Abstract
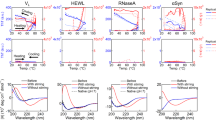
It is generally assumed that folding intermediates contain partially formed native-like secondary structures. However, if we consider the fact that the conformational stability of the intermediate state is simpler than that of the native state, it would be expected that the secondary structures in a folding intermediate would not necessarily be similar to those of the native state. β-Lactoglobulin is a predominantly β-sheet protein, although it has a markedly high intrinsic preference for α-helical structure. We have studied the refolding kinetics of bovine β-lactoglobulin using stopped-flow circular dichroism and find that a partly α-helical intermediate accumulates transiently before formation of the native β-sheets. The present results suggest that the folding reaction of β-lactoglobulin follows a non-hierarchical mechanism, in which non-native α-helical structures play important roles.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Anfinsen, C.B. Principles that govern the folding of protein chains. Science 181, 223–230 (1973).
Dill, K.A. et al. Principles of protein folding - A perspective from simple exact models. Protein Sci. 4, 561–602 (1995).
Kuwajima, K. The molten globule state as a clue for understanding the folding and cooperativity of globular-protein structure. Proteins: Struct. Funct. Genet. 6, 87–103 (1989).
Ptitsyn, O.B. The molten globule state. In Protein Folding (ed. Creighton, T.E.) 243–300 (W. H. Freeman and Company, New York, 1992).
Jennings, P.A. & Wright, P.E. Formation of a molten globule intermediate early in the kinetic folding pathway of apomyoglobin. Science 262, 892–896 (1993)
Balbach, J. et al. Following protein folding in real time using NMR spectroscopy. Nature Struct. Biol. 2, 865–870 (1995).
Fersht, A.R. Optimization of rates of protein folding: The nucleation-condensation mechanism and its implications. Proc. Natl. Acad. Sci. USA 92, 10869–10873 (1995).
Shakhnovich, E.I. Modeling protein folding: the beauty and power of simplicity. Folding & Design 1, R50–R54 (1996).
Baldwin, R.L. The nature of protein folding pathways: The classical versus the new view. J. Biochem. NMR 5, 103–109 (1995).
Itzhaki, L.S., Otzen, D.E. & Fersht, A.R. The structure of the transition state for folding of chymotrypsin inhibitor 2 analysed by protein engineering methods: Evidence for a nucleation-condensation mechanism for protein folding. J. Mol. Biol. 254, 260–288 (1995).
Pervaiz, S. & Brew, K. Homology of β-lactoglobulin, serum retinol-binding protein, and protein HC. Science 228, 335–337 (1985).
Papiz, M.Z. et al. The structure of β-lactoglobulin and its similarity to plasma retinol-binding protein. Nature 324, 383–385 (1986).
Monaco, H.L. et al. Crystal structure of the trigonal form of bovine β-lactoglobulin and of its complex with retinol at 2.5 Å resolution. J. Mol. Biol. 197, 695–706 (1987).
Fugate, R.G. & Song, P.-S. Spectroscopic characterization of β-lactoglobulin-retinol complex. Biochim. Biophys. Acta 625, 28–42 (1980).
Kuwajima, K., Yamaya, H., Miwa, S., Sugai, S. & Nagamura, T. Rapid formation of secondary structure framework in protein folding studied by stopped-flow circular dichroism. FEBS Lett. 221, 115–118 (1987).
Kuwajima, K. Stopped-flow circular dichroism. In Circular dichroism and the conformational analysis of biomolecules (ed. Fasman, G.D.), 159–182 (Plenum, New York, 1996).
Nishikawa, K. & Noguchi, T. Predicting protein secondary structure based on amino acid sequence. Methods Enzymol. 202, 21–24 (1991).
Shiraki, K., Nishikawa, K. & Goto, Y. Trifluoroethanol-induced stabilization of the α-helical structure of β-lactoglobulin: Implication for non-hierarchical protein folding. J. Mol. Biol. 245, 180–194 (1995).
Hamada, D., Kuroda, Y., Tanaka, T. & Goto, Y. High helical propensity of the peptide fragments derived from β-lactoglobulin, a predominantly β-sheet protein. J. Mol. Biol. 254, 737–746 (1995).
Kuroda, Y., Hamada, D., Tanaka, T. & Goto, Y. High helicity of peptide fragments corresponding to β-strand regions of β-lactoglobulin observed by 2D-NMR spectroscopy. Folding & Design 1, 243–251 (1996).
