Key Points
-
Until recently, the maturation of the 3′-end of tRNA precursors in Bacteria has been assumed to follow the E. coli paradigm, in which a cleavage event downstream of tRNAs is followed by exonucleolytic removal of the residual nucleotides. However, the discovery of RNase Z in B. subtilis demonstrated the existence of a single-step endonucleolytic 3′-end maturation pathway in Bacteria, an activity previously thought to be confined to eukaryotic cells.
-
RNase Z generally hydrolyses the phosphodiester bond immediately downstream of the discriminator nucleotide. This results in a tRNA with a 3′-hydroxyl group and a trailer fragment bearing a 5′-phosphate residue. The activity of RNase Z is strongly inhibited by the presence of cytidines downstream of the discriminator. Therefore, the CCA motif present at the 3′-end of all mature tRNAs is an anti-determinant for RNase Z activity.
-
RNase Z is widely distributed in all prokaryotic families except the Proteobacteria. All Archaea and Eukaryotes sequenced so far contain an RNase Z paralogue. Eukaryotic RNase Z exists in two forms: a short form, and a long form that is thought to have evolved from the short form by gene duplication. Only the short form is present in Bacteria.
-
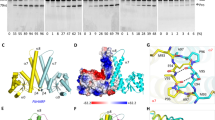
The structures of RNase Z from B. subtilis, T. maritima and E. coli have been solved at high resolution. RNase Z functions as a dimer of Zn-containing metallo-β-lactamase domains. The characteristic feature of RNase Z is a protruding flexible arm that has a role in tRNA binding. The solved structures demonstrate varied conformations of the catalytic site, from the apo-enzyme to a fully loaded active site with two zinc atoms. These variations, together with an observation that RNase Z has sigmoidal saturation kinetics, indicate that RNase Z is a member of those metallo-β-lactamases that are activated by the presence of substrate. The structure of RNase Z in complex with tRNA demonstrates that tRNA is primarily recognized by one protein subunit and cleaved by the other.
-
Despite rapid progress in the past few years on our knowledge of RNase Z function in precursor tRNA processing, we still have only a fragmentary understanding of its role in organisms such as E. coli, in which RNase Z is dispensable and does not have a significant role in tRNA maturation. Understanding the regulation of RNase Z at the level of its expression or its enzyme activity is also a challenging task for the future, as is understanding the molecular basis for inhibition by the CCA motif.
Abstract
RNase Z is a widely distributed and often essential endoribonuclease that is responsible for the maturation of the 3′-end of a large family of transfer RNAs (tRNAs). Although it has been the subject of study for more than 25 years, interest in this enzyme intensified dramatically with the identification of the encoding gene in 2002. This led to the discovery of RNase Z in bacteria, in which the final step in the generation of the mature 3′-end of tRNAs had previously been assumed to be catalysed by exoribonucleases. It also led inevitably to structural studies, and the recent resolution of the structure of RNase Z in complex with tRNA has provided a detailed understanding of the molecular mechanisms of RNase Z substrate recognition and cleavage. The identification of the RNase Z gene also allowed the search for alternative substrates for this enzyme to begin in earnest. In this Review, we outline the important recent developments that have contributed to our understanding of this enzyme, particularly in prokaryotes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kazantsev, A. V. & Pace, N. R. Bacterial RNase P: a new view of an ancient enzyme. Nature Rev. Microbiol. 4, 729–740 (2006).
Schedl, P., Roberts, J. & Primakoff, P. In vitro processing of E. coli tRNA precursors. Cell 8, 581–594 (1976).
Li, Z. & Deutscher, M. P. RNase E plays an essential role in the maturation of Escherichia coli tRNA precursors. RNA 8, 97–109 (2002).
Ow, M. C. & Kushner, S. R. Initiation of tRNA maturation by RNase E is essential for cell viability in E. coli. Genes Dev. 16, 1102–1115 (2002).
Deutscher, M. P. tRNA: Structure, Biosynthesis and Function (eds Soll, D. & Rajbhandary, U. L.) (American Society for Microbiology, Washington, DC, 1995).
Li, Z. & Deutscher, M. P. Maturation pathways for E. coli tRNA precursors: a random multienzyme process in vivo. Cell 86, 503–512 (1996).
Garber, R. L. & Gage, L. P. Transcription of a cloned Bombyx mori tRNA2Ala gene: nucleotide sequence of the tRNA precursor and its processing in vitro. Cell 18, 817–828 (1979).
Hagenbuchle, O., Larson, D., Hall, G. I. & Sprague, K. U. The primary transcription product of a silkworm alanine tRNA gene: identification of in vitro sites of initiation, termination and processing. Cell 18, 1217–1229 (1979).
