Abstract
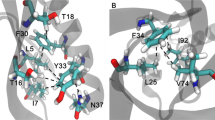
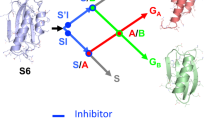
The energetic contributions of hydrogen bonding to protein folding are still unclear, despite more than 70 years of study. This is due partly to the difficulty of extracting thermodynamic information about specific interactions from protein mutagenesis data and partly to the context dependence of hydrogen bond strengths. Herein, we test the hypothesis that hydrogen bond strengths depend on the polarity of their microenvironment, with stronger hydrogen bonds forming in nonpolar surroundings. Double-mutant cycle analysis using a combination of amide-to-ester backbone mutagenesis and traditional side chain mutagenesis revealed that hydrogen bonds can be stronger by up to 1.2 kcal mol−1 when they are sequestered in hydrophobic surroundings than when they are solvent exposed. Such large coupling energies between hydrogen bond strengths and local polarity suggest that the context dependence of hydrogen bond strengths must be accounted for in any comprehensive account of the forces responsible for protein folding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dill, K.A. Dominant forces in protein folding. Biochemistry 29, 7133–7155 (1990).
Pauling, L., Corey, R.B. & Branson, H.R. The structure of proteins: two hydrogen-bonded helical configurations of the polypeptide chain. Proc. Natl. Acad. Sci. USA 37, 205–211 (1951).
Kauzmann, W. Some factors in the interpretation of protein denaturation. Adv. Protein Chem. 14, 1–63 (1959).
Bolen, D.W. & Rose, G.D. Structure and energetics of the hydrogen-bonded backbone in protein folding. Annu. Rev. Biochem. 77, 339–362 (2008).
Avbelj, F. & Baldwin, R.L. Limited validity of group additivity for the folding energetics of the peptide group. Proteins 63, 283–289 (2006).
Avbelj, F. & Baldwin, R.L. Origin of the change in solvation enthalpy of the peptide group when neighboring peptide groups are added. Proc. Natl. Acad. Sci. USA 106, 3137–3141 (2009).
Klotz, I.M. & Franzen, J.S. Hydrogen bonds between model peptide groups in solution. J. Am. Chem. Soc. 84, 3461–3466 (1962).
Krescheck, G.C. & Klotz, I.M. Thermodynamics of transfer of amides from an apolar to an aqueous solution. Biochemistry 8, 8–12 (1969).
Umeyama, H. & Morokuma, K. The origin of hydrogen bonding: an energy decomposition study. J. Am. Chem. Soc. 99, 1316–1332 (1977).
Némethy, G., Steinberg, I.Z. & Scheraga, H.A. Influence of water structure and of hydrophobic interactions on the strength of side-chain hydrogen bonds in proteins. Biopolymers 1, 43–69 (1963).
Fernández, A. & Berry, R.S. Extent of hydrogen-bond protection in folded proteins: a constraint on packing architectures. Biophys. J. 83, 2475–2481 (2002).
Fernandez, A., Zhang, X. & Chen, J.P. Folding and wrapping soluble proteins: Exploring the molecular basis of cooperativity and aggregation. Prog. Nucleic Acid Res. Mol. Biol. 83, 53–87 (2008).
Joh, N.H. et al. Modest stabilization by most hydrogen-bonded side-chain interactions in membrane proteins. Nature 453, 1266–1270 (2008).
Myers, J.K. & Pace, C.N. Hydrogen bonding stabilizes globular proteins. Biophys. J. 71, 2033–2039 (1996).
Albeck, S., Unger, R. & Schreiber, G. Evaluation of direct and cooperative contributions towards the strength of buried hydrogen bonds and salt bridges. J. Mol. Biol. 298, 503–520 (2000).
Blankenship, J.W., Balambika, R. & Dawson, P.E. Probing backbone hydrogen bonds in the hydrophobic core of GCN4. Biochemistry 41, 15676–15684 (2002).
Takano, K., Scholtz, J.M., Sacchettini, J.C. & Pace, C.N. The contribution of polar group burial to protein stability is strongly context-dependent. J. Biol. Chem. 278, 31790–31795 (2003).
Scheike, J.A. et al. Amide-to-ester substitution in coiled coils: the effect of removing hydrogen bonds on protein structure. Angew. Chem. Int. Edn Engl. 46, 7766–7769 (2007).
Deechongkit, S., Dawson, P.E. & Kelly, J.W. Toward assessing the position-dependent contributions of backbone hydrogen bonding to β-sheet folding thermodynamics employing amide-to-ester perturbations. J. Am. Chem. Soc. 126, 16762–16771 (2004).
Deechongkit, S. et al. Context-dependent contributions of backbone hydrogen bonding to β-sheet folding energetics. Nature 430, 101–105 (2004).
