Abstract
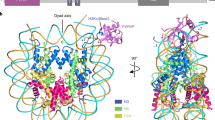
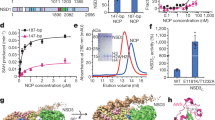
The WD40-repeat protein WDR5 is a conserved subunit of Trithorax (TRX) histone methyltransferase complexes. WDR5 has been reported to selectively bind dimethylated Lys4 (K4me2) in histone H3 to promote K4 trimethylation by TRX. To elucidate the basis of this binding specificity, we have determined the crystal structure of WDR5 bound to a histone H3 peptide bearing K4me2. The structure reveals that the N terminus of histone H3 binds as a 310-helix in the central depression formed by the WD40 repeats. R2 in histone H3 is bound in the acidic channel in the protein's core, whereas K4me2 is solvent exposed and does not engage in direct interactions with WDR5. Functional studies confirm that WDR5 recognizes A1, R2 and T3 in histone H3 but has virtually identical affinities for the unmodified and mono-, di- and trimethylated forms of K4, demonstrating that it does not discriminate among different degrees of methylation of this residue.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Luger, K. Structure and dynamic behavior of nucleosomes. Curr. Opin. Genet. Dev. 13, 127–135 (2003).
Woodcock, C.L. Chromatin architecture. Curr. Opin. Struct. Biol. 16, 213–220 (2006).
Iizuka, M. & Smith, M.M. Functional consequences of histone modifications. Curr. Opin. Genet. Dev. 13, 154–160 (2003).
Fischle, W., Wang, Y. & Allis, C.D. Histone and chromatin cross-talk. Curr. Opin. Cell Biol. 15, 172–183 (2003).
Zeng, L. & Zhou, M.M. Bromodomain: an acetyl-lysine binding domain. FEBS Lett. 513, 124–128 (2002).
Brehm, A., Tufteland, K.R., Aasland, R. & Becker, P.B. The many colours of chromodomains. Bioessays 26, 133–140 (2004).
Sims, R.J., III et al. Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J. Biol. Chem. 280, 41789–41792 (2005).
Huang, Y., Fang, J., Bedford, M.T., Zhang, Y. & Xu, R.M. Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A. Science 312, 748–751 (2006).
Wysocka, J. et al. WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121, 859–872 (2005).
Gori, F., Divieti, P. & Demay, M.B. Cloning and characterization of a novel WD-40 repeat protein that dramatically accelerates osteoblastic differentiation. J. Biol. Chem. 276, 46515–46522 (2001).
Gori, F. & Demay, M.B. BIG-3, a novel WD-40 repeat protein, is expressed in the developing growth plate and accelerates chondrocyte differentiation in vitro. Endocrinology 145, 1050–1054 (2004).
Gori, F. & Demay, M.B. The effects of BIG-3 on osteoblast differentiation are not dependent upon endogenously produced BMPs. Exp. Cell Res. 304, 287–292 (2005).
Gori, F., Friedman, L. & Demay, M.B. Wdr5, a novel WD repeat protein, regulates osteoblast and chondrocyte differentiation in vivo. J. Musculoskelet. Neuronal Interact. 5, 338–339 (2005).
Milne, T.A. et al. MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol. Cell 10, 1107–1117 (2002).
Nakamura, T. et al. ALL-1 is a histone methyltransferase that assembles a supercomplex of proteins involved in transcriptional regulation. Mol. Cell 10, 1119–1128 (2002).
Goo, Y.H. et al. Activating signal cointegrator 2 belongs to a novel steady-state complex that contains a subset of trithorax group proteins. Mol. Cell. Biol. 23, 140–149 (2003).
Wysocka, J., Myers, M.P., Laherty, C.D., Eisenman, R.N. & Herr, W. Human Sin3 deacetylase and trithorax-related Set1/Ash2 histone H3–K4 methyltransferase are tethered together selectively by the cell-proliferation factor HCF-1. Genes Dev. 17, 896–911 (2003).
Hughes, C.M. et al. Menin associates with a trithorax family histone methyltransferase complex and with the hoxc8 locus. Mol. Cell 13, 587–597 (2004).