Castano, J. G., Tobian, J. A. & Zasloff, M. Purification and characterization of an endonuclease from Xenopus laevis ovaries which accurately processes the 3′ terminus of human pre-tRNA-Met(i) (3′-pre-tRNase). J. Biol. Chem. 260, 9002–9008 (1985).
Frendewey, D., Dingermann, T., Cooley, L. & Soll, D. Processing of precursor tRNAs in Drosophila. Processing of the 3′-end involves an endonucleolytic cleavage and occurs after 5′-end maturation. J. Biol. Chem. 260, 449–454 (1985).
Stange, N. & Beier, H. A cell-free plant extract for accurate pre-tRNA processing, splicing and modification. EMBO J. 6, 2811–2818 (1987).
Oommen, A., Li, X. Q. & Gegenheimer, P. Cleavage specificity of chloroplast and nuclear tRNA 3′-processing nucleases. Mol. Cell. Biol. 12, 865–875 (1992).
Franklin, S. E., Zwick, M. G. & Johnson, J. D. Characterization and partial purification of two pre-tRNA 5′-processing activities from Daucus carrota (carrot) suspension cells. Plant J. 7, 553–563 (1995).
Han, S. J. & Kang, H. S. Purification and characterization of the precursor tRNA 3′-end processing nuclease from Aspergillus nidulans. Biochem. Biophys. Res. Commun. 233, 354–358 (1997).
Nashimoto, M. Distribution of both lengths and 5′-terminal nucleotides of mammalian pre-tRNA 3′-trailers reflects properties of 3′-processing endoribonuclease. Nucleic Acids Res. 25, 1148–1154 (1997).
Mayer, M., Schiffer, S. & Marchfelder, A. tRNA 3′-processing in plants: nuclear and mitochondrial activities differ. Biochemistry 39, 2096–2105 (2000).
Deutscher, M. P. & Li, Z. Exoribonucleases and their multiple roles in RNA metabolism. Prog. Nucleic Acid Res. Mol. Biol. 66, 67–105 (2001).
Schiffer, S., Rosch, S. & Marchfelder, A. Assigning a function to a conserved group of proteins: the tRNA 3′-processing enzymes. EMBO J. 21, 2769–2777 (2002). First identification of the RNase Z gene in A. thaliana and M. janaschii showing that RNase Z is a ubiquitous enzyme present in all domains of life.
Wen, T., Oussenko, I. A., Pellegrini, O., Bechhofer, D. H. & Condon, C. Ribonuclease PH plays a major role in the exonucleolytic maturation of CCA-containing tRNA precursors in Bacillus subtilis. Nucleic Acids Res. 33, 3636–3643 (2005).
Engelke, D. R., Gegenheimer, P. & Abelson, J. Nucleolytic processing of a tRNA Arg-tRNA Asp dimeric precursor by a homologous component from Saccharomyces cerevisiae. J. Biol. Chem. 260, 1271–1279 (1985).
Papadimitriou, A. & Gross, H. J. Pre-tRNA 3′-processing in Saccharomyces cerevisiae. Purification and characterization of exo- and endoribonucleases. Eur. J. Biochem. 242, 747–759 (1996).
Garber, R. L. & Altman, S. In vitro processing of B. mori transfer RNA precursor molecules. Cell 17, 389–397 (1979).
Mohan, A., Whyte, S., Wang, X., Nashimoto, M. & Levinger, L. The 3′-end CCA of mature tRNA is an antideterminant for eukaryotic 3′-tRNase. RNA 5, 245–256 (1999).
Kunzmann, A., Brennicke, A. & Marchfelder, A. 5′-end maturation and RNA editing have to precede tRNA 3′-processing in plant mitochondria. Proc. Natl Acad. Sci. USA 95, 108–113 (1998).
Nashimoto, M., Tamura, M. & Kaspar, R. L. Minimum requirements for substrates of mammalian tRNA 3′-processing endoribonuclease. Biochemistry 38, 12089–12096 (1999).
Vogel, A., Schilling, O., Spath, B. & Marchfelder, A. The tRNase Z family of proteins: physiological functions, substrate specificity and structural properties. Biol. Chem. 386, 1253–1264 (2005). A recent comprehensive review on RNase Z in both eukaryotes and prokaryotes.
Pellegrini, O., Nezzar, J., Marchfelder, A., Putzer, H. & Condon, C. Endonucleolytic processing of CCA-less tRNA precursors by RNase Z in Bacillus subtilis. EMBO J. 22, 4534–4543 (2003). First identification of RNase Z activity in bacteria. Shows the inhibition of RNase Z by the CCA-motif in vivo and in vitro.
