Abstract
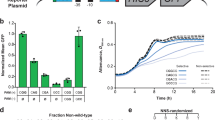
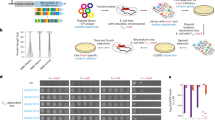
To optimize the in vivo folding of proteins, we linked protein stability to antibiotic resistance, thereby forcing bacteria to effectively fold and stabilize proteins. When we challenged Escherichia coli to stabilize a very unstable periplasmic protein, it massively overproduced a periplasmic protein called Spy, which increases the steady-state levels of a set of unstable protein mutants up to 700-fold. In vitro studies demonstrate that the Spy protein is an effective ATP-independent chaperone that suppresses protein aggregation and aids protein refolding. Our strategy opens up new routes for chaperone discovery and the custom tailoring of the in vivo folding environment. Spy forms thin, apparently flexible cradle-shaped dimers. The structure of Spy is unlike that of any previously solved chaperone, making it the prototypical member of a new class of small chaperones that facilitate protein refolding in the absence of energy cofactors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Kerner, M.J. et al. Proteome-wide analysis of chaperonin-dependent protein folding in Escherichia coli. Cell 122, 209–220 (2005).
Duguay, A.R. & Silhavy, T.J. Quality control in the bacterial periplasm. Biochim. Biophys. Acta 1694, 121–134 (2004).
Foit, L. et al. Optimizing protein stability in vivo. Mol. Cell 36, 861–871 (2009).
Stafford, S.J., Humphreys, D.P. & Lund, P.A. Mutations in dsbA and dsbB, but not dsbC, lead to an enhanced sensitivity of Escherichia coli to Hg2+ and Cd2+. FEMS Microbiol. Lett. 174, 179–184 (1999).
De Pascale, G. & Wright, G.D. Antibiotic resistance by enzyme inactivation: from mechanisms to solutions. ChemBioChem 11, 1325–1334 (2010).
Spence, G.R., Capaldi, A.P. & Radford, S.E. Trapping the on-pathway folding intermediate of Im7 at equilibrium. J. Mol. Biol. 341, 215–226 (2004).
Raffa, R.G. & Raivio, T.L. A third envelope stress signal transduction pathway in Escherichia coli. Mol. Microbiol. 45, 1599–1611 (2002).
MacRitchie, D.M., Buelow, D.R., Price, N.L. & Raivio, T.L. Two-component signaling and gram negative envelope stress response systems. Adv. Exp. Med. Biol. 631, 80–110 (2008).
Bury-Moné, S. et al. Global analysis of extracytoplasmic stress signaling in Escherichia coli. PLoS Genet. 5, e1000651 (2009).
Price, N.L. & Raivio, T.L. Characterization of the Cpx regulon in Escherichia coli strain MC4100. J. Bacteriol. 191, 1798–1815 (2009).
Raivio, T.L., Laird, M.W., Joly, J.C. & Silhavy, T.J. Tethering of CpxP to the inner membrane prevents spheroplast induction of the cpx envelope stress response. Mol. Microbiol. 37, 1186–1197 (2000).
Hartl, F.U. & Hayer-Hartl, M. Converging concepts of protein folding in vitro and in vivo. Nat. Struct. Mol. Biol. 16, 574–581 (2009).
Winter, J. & Jakob, U. Beyond transcription—new mechanisms for the regulation of molecular chaperones. Crit. Rev. Biochem. Mol. Biol. 39, 297–317 (2004).
Zoetendal, E.G., Smith, A.H., Sundset, M.A. & Mackie, R.I. The BaeSR two-component regulatory system mediates resistance to condensed tannins in Escherichia coli. Appl. Environ. Microbiol. 74, 535–539 (2008).
Scalbert, A. Antimicrobial properties of tannins. Phytochemistry 30, 3875–3883 (1991).
Hernes, P.J. et al. Tannin diagenesis in mangrove leaves from a tropical estuary: A novel molecular approach. Geochim. Cosmochim. Acta 65, 3109–3122 (2001).
Smith, A.H. & Mackie, R.I. Effect of condensed tannins on bacterial diversity and metabolic activity in the rat gastrointestinal tract. Appl. Environ. Microbiol. 70, 1104–1115 (2004).
Serrano, J., Puupponen-Pimia, R., Dauer, A., Aura, A.M. & Saura-Calixto, F. Tannins: current knowledge of food sources, intake, bioavailability and biological effects. Mol. Nutr. Food Res. 53, S310–S329 (2009).
Howell, A.B., Vorsa, N., Marderosian, A.D. & Foo, L.Y. Inhibition of the adherence of P-fimbriated Escherichia coli to uroepithelial-cell surfaces by proanthocyanidin extracts from cranberries. N. Engl. J. Med. 339, 1085–1086 (1998).
Zanchi, D. et al. Tannin-assisted aggregation of natively unfolded proteins. Europhys. Lett. 82, 58001 (2008).
Hofmann, T. et al. Protein binding and astringent taste of a polymeric procyanidin, 1,2,3,4,6-penta-O-galloyl-beta-D-glucopyranose, castalagin, and grandinin. J. Agric. Food Chem. 54, 9503–9509 (2006).
Ono, K., Hasegawa, K., Naiki, H. & Yamada, M. Anti-amyloidogenic activity of tannic acid and its activity to destabilize Alzheimer's β-amyloid fibrils in vitro. Biochim. Biophys. Acta 1690, 193–202 (2004).
Kim, W. et al. A high-throughput screen for compounds that inhibit aggregation of the Alzheimer's peptide. ACS Chem. Biol. 1, 461–469 (2006).
