Abstract
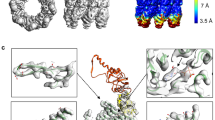
Improved refinement of the crystal structure of GroEL from Escherichia coli has resulted in a complete atomic model for the first 524 residues. A new torsion-angle dynamics method and non-crystallographic symmetry restraints were used in the refinement. The model indicates that conformational variability exists due to rigid-body movements between the apical and intermediate domains of GroEL, resulting in deviations from strict seven-fold symmetry. The regions of the protein involved in polypeptide and GroES binding show unusually high B factors; these values may indicate mobility or discrete disorder. The variability of these regions may play a role in the ability of GroEL to bind a wide variety of substrates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Anfinsen, C.B. Principles that govern the folding of protein chains. Science 181, 223–30 (1973).
Ellis, J. Proteins as molecular chaperones. Nature 328, 378–379 (1987).
Horwich, A.L., Low, K.B., Fenton, W.A., Hirshfield, I.N. & Furtak, K. Folding in vivo of bacterial cytoplasmic proteins: role of GroEL. Cell 74, 909–917 (1993).
Horwich, A.L. & Willison, K.R. Protein folding in the cell: functions of two families of molecular chaperone, hsp 60 and TF55-TCP1. Phil. Trans. R. Soc. London B 339 313–326 (1993).
Braig, K., Simon, M., Furuya, F., Hainfeld, J.F. & Horwich, A.L. A polypeptide bound by the chaperonin GroEL is localized within a central cavity. PNAS 90, 397–882 (1993).
Martin, J. et al. Chaperonin-mediated protein folding at the surface of GroEL through a “molten globule”-like intermediate. Nature 352, 36–42 (1991).
Landry, S.J., Jordan, R., McMacken, R. & Gierasch, L.M. Different conformations for the same polypeptide bound to chaperones DnaK and GroEL. Nature 355, 455–457 (1992).
Harris, J.R., Pluckthun, A. & Zahn, R. Transmission electron microscopy of GroEL, GroES, and the symmetrical GroEL/ES complex. J. of Struct. Biol. 112, 216–230 (1994).
Jackson, G.S. et al. Binding and hydrolysis of nucleotides in the chaperonin catalytic cycle: implications for the mechanism of assisted protein folding. Biochemistry 32, 2554–2563 (1993).
Todd, M.J., Viitanen, P.V. & Lorimer, G.H. Dynamics of the chaperonin ATPase cycle: implications for facilitated protein folding. Science 265, 659–666 (1993).
Weissman, J.S., Kashi, Y., Fenton, W.A. & Horwich, A.L. GroEL-mediated protein folding proceeds by multiple rounds of binding and release of nonnative forms. Cell 78, 693–702 (1994).
Braig, K. et al. The crystal structure of the bacterial chaperonin GroEL at 2.8 Å. Nature 371, 578–586 (1994).
Fenton, W.A., Kashi, Y., Furtak, K. & Horwich, A.L. Residues in chaperonin GroEL required for polypeptide binding and release. Nature 371, 614–619 (1994).
Hendrickson, W.A. Stereochemically restrained refinement of macromolecular structures. Meths. Enzymol. 115, 252–270 (1985).
Brünger, A.T., Kuriyan, J. & Karplus, M. Crystallographic R-factor refinement by molecular dynamics. Science 235, 458–460 (1987).
Brünger, A.T. The free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–474 (1992).
Rice, L.M. & Brünger, A.T. Torsion angle dynamics: reduced variable conformational sampling enhances crystallographic structure refinement. Proteins: Struct. Funct. Genet. 19, 277–290 (1994).
Weis, W.I., Brünger, A.T., Skehel, J.J. & Wiley, D.C. Refinement of the influenza virus haemagglutinin by simulated annealing. J. molec. Biol. 212, 737–761 (1990).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Cryst. A47, 110–119 (1991).
Brünger, A.T. X-PLOR Version 3.1 (Yale University, New Haven, CT, 1992)
Read, R.J., Fourier coefficients for maps using phases from partial structures with errors. Acta Cryst. A42, 140–149 (1986).