Dufour, E., Bertrand, H.C. & Haertle, T. Reversible effects of medium dielectric constant on structural transformation of β-lactoglobulin and its retinol binding. Biopolymers 33, 589–598 (1993).
Liu, Z.-P., Rizo, J. & Gierasch, L.M. Equilibrium folding studies of cellular retinoic acid binding protein, a predominantly β-sheet protein. Biochemistry 33, 134–142 (1994).
Chaffote, A.F., Guillou, Y. & Goldberg, M.E. Kinetic resolution of peptide bond and side chain far-UV circular dichroism during the folding of hen egg lysozyme. Biochemistry 31, 9694–9703 (1992).
Chen, Y.H., Yang, J.T. & Chau, K.H. Determination of the helix and β form of proteins in aqueous solution by circular dichroism. Biochemistry 20, 33–37 (1994).
Filimonov, V.V., Prieto, J., Martinez, J.C., Bruix, M., Mateo, P.L., Serrano, L. Thermodynamic analysis of the chemotactic protein from Escherichia coli, CheY. Biochemistry 47, 12906–12921 (1994).
López-Hernández, E. & Serrano, L. Structure of the transition state for folding of the 129aa protein CheY resembles that of a smaller protein, CI-2. Folding & Design 1, 43–55 (1996).
Wright, P.E., Dyson, H.J. & Lerner, R.A. Conformation of peptide fragments of proteins in aqueous solution: Implication for initiation of protein folding. Biochemistry 27, 7164–7175 (1988).
Dyson, H.J., Merutka, G., Waltho, J.P., Lerner, R.A. & Wright, P.E. Folding of peptide fragments comprising the complete sequence of proteins. Models for initiation of protein folding. I. Myohemerythrin. J. Mol. Biol. 226, 795–817 (1992).
Dyson, H.J. et al. Folding of peptide fragments comprising the complete sequence of proteins. Models for initiation of protein folding. II. Plastocyanin. J. Mol. Biol. 226, 819–835 (1992).
Waterhous, D.V. & Johnson, W.C., Jr. Importance of environment in determining secondary structure in proteins. Biochemistry 33, 2121–2128 (1994).
Minor Jr., D.L. & Kim, P.S. Context-dependent secondary structure formation of a designed protein sequence. Nature 380, 730–734 (1996).
Mottonen, J. et al. Structural basis of latency in plasminogen activator inhibitor-1. Nature 355, 27–273 (1992).
Sancho, E., Declerck, P.J., Price, N.C., Kelly, S.M. & Booth, N.A. Conformational studies on plasminogen activator inhibitor (PAI-1) in -active, latent, substrate, and cleaved forms. Biochemistry 34, 1064–1069 (1995).
Gutin, A.M., Abkevich, V.I. & Shakhnovich, E.I. Is burst hydrophobic collapse necessary for protein folding? Biochemistry 34, 3066–3076 (1995).
Arcus, V.L., Vuilleumier, S., Freund, S.M.V., Bycroft, M. & Fersht, A.R. A comparison of the pH, urea, and temperature-denatured states of barnase by heteronuclear NMR: Implications for the initiation of protein folding. J. Mol. Biol. 254, 305–321 (1995).
Radford, S.E. & Dobson, C.M. Insight into protein folding using physical techniques: studies of lysozyme and α-lactalbumin. Phil. Trans. R. Soc. Lond. B. 348, 17–25 (1995).
Kiefhaber, T. Kinetic traps in lysozyme folding. Proc. Natl. Acad. Sci. USA 92, 9029–9033 (1995).
Bryngelson, J.P., Onuchic, J.N., Socci, N.D. & Wolynes, P.G. Funnels, pathways, and the energy landscape of protein folding: a synthesis. Proteins: Struct. Funct. Genet. 21, 167–195 (1995).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 956–950 (1991).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: Pattern recognition of hydrogen–bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hamada, D., Segawa, Si. & Goto, Y. Non-native α-helical intermediate in the refolding of β-lactoglobulin, a predominantly β-sheet protein. Nat Struct Mol Biol 3, 868–873 (1996). https://doi.org/10.1038/nsb1096-868
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1096-868
This article is cited by
-
Characterization and structure of cold-extruded whey protein isolate: impact of ball milling
Applied Nanoscience (2019)
-
Rapid α-oligomer formation mediated by the Aβ C terminus initiates an amyloid assembly pathway
Nature Communications (2016)
-
Amphiphilic α-Helical Potential: A Putative Folding Motif Adding Few Constraints to Protein Evolution
Journal of Molecular Evolution (2011)
-
From Levinthal to pathways to funnels
Nature Structural & Molecular Biology (1997)