Powers, E.T., Deechongkit, S. & Kelly, J.W. Backbone-backbone H-bonds make context-dependent contributions to protein folding kinetics and thermodynamics: Lessons from amide-to-ester mutations. Adv. Protein Chem. 72, 39–78 (2006).
Ferguson, N., Johnson, C.M., Macias, M., Oschkinat, H. & Fersht, A. Ultrafast folding of WW domains without structured aromatic clusters in the denatured state. Proc. Natl. Acad. Sci. USA 98, 13002–13007 (2001).
Ferguson, N. et al. Using flexible loop mimetics to extend φ-value analysis to secondary structure interactions. Proc. Natl. Acad. Sci. USA 98, 13008–13013 (2001).
Socolich, M. et al. Evolutionary information for specifying a protein fold. Nature 437, 512–518 (2005).
Jäger, M., Nguyen, H., Crane, J.C., Kelly, J.W. & Gruebele, M. The folding mechanism of a β-sheet: the WW domain. J. Mol. Biol. 311, 373–393 (2001).
Jäger, M. et al. Structure-function-folding relationship in a WW domain. Proc. Natl. Acad. Sci. USA 103, 10648–10653 (2006).
Deechongkit, S. et al. β-sheet folding mechanisms from perturbation energetics. Curr. Opin. Struct. Biol. 16, 94–101 (2006).
Nguyen, H., Jager, M., Kelly, J.W. & Gruebele, M. Engineering a β-sheet protein toward the folding speed limit. J. Phys. Chem. B 109, 15182–15186 (2005).
Ranganathan, R., Lu, K.P., Hunter, T. & Noel, J.P. Structural and functional analysis of the mitotic rotamase Pin1 suggests substrate recognition is phosphorylation dependent. Cell 89, 875–886 (1997).
Bai, Y., Karimi, A., Dyson, H.J. & Wright, P.E. Absence of a stable intermediate on the folding pathway of protein A. Protein Sci. 6, 1449–1457 (1997).
Sato, S., Religa, T.L., Daggett, V. & Fersht, A.R. Testing protein-folding simulations by experiment: B domain of protein A. Proc. Natl. Acad. Sci. USA 101, 6952–6956 (2004).
Fersht, A.R., Matouschek, A. & Serrano, L. The folding of an enzyme. I. Theory of protein engineering analysis of stability and pathway of protein folding. J. Mol. Biol. 224, 771–782 (1992).
Carter, P.J., Winter, G., Wilkinson, A.J. & Fersht, A.R. The use of double mutants to detect structural changes in the active site of the tyrosyl-tRNA synthetase (Bacillus stearothermophilus). Cell 38, 835–840 (1984).
Horovitz, A. & Fersht, A.R. Co-operative interactions during protein folding. J. Mol. Biol. 224, 733–740 (1992).
Seebach, D., Beck, A.K. & Bierbaum, D.J. The world of β- and γ-peptides comprised of homologated proteinogenic amino acids and other components. Chem. Biodivers. 1, 1111–1239 (2004).
Horne, W.S., Price, J.L. & Gellman, S.H. Interplay among side chain sequence, backbone composition, and residue rigidification in polypeptide folding and assembly. Proc. Natl. Acad. Sci. USA 105, 9151–9156 (2008).
Price, J.L., Horne, W.S. & Gellman, S.H. Discrete heterogeneous quaternary structure formed by α/β-peptide foldamers and α-peptides. J. Am. Chem. Soc. 129, 6376–6377 (2007).
Chatterjee, S., Roy, R.S. & Balaram, P. Expanding the polypeptide backbone: hydrogen-bonded conformations in hybrid polypeptides containing the higher homologues of α-amino acids. J. R. Soc. Interface 4, 587–606 (2007).
Hann, M.M., Sammes, P.G., Kennewell, P.D. & Taylor, J.B. On double bond isosteres of the peptide bond; an enkephalin analogue. J. Chem. Soc. Chem. Commun. 5, 234–235 (1980).
Xiao, J., Weisblum, B. & Wipf, P. Trisubstituted (E)-alkene dipeptide isosteres as β-turn promoters in the gramicidin S cyclodecapeptide scaffold. Org. Lett. 8, 4731–4734 (2006).
Jenkins, C.L., Vasbinder, M.M., Miller, S.J. & Raines, R.T. Peptide bond isosteres: ester or E-alkene in the backbone of the collagen triple helix. Org. Lett. 7, 2619–2622 (2005).
Fu, Y., Gao, J.M., Bieschke, J., Dendle, M.A. & Kelly, J.W. Amide-to-E-olefin versus amide-to-ester backbone H-bond perturbations: Evaluating the O-O repulsion for extracting H-bond energies. J. Am. Chem. Soc. 128, 15948–15949 (2006).