Yokoyama, A. et al. Leukemia proto-oncoprotein MLL forms a SET1-like histone methyltransferase complex with menin to regulate Hox gene expression. Mol. Cell. Biol. 24, 5639–5649 (2004).
Milne, T.A. et al. MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol. Cell 10, 1107–1117 (2002).
Milne, T.A. et al. MLL associates specifically with a subset of transcriptionally active target genes. Proc. Natl. Acad. Sci. USA 102, 14765–14770 (2005).
Guenther, M.G. et al. Global and Hox-specific roles for the MLL1 methyltransferase. Proc. Natl. Acad. Sci. USA 102, 8603–8608 (2005).
Shannon, M.P., Kaufman, T.C., Shen, M.W. & Judd, B.H. Lethality patterns and morphology of selected lethal and semi-lethal mutations in the zeste-white region of Drosophila melanogaster. Genetics 72, 615–638 (1972).
Hollmann, M., Simmerl, E., Schafer, U. & Schafer, M.A. The essential Drosophila melanogaster gene wds (will die slowly) codes for a WD-repeat protein with seven repeats. Mol. Genet. Genomics 268, 425–433 (2002).
Nielsen, P.R. et al. Structure of the HP1 chromodomain bound to histone H3 methylated at lysine 9. Nature 416, 103–107 (2002).
Jacobs, S.A. & Khorasanizadeh, S. Structure of HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 295, 2080–2083 (2002).
Schurter, B.T. et al. Methylation of histone H3 by coactivator-associated arginine methyltransferase 1. Biochemistry 40, 5747–5756 (2001).
Wang, Y. et al. Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 306, 279–283 (2004).
Cuthbert, G.L. et al. Histone deimination antagonizes arginine methylation. Cell 118, 545–553 (2004).
Dai, J., Sultan, S., Taylor, S.S. & Higgins, J.M. The kinase haspin is required for mitotic histone H3 Thr 3 phosphorylation and normal metaphase chromosome alignment. Genes Dev. 19, 472–488 (2005).
Fischle, W., Wang, Y. & Allis, C.D. Binary switches and modification cassettes in histone biology and beyond. Nature 425, 475–479 (2003).
Lambright, D.G. et al. The 2.0 Å crystal structure of a heterotrimeric G protein. Nature 379, 311–319 (1996).
Orlicky, S., Tang, X., Willems, A., Tyers, M. & Sicheri, F. Structural basis for phosphodependent substrate selection and orientation by the SCFCdc4 ubiquitin ligase. Cell 112, 243–256 (2003).
Wu, G. et al. Structure of a beta-TrCP1-Skp1-beta-catenin complex: destruction motif binding and lysine specificity of the SCF(beta-TrCP1) ubiquitin ligase. Mol. Cell 11, 1445–1456 (2003).
Verreault, A., Kaufman, P.D., Kobayashi, R. & Stillman, B. Nucleosome assembly by a complex of CAF-1 and acetylated histones H3/H4. Cell 87, 95–104 (1996).
Parthun, M.R., Widom, J. & Gottschling, D.E. The major cytoplasmic histone acetyltransferase in yeast: links to chromatin replication and histone metabolism. Cell 87, 85–94 (1996).
Verreault, A., Kaufman, P.D., Kobayashi, R. & Stillman, B. Nucleosomal DNA regulates the core-histone-binding subunit of the human Hat1 acetyltransferase. Curr. Biol. 8, 96–108 (1998).
Zhang, Q., Vo, N. & Goodman, R.H. Histone binding protein RbAp48 interacts with a complex of CREB binding protein and phosphorylated CREB. Mol. Cell. Biol. 20, 4970–4978 (2000).
Martinez-Balbas, M.A., Tsukiyama, T., Gdula, D. & Wu, C. Drosophila NURF-55, a WD repeat protein involved in histone metabolism. Proc. Natl. Acad. Sci. USA 95, 132–137 (1998).
Zhang, Y., Iratni, R., Erdjument-Bromage, H., Tempst, P. & Reinberg, D. Histone deacetylases and SAP18, a novel polypeptide, are components of a human Sin3 complex. Cell 89, 357–364 (1997).