Lee, L.L., Ha, H., Chang, Y.T. & DeLisa, M.P. Discovery of amyloid-β aggregation inhibitors using an engineered assay for intracellular protein folding and solubility. Protein Sci. 18, 277–286 (2009).
Raivio, T.L., Popkin, D.L. & Silhavy, T.J. The Cpx envelope stress response is controlled by amplification and feedback inhibition. J. Bacteriol. 181, 5263–5272 (1999).
Danese, P.N. & Silhavy, T.J. CpxP, a stress-combative member of the Cpx regulon. J. Bacteriol. 180, 831–839 (1998).
DiGiuseppe, P.A. & Silhavy, T.J. Signal detection and target gene induction by the CpxRA two-component system. J. Bacteriol. 185, 2432–2440 (2003).
Isaac, D.D., Pinkner, J.S., Hultgren, S.J. & Silhavy, T.J. The extracytoplasmic adaptor protein CpxP is degraded with substrate by DegP. Proc. Natl. Acad. Sci. USA 102, 17775–17779 (2005).
Kwon, E., Kim, D.Y., Gross, C.A., Gross, J.D. & Kim, K.K. The crystal structure Escherichia coli Spy. Protein Sci. 19, 2252–2259 (2010).
Powl, A.M., Wright, J.N., East, J.M. & Lee, A.G. Identification of the hydrophobic thickness of a membrane protein using fluorescence spectroscopy: Studies with the mechanosensitive channel MscL. Biochemistry 44, 5713–5721 (2005).
Tapley, T.L. et al. Structural plasticity of an acid-activated chaperone allows promiscuous substrate binding. Proc. Natl. Acad. Sci. USA 106, 5557–5562 (2009).
Brynildsen, M.P. & Liao, J.C. An integrated network approach identifies the isobutanol response network of Escherichia coli. Mol. Syst. Biol. 5, 277 (2009); doi:10.1038/msb.2009.34.
Rutherford, B.J. et al. Functional genomic study of exogenous n-butanol stress in Escherichia coli. Appl. Environ. Microbiol. 76, 1935–1945 (2010).
Miyawaki, O. & Tatsuno, M. Thermodynamic analysis of alcohol effect on thermal stability of proteins. J. Biosci. Bioeng. 111, 198–203 (2011).
Neidhardt, F.C., VanBogelen, R.A. & Vaughn, V. The genetics and regulation of heat-shock proteins. Annu. Rev. Genet. 18, 295–329 (1984).
Neidhardt, F.C. et al. Identity of the B56.5 protein, the A-protein, and the groE gene product of Escherichia coli. J. Bacteriol. 145, 513–520 (1981).
McNulty, B.C., Young, G.B. & Pielak, G.J. Macromolecular crowding in the Escherichia coli periplasm maintains α-synuclein disorder. J. Mol. Biol. 355, 893–897 (2006).
Tapley, T.L., Franzmann, T.M., Chakraborty, S., Jakob, U. & Bardwell, J.C.A. Protein refolding by pH-triggered chaperone binding and release. Proc. Natl. Acad. Sci. USA 107, 1071–1076 (2010).
Miller, J.H. in A Short Course in Bacterial Genetics: A Laboratory Manual and Handbook for Escherichia coli and Related Bacteria (Cold Spring Harbor Laboratory Press, Woodbury, NY, 1992).
Hiniker, A. & Bardwell, J.C.A. In vivo substrate specificity of periplasmic disulfide oxidoreductases. J. Biol. Chem. 279, 12967–12973 (2004).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Meth. Enzymol. 276, 307–326 (1997).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We thank G. Morgan and S. Radford for communicating results before publication and for the Im7-L53A I54A protein, T. Franzmann for conducting the ultracentrifugation experiments, D. Reichmann, C. Cremers and A. Malik for advice, M. Lei and Y. Chen for providing vector and reagents for Spy purification and C. Munger for initial purification and crystallization of Spy. Protein identification was done by the Protein Structure Facility at the University of Michigan. Diffraction data were collected at the Canadian Macromolecular Crystallography Facility-1 beamline at the Canadian Light Source. We would like to thank S. Labiuk for data collection. The Howard Hughes Medical Institute funded this work (to J.C.A.B). M.C. acknowledges financial support from Canadian Institutes of Health Research grant GSP-48370.
Author information
Authors and Affiliations
Contributions
J.C.A.B. designed the study and wrote the manuscript, with contributions from M.C. and S.Q. S.Q., P.K., T.T., N.K., K.M.R., R.S., J.P., S.H. and G.R. conducted the experiments and collected and analyzed the data. J.C.A.B., Z.X. and M.C. further analyzed the data. L.F. and U.J. provided technical support and conceptual advice.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1 and Supplementary Methods (PDF 2160 kb)
Rights and permissions
About this article
Cite this article
Quan, S., Koldewey, P., Tapley, T. et al. Genetic selection designed to stabilize proteins uncovers a chaperone called Spy. Nat Struct Mol Biol 18, 262–269 (2011). https://doi.org/10.1038/nsmb.2016
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2016
This article is cited by
-
Dynamic coupling of fast channel gating with slow ATP-turnover underpins protein transport through the Sec translocon
The EMBO Journal (2023)
-
Global transcriptomic response of Escherichia coli to p-coumaric acid
Microbial Cell Factories (2022)
-
Trigger factor both holds and folds its client proteins
Nature Communications (2022)
-
Insights into the client protein release mechanism of the ATP-independent chaperone Spy
Nature Communications (2022)
-
Mechanism of the small ATP-independent chaperone Spy is substrate specific
Nature Communications (2021)