Levinger, L. et al. Matrices of paired substitutions show the effects of tRNA D/T loop sequence on Drosophila RNase P and 3′-tRNase processing. J. Biol. Chem. 273, 1015–1025 (1998).
Hardt, W. D., Schlegl, J., Erdmann, V. A. & Hartmann, R. K. Role of the D arm and the anticodon arm in tRNA recognition by eubacterial and eukaryotic RNase P enzymes. Biochemistry 32, 13046–13053 (1993).
Schiffer, S., Helm, M., Theobald-Dietrich, A., Giege, R. & Marchfelder, A. The plant tRNA 3′-processing enzyme has a broad substrate spectrum. Biochemistry 40, 8264–8272 (2001).
Minagawa, A., Takaku, H., Takagi, M. & Nashimoto, M. A novel endonucleolytic mechanism to generate the CCA 3′ termini of tRNA molecules in Thermotoga maritima. J. Biol. Chem. 279, 15688–15697 (2004).
Ezraty, B., Dahlgren, B. & Deutscher, M. P. The RNase Z homologue encoded by Escherichia coli elaC gene is RNase BN. J. Biol. Chem. 280, 16542–16545 (2005). Shows the reassignment of RNase Z function to the previously characterized RNase BN.
Schilling, O. et al. Exosite modules guide substrate recognition in the ZiPD/ElaC protein family. J. Biol. Chem. 280, 17857–17862 (2005).
Ceballos-Chavez, M. & Vioque, A. Sequence-dependent cleavage site selection by RNase Z from the cyanobacterium Synechocystis sp. PCC 6803. J. Biol. Chem. 280, 33461–33469 (2005).
Tavtigian, S. V. et al. A candidate prostate cancer susceptibility gene at chromosome 17p. Nature Genet. 27, 172–180 (2001). Shows the potential clinical significance of RNase Z (ElaC) in prostate cancer susceptibility.
Vogel, A., Schilling, O., Niecke, M., Bettmer, J. & Meyer-Klaucke, W. ElaC encodes a novel binuclear zinc phosphodiesterase. J. Biol. Chem. 277, 29078–29085 (2002).
Schilling, O. et al. Characterization of an Escherichia coli elaC deletion mutant. Biochem. Biophys. Res. Commun. 320, 1365–1373 (2004).
Asha, P. K., Blouin, R. T., Zaniewski, R. & Deutscher, M. P. Ribonuclease BN: identification and partial characterization of a new tRNA processing enzyme. Proc. Natl Acad. Sci. USA 80, 3301–3304 (1983).
Seidman, J. G., Schmidt, F. J., Foss, K. & McClain, W. H. A mutant of Escherichia coli defective in removing 3′-terminal nucleotides from some transfer RNA precursor molecules. Cell 5, 389–400 (1975).
Zaniewski, R., Petkaitis, E. & Deutscher, M. P. A multiple mutant of Escherichia coli lacking the exoribonucleases RNase II, RNase D, and RNase BN. J. Biol. Chem. 259, 11651–11653 (1984).
Kelly, K. O. & Deutscher, M. P. The presence of only one of five exoribonucleases is sufficient to support the growth of Escherichia coli. J. Bacteriol. 174, 6682–6684 (1992).
Reuven, N. B. & Deutscher, M. P. Multiple exoribonucleases are required for the 3′ processing of Escherichia coli tRNA precursors in vivo. FASEB J. 7, 143–148 (1993).
Perwez, T. & Kushner, S. R. RNase Z in Escherichia coli plays a significant role in mRNA decay. Mol. Microbiol. 60, 723–737 (2006).
Shibata, H. S., Minagawa, A., Takaku, H., Takagi, M. & Nashimoto, M. Unstructured RNA is a substrate for tRNase Z. Biochemistry 45, 5486–5492 (2006).
Schierling, K., Rosch, S., Rupprecht, R., Schiffer, S. & Marchfelder, A. tRNA 3′ end maturation in archaea has eukaryotic features: the RNase Z from Haloferax volcanii. J. Mol. Biol. 316, 895–902 (2002).
Takaku, H., Minagawa, A., Takagi, M. & Nashimoto, M. A candidate prostate cancer susceptibility gene encodes tRNA 3′-processing endoribonuclease. Nucleic Acids Res. 31, 2272–2278 (2003).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
de la Sierra-Gallay, I. L., Pellegrini, O. & Condon, C. Structural basis for substrate binding, cleavage and allostery in the tRNA maturase RNase Z. Nature 433, 657–661 (2005). Shows the first crystal structure of RNase Z at 2.1 Å resolution.
Ishii, R. et al. Crystal structure of the tRNA 3′-processing endoribonuclease tRNase Z from Thermotoga maritima. J. Biol. Chem. 280, 14138–14144 (2005).