Xue, Y. et al. NURD, a novel complex with both ATP-dependent chromatin-remodeling and histone deacetylase activities. Mol. Cell 2, 851–861 (1998).
Zhang, Y., LeRoy, G., Seelig, H.P., Lane, W.S. & Reinberg, D. The dermatomyositis-specific autoantigen Mi2 is a component of a complex containing histone deacetylase and nucleosome remodeling activities. Cell 95, 279–289 (1998).
Zhang, Y. et al. Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation. Genes Dev. 13, 1924–1935 (1999).
Kuzmichev, A., Nishioka, K., Erdjument-Bromage, H., Tempst, P. & Reinberg, D. Histone methyltransferase activity associated with a human multiprotein complex containing the Enhancer of Zeste protein. Genes Dev. 16, 2893–2905 (2002).
Cao, R. et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298, 1039–1043 (2002).
Sewalt, R.G. et al. Characterization of interactions between the mammalian polycomb-group proteins Enx1/EZH2 and EED suggests the existence of different mammalian polycomb-group protein complexes. Mol. Cell. Biol. 18, 3586–3595 (1998).
Montgomery, N.D. et al. The murine polycomb group protein Eed is required for global histone H3 lysine-27 methylation. Curr. Biol. 15, 942–947 (2005).
Sheffield, P., Garrard, S. & Derewenda, Z. Overcoming expression and purification problems of RhoGDI using a family of “parallel” expression vectors. Protein Expr. Purif. 15, 34–39 (1999).
Messerschmidt, A. & Pflugrath, J.W. Crystal orientation and X-ray pattern prediction routines for area-detector diffractometer systems in macromolecular crystallography. J. Appl. Crystallogr. 20, 306–315 (1987).
Lambert, C., Leonard, N., De Bolle, X. & Depiereux, E. ESyPred3D: prediction of proteins 3D structures. Bioinformatics 18, 1250–1256 (2002).
McCoy, A.J., Grosse-Kunstleve, R.W., Storoni, L.C. & Read, R.J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D Biol. Crystallogr. 61, 458–464 (2005).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Fenn, T.D., Ringe, D. & Petsko, G.A. POVScript+: a program for model and data visualization using persistence of vision ray-tracing. J. Appl. Crystallogr. 36, 944–947 (2003).
Acknowledgements
We thank J. Brunzelle for assistance in X-ray data collection, D. Peisach for assistance with rendering figures, D. Peisach for reading the manuscript and providing useful comments, the ESRF for provision of synchrotron radiation facilities and L. Serre and M. Walsh for their assistance in using beamlines FIP BM30A and BM14, respectively. Use of the University of Michigan DNA Sequencing Core was supported by the US National Institutes of Health through the University of Michigan's Cancer Center Support Grant (5 P30 CA46592). J.-F.C. is a Canadian Institutes of Health Research Postdoctoral Fellow. This work was supported by the University of Michigan's Office of the Vice President for Research and US National Institutes of Health grant GM073839 to R.C.T.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
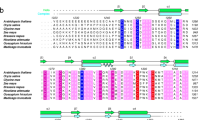
Supplementary Fig. 1
Sequence alignment of metazoan orthologs of WDR5. (PDF 245 kb)
Rights and permissions
About this article
Cite this article
Couture, JF., Collazo, E. & Trievel, R. Molecular recognition of histone H3 by the WD40 protein WDR5. Nat Struct Mol Biol 13, 698–703 (2006). https://doi.org/10.1038/nsmb1116
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1116
This article is cited by
-
Histone 3.3-related chromatinopathy: missense variants throughout H3-3A and H3-3B cause a range of functional consequences across species
Human Genetics (2023)
-
Rescue of deficits by Brwd1 copy number restoration in the Ts65Dn mouse model of Down syndrome
Nature Communications (2022)
-
Targeting non-bromodomain chromatin readers
Nature Structural & Molecular Biology (2019)
-
WDR5 supports colon cancer cells by promoting methylation of H3K4 and suppressing DNA damage
BMC Cancer (2018)
-
The ZZ-type zinc finger of ZZZ3 modulates the ATAC complex-mediated histone acetylation and gene activation
Nature Communications (2018)