Brünger, A.T. Crystallographic refinement by simulated annealing:application to a 2.8Å resolution structure of aspartate aminotransferase. J. molec. Biol. 203, 803–816 (1988).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. Main-chain bond lengths and bond angles in protein structures. J. app. Cryst. 26, 283–291 (1993).
Morris, A.L., MacArthur, M.W., Hutchinson, E.G. & Thornton, J.M. Stereochemical quality of protein structure coordinates. Proteins 12, 345–364 (1992).
Luzzati, P.V. Traitement statistique des erreurs dans la determination des structures cristallines. Acta Crystallogr. 5, 802–810 (1952).
Weissman, J.S. et al. Mechanism of GroEL action: productive release of polypeptide from a sequestered position under GroES. Cell 83, 1–20 (1995).
Kleywegt, G.J. et al. Crystal structure of cellular retinoic acid binding proteins I and II in complex with all- trans -retinoic acid and a synthetic retinoid. Structure 2, 1241–1258 (1994).
Brünger, A.T. The free Rvalue: A more objective statistic for crystallography. Meths. Enzymol. in the press.
Viitanen, P.V., Gatenby, A.A. & Lorimer, G.H. Purified chaperonin 60 (GroEL) interacts with the nonnative states of a multitude of Escherichia coli proteins. Prot. Sci. 1, 363–369 (1992).
Chen, S., Roseman, A.M., Hunter, A.S., Wood, S.P., Burston, S.G., Ranson, N.A., Clarke, A.R. & Saibil, H.R. Location of a folding protein and shape changes in GroEL-GroES complexes imaged by cryo-electron microscopy. Nature 371, 261–264 (1994).
Langer, T., Pfeifer, G., Martin, J., Baumeister, W. & Haiti, F.U. Chaperonin-mediated protein folding: GroES binds to one end of the GroEL cylinder, which accomodates the protein substrate within its central cavity. EMBO J. 11, 4757–4765 (1992).
Nilges, M., Clore, G.M. & Gronenborn, A.M. Determination of three-dimensional structures of proteins from interproton distance data by hybrid distance geometry-dynamical simulated annealing calculations. FEBS Letts. 229, 317–324 (1988).
Engh, R.A. & Huber, R. Accurate bond and angle parameters for X-ray protein-structure refinement. Acta Cryst. A47, 392–400 (1991).
Berendsen, H.J.C., Postma, J.P.M., van Gunsteren, W.F., DiNola, A. & Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Hockney, R.W. & Eastwood, J.W. Computer simulation using particles. (McGraw-Hill, New York, 1981).
Hutchinson, G. & Thornton, J. Promotif A program to analyse structural motifs in a protein, (manuscript in preparation).
Lüthy, R., Bowie, J.U. & Eisenberg, D. Assessment of protein models with three-dimensional profiles. Nature 356, 83–85 (1992).
Evans, S.V. SETOR: hardware lighted three-dimensional solid model representations of macromolecules. J. molec. Graphics 11, 134–138 (1993).
Kraulis, P.J. Molscript: a program to produce both detailed and schematic plots of protein structures. J. appl. Cryst. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. Raster3D version 2. 0. A program for photorealistic molecular graphics. Acta Crystallogr. D50, 869–873 (1994).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Braig, K., Adams, P. & Brünger, A. Conformational variability in the refined structure of the chaperonin GroEL at 2.8 Å resolution. Nat Struct Mol Biol 2, 1083–1094 (1995). https://doi.org/10.1038/nsb1295-1083
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1295-1083
This article is cited by
-
Ten computer codes that transformed science
Nature (2021)
-
Crystal structure of P. falciparum Cpn60 bound to ATP reveals an open dynamic conformation before substrate binding
Scientific Reports (2021)
-
GroEL/ES mediated the in vivo recovery of TRAIL inclusion bodies in Escherichia coli
Scientific Reports (2018)
-
Unraveling low-resolution structural data of large biomolecules by constructing atomic models with experiment-targeted parallel cascade selection simulations
Scientific Reports (2016)
-
Suppression of amyloid fibrils using the GroEL apical domain
Scientific Reports (2016)