Abstract
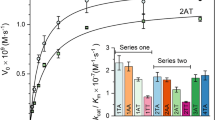
Flap endonucleases (FENs) have essential roles in DNA processing. They catalyze exonucleolytic and structure-specific endonucleolytic DNA cleavage reactions. Divalent metal ions are essential cofactors in both reactions. The crystal structure of FEN shows that the protein has two conserved metal-binding sites. Mutations in site I caused complete loss of catalytic activity. Mutation of crucial aspartates in site II abolished exonuclease action, but caused enzymes to retain structure-specific (flap endonuclease) activity. Isothermal titration calorimetry revealed that site I has a 30-fold higher affinity for cofactor than site II. Structure-specific endonuclease activity requires binding of a single metal ion in the high-affinity site, whereas exonuclease activity requires that both the high- and low-affinity sites be occupied by divalent cofactor. The data suggest that a novel two-metal mechanism operates in the FEN-catalyzed exonucleolytic reaction. These results raise the possibility that local concentrations of free cofactor could influence the endo- or exonucleolytic pathway in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Cowan, J.A. Metal activation of enzymes in nucleic acid biochemistry. Chem. Rev. 98, 1067–1088 (1998).
Lieber, M.R. The FEN1 family of structure-specific nucleases in eukaryotic DNA replication, recombination and repair. Bioessays 19, 233–240 (1997).
Ceska, T.A. & Sayers, J.R. Structure-specific DNA cleavage by 5′ nucleases. Trends Biochem. Sci. 23, 331–336 (1998).
Kucherlapati, M. et al. Haploinsufficiency of flap endonuclease (Fen1) leads to rapid tumor progression. Proc. Natl. Acad. Sci. USA 99, 9924–9929 (2002).
Harrington, J.J. & Lieber, M.R. The characterisation of a mammalian DNA structure-specific endonuclease. EMBO J. 13, 1235–1246 (1994).
Lyamichev, V., Brow, M.A.D. & Dahlberg, J.E. Structure-specific endonucleolytic cleavage of nucleic acids by eubacterial DNA polymerases. Science 260, 778–783 (1993).
Diaz, A., Pons, M.E., Lacks, S.A. & Lopez, P. Streptococcus pneumoniae DNA polymerase I lacks 3′–5′ exonuclease activity: localization of the 5′–3′ exonucleolytic domain. J. Bacteriol. 174, 2014–2024 (1992).
Bhagwat, M. & Nossal, N.G. Bacteriophage T4 RNase H removes both RNA primers and adjacent DNA from the 5′ end of lagging strand fragments. J. Biol. Chem. 276, 28516–28524 (2001).
Rumbaugh, J.A., Murante, R.S., Shi, S. & Bambara, R.A. Creation and removal of embedded ribonucleotides in chromosomal DNA during mammalian Okazaki fragment processing. J. Biol. Chem. 272, 22591–22599 (1997).
Sayers, J.R. & Artymiuk, P.J. Flexible loops and helical arches. Nat. Struct. Biol. 5, 668–670 (1998).
Kim, Y. et al. Crystal structure of Thermus aquaticus DNA polymerase. Nature 376, 612–616 (1995).
Mueser, T.C., Nossal, N.G. & Hyde, C.C. Structure of bacteriophage T4 RNase H, a 5′–3′ RNA-DNA and DNA-DNA exonuclease with sequence similarity to the RAD2 family of eukaryotic proteins. Cell 85, 1101–1112 (1996).
Ceska, T.A., Sayers, J.R., Stier, G. & Suck, D. A helical arch allowing single-stranded DNA to thread through T5 5′ exonuclease. Nature 382, 90–93 (1996).
Hwang, K.Y., Baek, K., Kim, H.Y. & Cho, Y. The crystal structure of flap endonuclease-1 from Methanococcus jannaschii . Nat. Struct. Biol. 5, 707–713 (1998).
Hosfield, D.J., Mol, C.D., Shen, B. & Tainer, J.A. Structure of the DNA repair and replication endonuclease and exonuclease FEN1: coupling DNA and PCNA binding to FEN1 activity. Cell 95, 135–146 (1998).
Matsui, E., et al. Molecular structure and novel DNA binding sites located in loops of flap endonuclease-1 from Pyrococcus horikoshii . J. Biol. Chem. 277, 37840–37847 (2002).
Beese, L.S. & Steitz, T.A. Structural basis for the 3′–5′ exonuclease activity of Escherichia coli DNA polymerase I: a two metal ion mechanism. EMBO J. 10, 25–33 (1991).
Bhagwat, M., Meara, D. & Nossal, N.G. Identification of residues of T4 RNase H required for catalysis and DNA binding J. Biol. Chem. 272, 28531–28538 (1997).
Amblar, M. & Lopez, P. Purification and properties of the 5′–3′ exonuclease D190A mutant of DNA polymerase I from Streptococcus pneumoniae . Eur. J. Biochem. 252, 124–132 (1998).
Mizrahi, V. & Huberts, P. Deoxy- and dideoxynucleotide discrimination and identification of critical 5′ nuclease domain residues of the DNA polymerase I from Mycobacterium tuberculosis . Nucleic Acids Res. 24, 4845–4852 (1996).
Xu, Y. et al. Biochemical and mutational studies of the 5′–3′ exonuclease of DNA polymerase I of Escherichia coli . J. Mol. Biol. 268, 284–302 (1997).
Xu, Y., Potapova, O., Leschziner, A.E., Grindley, N.D. & Joyce, C.M. Contacts between the 5′ nuclease of DNA polymerase I and its DNA substrate. J. Biol. Chem. 276, 30167–30177 (2001).
