gangliosidosis variant. J. Neurochem. 51, 984–987 (1988).
Akalin, N. et al. Tay–Sachs Disease in China: two new mutations and a “Macro Polo” allele. Am. J. Hum. Genet. 49, A2246 (1991).
Ainsworth, P.J. & Coulter-Mackie, M.B. A double mutation in exon 6 of the β-hexosaminidase α subunit in a patient with the B1 variant of Tay–Sachs Disease. Am. J. Hum. Genet. 51, 802–809 (1992).
Coulter-Mackie, M.B. Molecular characterization of both alleles in an unusual Tay–Sachs Disease B1 variant. Am. J. Hum. Genet. 54, 1126–1127 (1994).
Triggs-Raine, B.L. et al. A pseudodeficiency allele common in non-Jewish Tay–Sachs carriers: implications for carrier screening. Am. J. Hum. Genet. 51, 793–801 (1992).
Cao, Z. et al. A second mutation associated with apparent β-hexosaminidase A pseudodeficiency: identification and frequency estimation. Am. J. Hum. Genet. 53, 1198–1205 (1993).
Bolhuis, P.A., Ponne, N.J., Bikker, H., Baas, F. & deJong, V.J.M.B. Molecular basis of an adult form of Sandhoff disease: substitution of glutamine for arginine at position 505 of the β-chain of β-hexosaminidase results in a labile enzyme. Biochim. Biophys. Acta. 1182, 142–146 (1993).
Petroulakis, E., Cao, Z., Salo, T., Clarke, J. & Triggs-Raine, B. A Trp474Cys mutation in the alpha-subunit of beta-hexosaminidase causes the subacute encephalopathic form of GM2 gangliosidosis, Type 1. Am. J. Hum. Genet. 55, A2129 (1994).
Cao, Z., Petroulakis, E., Salo, T. & Triggs-Raine, B. Expression of the HEXA mutations Arg247Trp and Arg249Trp, associated with beta-hexosaminidase A pseudodeficiency. Am. J. Hum. Genet. 55, A1251 (1994).
Navon, R. & Proia, R.L. The mutations in Ashkenazi Jews with adult GM2 gangliosidosis, the adult form of Tay–Sachs Disease. Science 243, 1471–1474 (1989).
Paw, B.H., Kaback, M.M. & Neufeld, E.F. Molecular basis of adult-onset and chronic GM2 gangliosidosis of Ashkenazi Jewish origin: substitution of serine for glycine at position 269 of the α-subunit of β-hexosaminidase. Proc. Natl. Acad. Sci. USA 86, 2413–2417 (1989).
Brown, C.A. & Mahuran, D.J. β-hexosaminidase Isozymes from cells cotransfected with α and β cDNA constructs: analysis of the α-subunit missense mutation associated with the adult form of Tay–Sachs Disease. Am. J. Hum. Genet. 53, 497–508 (1993).
Kless, H., Sitrit, Y., Chet, I. & Oppenheim, A.B. Cloning of the gene coding for chitobiase of Serratia marcescens. Mol. Gen. Genet. 217, 471–473 (1989).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Meths. Enzymol. 276 (in the press).
Furey, W. & Swaminathan, S. PHASES - a program package for the processing and analysis of diffraction data from macromolecules. American Ciystallographic Association Meeting Abstracts 18, 73 (1990).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Brünger, A.T. Assessment of phase accuracy by cross validation: the free R value. Methods and application. Acta Crystallogr. D49, 24–36 (1993).
Konnert, J.H. & Hendrickson, W.A. A restrained-parameter thermal-factor refinement procedure. Acta Crystallogr. A36, 344–350 (1980).
Collaborative Computer Project, Number 4 (CCP4) The CCP4 Suite: Programs for Protein Crystallography. Acta Crystallogr. D50, 760–763 (1994).
Lamzin, V.S. & Wilson, K.S. Automated refinement of protein models. Acta Crystallogr. D49, 129–147 (1993).
Holm, L. & Sander, C. Protein Structure Comparison by Alignment of Distance Matrices. J. Mol. Biol. 233, 123–138 (1993).
van Gunsteren, W.F. & Berendsen, H.J. GROMOS. BIOMOS, Biomolecular Software, University of Groningen, the Netherlands.
Kraulis, P. MOLSCRIPT: a program to produce both detailed and schematic plot of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Navon, R. & Proia, R.L. Tay–Sachs Disease in Moroccan Jews: deletion of a phenylalanine in the α-subunit of β-hexosaminidase. Am. J. Hum. Genet. 48, 412–419 (1991).
Banerjee, P. et al. Molecular basis of an adult form of β-hexosaminidase B deficiency with motor neuron disease. Biochem. Biophys. Res. Comm. 181, 108–115 (1991).
Mules, E.H., Hayflick, S., Miller, C.S., Reynolds, L.W. & Thomas, G.H. Six novel deleterious and three neutral mutations in the gene encoding the α-subunit of the hexosaminidase A in non-Jewish individuals. Am. J. Hum. Genet. 50, 834–841 (1992).
Paw, B.H. et al. Juvenile GM2 gangliosidosis caused by substitution of histidine for arginine at position 499 or 504 of the α-subunit of β-hexosaminidase. J. Biol. Chem. 265, 9452–9457 (1990).
Paw, B.H., Wood, L.C. & Neufeld, E.F. A third mutation at the CpG dinucleotide of codon 504 and a silent mutation at codon 506 of the HEXA Gene. Am. J. Hum. Genet. 48, 1139–1146 (1991).
Fernandes, M., Boulay, B., Hechtmann, P., Kaplan, F. & Strasberg, P. Five novel HEXA mutations in non-Jewish Tay–Sachs disease (TSD) patients. Am. J. Hum. Genet. 51, A656 (1992).
Author information
Authors and Affiliations
Author notes
From September 1996 C.E.V will move to: Athens University, Biology Department, Biochemistry Laboratory, Panepistimiopoli, Kouponia, 15701 Athens, Greece
- Constantin E. Vorgias
Rights and permissions
About this article
Cite this article
Tews, I., Perrakis, A., Oppenheim, A. et al. Bacterial chitobiase structure provides insight into catalytic mechanism and the basis of Tay–Sachs disease. Nat Struct Mol Biol 3, 638–648 (1996). https://doi.org/10.1038/nsb0796-638
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0796-638