Gao, J. & Kelly, J.W. Toward quantification of protein backbone-backbone hydrogen bonding energies: An energetic analysis of an amide-to-ester mutation in an α-helix within a protein. Protein Sci. 17, 1096–1101 (2008).
Stigers, K.D., Soth, M.J. & Nowick, J.S. Designed molecules that fold to mimic protein secondary structures. Curr. Opin. Chem. Biol. 3, 714–723 (1999).
Yang, X., Wang, M. & Fitzgerald, M.C. Analysis of protein folding and function using backbone modified proteins. Bioorg. Chem. 32, 438–449 (2004).
Myers, J.K. & Oas, T.G. Preorganized secondary structure as an important determinant of fast protein folding. Nat. Struct. Biol. 8, 552–558 (2001).
Dawson, P.E. & Kent, S.B. Synthesis of native proteins by chemical ligation. Annu. Rev. Biochem. 69, 923–960 (2000).
Bai, Y., Milne, J.S., Mayne, L. & Englander, S.W. Primary structure effects on peptide group hydrogen exchange. Proteins 17, 75–86 (1993).
Dyson, H.J., Rance, M., Houghten, R.A., Lerner, R.A. & Wright, P.E. Folding of immunogenic peptide fragments of proteins in water solution. I. Sequence requirements for the formation of a reverse turn. J. Mol. Biol. 201, 161–200 (1988).
Wright, P.E., Dyson, H.J. & Lerner, R.A. Conformation of peptide fragments of proteins in aqueous solution: implications for initiation of protein folding. Biochemistry 27, 7167–7175 (1988).
Yi, Q., Scalley-Kim, M.L., Alm, E.J. & Baker, D. NMR characterization of residual structure in the denatured state of protein L. J. Mol. Biol. 299, 1341–1351 (2000).
Zhang, O. & Forman-Kay, J.D. NMR studies of unfolded states of an SH3 domain in aqueous solution and denaturing conditions. Biochemistry 36, 3959–3970 (1997).
Anil, B., Li, Y., Cho, J.H. & Raleigh, D.P. The unfolded state of NTL9 is compact in the absence of denaturant. Biochemistry 45, 10110–10116 (2006).
Blaber, M. et al. Energetic cost and structural consequences of burying a hydroxyl group within the core of a protein determined from Ala → Ser and Val → Thr substitutions in T4 lysozyme. Biochemistry 32, 11363–11373 (1993).
Alber, T. et al. Contributions of hydrogen bonds of Thr 157 to the thermodynamic stability of phage T4 lysozyme. Nature 330, 41–46 (1987).
McDonald, I.K. & Thornton, J.M. Satisfying hydrogen bonding potential in proteins. J. Mol. Biol. 238, 777–793 (1994).
Fleming, P.J. & Rose, G.D. Do all backbone polar groups in proteins form hydrogen bonds? Protein Sci. 14, 1911–1917 (2005).
Bai, Y. & Englander, S.W. Hydrogen bond strength and β-sheet propensities—the role of a side chain blocking effect. Proteins 18, 262–266 (1994).
Franzen, J.S. & Stephens, R.E. Effect of a dipolar solvent system on interamide hydrogen bonds. Biochemistry 2, 1321–1327 (1963).
Pace, C.N. The stability of globular proteins. CRC Crit. Rev. Biochem. 3, 1–43 (1975).
Acknowledgements
This work was supported by a grant from the US National Institutes of Health (GM051105), the Lita Annenberg Hazen Foundation and The Skaggs Institute for Chemical Biology. D.A.B. thanks the National Institutes of Health for a postdoctoral fellowship (NS047024).
Author information
Authors and Affiliations
Contributions
J.W.K., E.T.P. and J.G. designed the project, analyzed the data and wrote the manuscript; J.G. performed the protein folding studies; D.A.B. and J.G. obtained and analyzed the NMR data.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1 and 2 (PDF 395 kb)
Rights and permissions
About this article
Cite this article
Gao, J., Bosco, D., Powers, E. et al. Localized thermodynamic coupling between hydrogen bonding and microenvironment polarity substantially stabilizes proteins. Nat Struct Mol Biol 16, 684–690 (2009). https://doi.org/10.1038/nsmb.1610
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1610
This article is cited by
-
Inhibitor design for TMPRSS2: insights from computational analysis of its backbone hydrogen bonds using a simple descriptor
European Biophysics Journal (2024)
-
Switchable bifunctional molecular recognition in water using a pH-responsive Endo-functionalized cavity
Nature Communications (2022)
-
Metabolomics analysis of milk thistle lipids to identify drought-tolerant genes
Scientific Reports (2022)
-
Hydrogen-bonded networks in alcohol-acetone binary mixtures: molecular dynamics study
Journal of Molecular Modeling (2022)
-
Stereoelectronic effects in stabilizing protein–N-glycan interactions revealed by experiment and machine learning
Nature Chemistry (2021)