Shen, B., Nolan, J.P., Sklar, L.A. & Park, M.S. Functional analysis of point mutations in human flap endonuclease-1 active site. Nucleic Acids Res. 25, 3332–3338 (1997).
Amblar, M., de Lacoba, M.G., Corrales, M.A. & Lopez, P. Biochemical analysis of point mutations in the 5′–3′ exonuclease of DNA polymerase I of Streptococcus pneumoniae. Functional and structural implications. J. Biol. Chem. 276, 19172–19181 (2001).
Tock, M.R., Frary, E., Sayers, J.R. & Grasby, J.A. Dynamic evidence for metal ion catalysis in the reaction mediated by a flap endonuclease. EMBO J. 22, 995–1004 (2003).
Haq, I., Chowdhry, B.Z. & Jenkins, T.C. Calorimetric techniques in the study of high-order DNA-drug interactions. Methods Enzymol. 340, 109–149 (2001).
Wiseman, T., Williston, S., Brandts, J.F. & Lin, L.N. Rapid measurement of binding constants and heats of binding using a new titration calorimeter. Anal. Biochem. 179, 131–137 (1989).
Milos, M., Schaer, J.J., Comte, M. & Cox, J.A. Microcalorimetric investigation of the interactions in the ternary complex calmodulin-calcium-melittin. J. Biol. Chem. 262, 2746–2749 (1987).
Ogawa, Y. & Tanokura, M. Calcium binding to calmodulin: effects of ionic strength, Mg2+, pH and temperature. J. Biochem. 95, 19–28 (1984).
Jose, T.J., Conlan, L.H. & Dupureur, C.M. Quantitative evaluation of metal ion binding to PvuII restriction endonuclease. J. Biol. Inor. Chem. 4, 814–823 (1999).
Zheng, L., Li, M., Shan, J., Krishnamoorthi, R. & Shen, B. Distinct roles of two Mg2+ binding sites in regulation of murine flap endonuclease-1 activities. Biochemistry 41, 10323–10331 (2002).
Hosfield, D.J., Frank, G., Weng, Y., Tainer, J.A. & Shen, B. Newly discovered archaebacterial flap endonucleases show a structure-specific mechanism for DNA substrate binding and catalysis resembling human flap endonuclease-1. J. Biol. Chem. 273, 27154–27161 (1998).
Sayers, J.R. & Eckstein, F. Properties of overexpressed phage T5 D15 exonuclease. J. Biol. Chem. 265, 18311–18317 (1990).
Vipond, I.B. & Halford, S.E. Specific DNA recognition by EcoRV restriction endonuclease induced by calcium ions. Biochemistry (US), 34, 1113–1119 (1995).
Sayers, J.R. & Eckstein, F. A single-strand specific endonuclease activity copurifies with overexpressed T5 D15 exonuclease. Nucleic Acids Res. 19, 4127–4132 (1991).
Garforth, S.J. & Sayers, J.R. Structure-specific DNA binding by bacteriophage T5 5′–3′ exonuclease. Nucleic Acids Res. 25, 3801–3807 (1997).
Sayers, J.R. Viral polymerase-associated 5′–3′ exonucleases: expression, purification, and uses. Methods Enzymol. 275, 227–238 (1996).
Fraser, M.J. Purification and properties of Neurospora crassa endo-exonuclease, an enzyme which can be converted to a single-strand specific endonuclease. Methods Enzymol. 65, 255–263 (1980).
Garforth, S.J., Ceska, T.A., Suck, D. & Sayers, J.R. Mutagenesis of conserved lysine residues in bacteriophage T5 5′–3′ exonuclease suggests separate mechanisms of endo- and exonucleolytic cleavage. Proc. Natl. Acad. Sci. USA 96, 38–43 (1999).
Ceska, T.A., Sayers, J.R., Eckstein, F. & Suck, D. Preliminary crystallographic studies on the D15 5′–3′ exonuclease from phage T5. J. Mol. Biol. 233, 179–182 (1993).
Kabsch, W. Evaluation of single crystal X-ray diffraction data from a position sensitive detector. J. Appl. Crystallogr. 21, 916–924 (1988).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Badger, J., Kumar, R.A., Yip, P. & Szalma, S. New features and enhancements in the X-PLOR computer program. Proteins 35, 25–33 (1999).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Acknowledgements
We thank J. Grasby for informal discussions and critical reading of the manuscript. This paper is dedicated to the memory of A. Jones. The support of The Wellcome Trust (grant numbers 052123 and 058034) is gratefully acknowledged. The White Rose University Consortium and the BBSRC supported studentships for M.F. and J.J.D., respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Feng, M., Patel, D., Dervan, J. et al. Roles of divalent metal ions in flap endonuclease–substrate interactions. Nat Struct Mol Biol 11, 450–456 (2004). https://doi.org/10.1038/nsmb754
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb754
This article is cited by
-
Direct observation of DNA threading in flap endonuclease complexes
Nature Structural & Molecular Biology (2016)
-
A novel chemoreceptor MCP2983 from Comamonas testosteroni specifically binds to cis-aconitate and triggers chemotaxis towards diverse organic compounds
Applied Microbiology and Biotechnology (2015)
-
Structural and biochemical studies of the 5′→3′ exoribonuclease Xrn1
Nature Structural & Molecular Biology (2011)
-
Metal ion and DNA binding by single-chain PvuII endonuclease: lessons from the linker
JBIC Journal of Biological Inorganic Chemistry (2011)
-
Zc3h12a is an RNase essential for controlling immune responses by regulating mRNA decay
Nature (2009)