Kostelecky, B., Pohl, E., Vogel, A., Schilling, O. & Meyer-Klaucke, W. The crystal structure of the zinc phosphodiesterase from Escherichia coli provides insight into function and cooperativity of tRNase Z-family proteins. J. Bacteriol. 188, 1607–1614 (2006).
Li de la Sierra-Gallay, I., Mathy, N., Pellegrini, O. & Condon, C. Structure of the ubiquitous 3′ processing enzyme RNase Z bound to transfer RNA. Nature Struct. Mol. Biol. 13, 376–377 (2006). Describes the crystal structure of RNase Z in a complex with tRNA at 2.9 Å resolution.
Cameron, A. D., Ridderstrom, M., Olin, B. & Mannervik, B. Crystal structure of human glyoxalase II and its complex with a glutathione thiolester substrate analogue. Structure 7, 1067–1078 (1999).
Minagawa, A., Takaku, H., Takagi, M. & Nashimoto, M. The missense mutations in the candidate prostate cancer gene ELAC2 do not alter enzymatic properties of its product. Cancer Lett. 222, 211–215 (2005).
Ricard, J., Meunier, J. C. & Buc, J. Regulatory behavior of monomeric enzymes. 1. The mnemonical enzyme concept. Eur. J. Biochem. 49, 195–208 (1974).
Kamata, K., Mitsuya, M., Nishimura, T., Eiki, J. & Nagata, Y. Structural basis for allosteric regulation of the monomeric allosteric enzyme human glucokinase. Structure 12, 429–438 (2004).
Wommer, S. et al. Substrate-activated zinc binding of metallo-β-lactamases: physiological importance of mononuclear enzymes. J. Biol. Chem. 277, 24142–24147 (2002). Suggests substrate-mediated activation of Zn binding by metallo-β-lactamases.
Zhu, L. & Deutscher, M. P. tRNA nucleotidyltransferase is not essential for Escherichia coli viability. EMBO J. 6, 2473–2477 (1987).
Weiner, A. M. tRNA maturation: RNA polymerization without a nucleic acid template. Curr. Biol. 14, R883–R885 (2004).
Sabath, L. D. & Abraham, E. P. Zinc as a cofactor for cephalosporinase from Bacillus cereus 569. Biochem. J. 98, 11C–13C (1966).
Daiyasu, H., Osaka, K., Ishino, Y. & Toh, H. Expansion of the zinc metallo-hydrolase family of the β-lactamase fold. FEBS Lett. 503, 1–6 (2001).
Heinz, U. & Adolph, H. W. Metallo-β-lactamases: two binding sites for one catalytic metal ion? Cell. Mol. Life Sci. 61, 2827–2839 (2004).
Acknowledgements
We thank O. Pellegrini and N. Mathy for their contributions to our work in this domain. This work was supported by funds from the Association pour la Recherche sur le Cancer, the Centre National de Recherche Scientifique (CNRS), Université Paris VII-Denis Diderot, ACI Jeunes Chercheurs from the Ministère de l'Education Nationale, the Agence Nationale de la Recherche (ANR) and the Conseil Régionale de l'Ile de France.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez Genome Project
FURTHER INFORMATION
Glossary
- Discriminator nucleotide
-
The single unpaired nucleotide at the 3′-end of the acceptor stem.
- T arm
-
The tRNA T arm is a 5-base-pair stem that is the recognition site for the ribosome, allowing the formation of the tRNA–ribosome complex.
- Acceptor stem
-
The tRNA acceptor stem is a 7-base-pair stem that includes the 5′-terminal nucleotide, with a 4-nucleotide single-stranded extension containing the 3′-terminal CCA motif. The OH group of the 3′-terminal adenosine serves as the amino-acid attachment site.
- D arm
-
The tRNA D arm is a 4-base-pair stem that often contains the modified base dihydrouridine.
- Anticodon loop
-
The tRNA anticodon loop is a 5-base-pair stem ending in a loop containing the anticodon triplet.
- Sigmoidal saturation kinetics
-
A kinetic model indicative of a change in enzyme conformation to optimize catalytic activity as substrate concentration increases.
- Michaelis–Menten kinetics
-
A kinetic model describing the relationship between enzyme and substrate concentration that is valid when the enzyme concentration is the limiting factor. Used to define the maximum velocity (Vmax) of the enzyme and the Km, the substrate concentration at which the rate of reaction is half its maximum.
Rights and permissions
About this article
Cite this article
Redko, Y., Li de la Sierra-Gallay, I. & Condon, C. When all's zed and done: the structure and function of RNase Z in prokaryotes. Nat Rev Microbiol 5, 278–286 (2007). https://doi.org/10.1038/nrmicro1622
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1622