Abstract
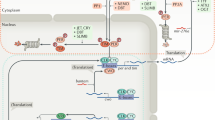
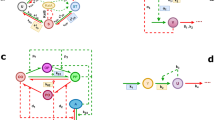
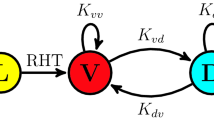
The spectacularly successful race over the past three years to place our understanding of the circadian clockwork of mammals into a molecular framework is beginning to yield the cardinal example of the molecular-genetic control of behaviour. This perspective describes recent evidence for the conservation of a double-loop, autoregulatory feedback mechanism across the best understood eukaryotic circadian systems, and discusses how these findings may illuminate some long-standing puzzles concerning our subliminal sense of circadian time.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Weaver, D. R. The suprachiasmatic nucleus: a 25-year retrospective. J. Biol. Rhythms 2, 100–112 ( 1998).
Welsh, D. K., Logothetis, D. E., Meister, M. & Reppert, S. M. Individual neurons dissociated from rat suprachiasmatic nucleus express independently phased circadian firing rhythms. Neuron 4, 697–706 (1995).
Dunlap, J. C. Molecular bases for circadian clocks. Cell 2, 271–290 (1999).
Taylor, B. L. & Zhulin, I. B. PAS domains: internal sensors of oxygen, redox potential, and light. Microbiol. Mol. Biol. Rev. 63, 479–506 ( 1999).
Lee C., Bae, K. & Edery, I. PER and TIM inhibit the DNA binding activity of a Drosophila CLOCK-CYC/dBMAL1 heterodimer without disrupting formation of the heterodimer: a basis for circadian transcription. Mol. Cell. Biol. 19, 5316 –5325 (1999).
Price, J. L. et al. double-time is a novel Drosophila clock gene that regulates PERIOD protein accumulation. Cell 94 , 83–96 (1998).
Lowrey, P. L. et al. Positional syntenic cloning and functional characterisation of the mammalian circadian mutation, tau. Science 288 , 483–491 (2000).
Glossop, N. R., Lyons, L. C. & Hardin, P. E. Interlocked feedback loops within the Drosophila circadian oscillator. Science 286, 766–768 (1999).
Shearman L. P. et al. Interacting molecular loops in the mammalian circadian clock . Science 288, 1013–1019 (2000).
Lee, K., Loros, J. J. & Dunlap, J. C. Interconnected feedback loops in the Neurospora circadian system. Science 289, 107– 110 (2000).
Bae, K., Lee, C., Hardin, P. E. & Edery, I. dCLOCK is present in limiting amounts and likely mediates daily interactions between the dCLOCK-CYC transcription factor and the PER–TIM complex. J. Neurosci. 5, 1746–1753 ( 2000).
Gotter, A. L. et al. A time-less function for mouse timeless. Nature Neurosci. 8, 755–756 ( 2000).
Kume, K. et al. mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell 98, 193–205 (1999).
Emery, P. et al. Drosophila CRY is a deep brain circadian photoreceptor . Neuron 26, 493–504 (2000).
Field, M. D. et al. Analysis of clock proteins in mouse SCN demonstrates phylogenetic divergence of the circadian clockwork and resetting mechanisms. Neuron 25, 437–447 ( 2000).
Monaco, L. M. et al. CREM: a master-switch in the transcriptional response to cAMP . Phil. Trans. R. Soc. Lond. B 351, 561– 567 (1996).
Jin, X. et al. A molecular mechanism regulating rhythmic output from the suprachiasmatic circadian clock. Cell 96, 57– 68 (1999).
Ripperberger, J. A., Sheraman, L. P., Repert, S. M. & Schibler, U. CLOCK, an essential pacemaker component, controls expression of the circadian transcription factor DBP. Genes Dev. 14, 679–689 (2000).
Yamaguchi, S. & Okamura, H. Role of DBP in the circadian oscillatory mechanism. Mol. Cell. Biol. 13, 4773– 4781 (2000).
Bell-Pedersen, D., Shinohara, M. L., Loros, J. J. & Dunlap, J. C. Circadian clock-controlled genes isolated from Neurospora crassa are late night- to early morning-specific. Proc. Natl Acad. Sci. USA 23, 13096–13101 ( 1996).
Renn, S. C., Park, J. H., Rosbash, M., Hall, J. C. & Taghert, P. H. A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila. Cell 7, 791– 802 (1999).
Blau, J. & Young, M. W. Cycling vrille expression is required for a functional Drosophila clock. Cell 6, 661–671 (1999).
Park, J. H. et al. Differential regulation of circadian pacemaker output by separate clock genes in Drosophila. Proc. Natl Acad. Sci. USA 97, 3608–3613 (2000).
Ueyama, T. et al. Suprachiasmatic nucleus: a central autonomic clock. Nature Neurosci. 2, 1051–1053 (1999).
Smolensky, M. H. & Portaluppi, F. Chronopharmacology and chronotherapy of cardiovascular medications: relevance to prevention and treatment of coronary heart disease. Am. Heart J.. 137, S14–S24 (1999).
Plautz, J. D., Kaneko, M., Hall, J. C. & Kay, S. A. Independent photoreceptive circadian clocks throughout Drosophila. Science 278, 1632–1635 (1997).
Whitmore, D., Foulkes, N. S. & Sassone-Corsi, P. Light acts directly on organs and cells in culture to set the vertebrate circadian clock. Nature 404, 87–91 (2000).
Balsalabre, A., Damiola, F. & Schibler, U. A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 93, 929– 937 (1998).
Yagita, K. & Okamura, H. Forskolin induces circadian gene expression of rPer1, rPer2 and dbp in mammalian rat-1 fibroblasts. FEBS Lett. 465, 79– 82 (2000).
Yamazaki, S. et al. Resetting central and peripheral circadian oscillators in transgenic rats. Science 288, 682– 685 (2000).
Yamaguchi, S. et al. The 5′ upstream region of mPer1 gene contains two promoters and is responsible for circadian oscillation. Curr. Biol. 14, 873–876 ( 2000).
Balsalobre, A. et al. Resetting of circadian time in peripheral tissues by glucocorticoid signalling. Science 289, 2344– 2347 (2000).
Author information
Authors and Affiliations
Supplementary information
Rights and permissions
About this article
Cite this article
Hastings, M. Circadian clockwork: two loops are better than one. Nat Rev Neurosci 1, 143–146 (2000). https://doi.org/10.1038/35039080
Issue Date:
DOI: https://doi.org/10.1038/35039080
This article is cited by
-
Diurnal expression of circadian clock genes period 1 and period 3 in Pelteobagrus vachellii
Journal of Oceanology and Limnology (2021)
-
Parameter estimation in models of biological oscillators: an automated regularised estimation approach
BMC Bioinformatics (2019)
-
No evidence for an association between Clock gene allelic variation and migration timing in a long-distance migratory shorebird (Limosa lapponica baueri)
Oecologia (2019)
-
VIP receptors control excitability of suprachiasmatic nuclei neurones
Pflügers Archiv - European Journal of Physiology (2006)
-
Role for antisense RNA in regulating circadian clock function in Neurospora crassa
Nature (2003)