Abstract
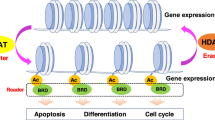
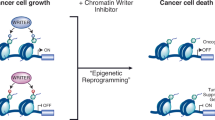
This review focuses on the discovery and development of the histone deacetylase (HDAC) inhibitor, suberoylanilide hydroxamic acid (SAHA). Post-translational modifications of the histones of chromatin are important factors in regulating gene expression—so-called epigenetic gene regulation. Acetylation and deacetylation of lysine residues in histone tails, controlled by the activities of HDACs and histone acetyltransferases, are among the most studied post-translational modification of histones. In addition to chromatin protein, transcription factors, cell-signaling regulatory proteins, and proteins regulating cell death are substrates of HDACs and may be altered in function by HDAC inhibitors. HDAC inhibitors have several remarkable aspects. For instance, despite HDACs being ubiquitously distributed through chromatin, SAHA selectively alters the transcription of relatively few genes, and normal cells are at least 10-fold more resistant than transformed cells to SAHA and related HDAC inhibitor-induced cell death. HDAC inhibitors represent a relatively new group of targeted anticancer compounds, which are showing significant promise as agents with activity against a broad spectrum of neoplasms, at doses that are well tolerated by cancer patients. SAHA is one of the HDAC inhibitors most advanced in development. It is in phase I and II clinical trials for patients with both hematologic and solid tumors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Miller TA et al. (2003) Histone deacetylase inhibitors. J Med Chem 46: 5097–5116
Marks PA et al. (2004) Histone deacetylase inhibitors. Adv Cancer Res 91: 137–168
Yoshida M et al. (1990) Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. J Biol Chem 265: 17174–17179
Kyrylenko S et al. (2003) Differential regulation of the Sir2 histone deacetylase gene family by inhibitors of class I and II histone deacetylases. Cell Mol Life Sci 60: 1990–1997
Remiszewski SW (2003) The discovery of NVP-LAQ824: from concept to clinic. Curr Med Chem 10: 2393–2402
Furumai R et al. (2002) FK228 (depsipeptide) as a natural prodrug that inhibits class I histone deacetylases. Cancer Res 62: 4916–4921
Piekarz R and Bates S (2004) A review of depsipeptide and other histone deacetylase inhibitors in clinical trials. Curr Pharm Des 10: 2289–2298
Thibault A et al. (1994) A phase I and pharmacokinetic study of intravenous phenylacetate in patients with cancer. Cancer Res 54: 1690–1694
Chang SM et al. (1999) Phase II study of phenylacetate in patients with recurrent malignant glioma: a North American Brain Tumor Consortium report. J Clin Oncol 17: 984–990
Reid T et al. (2004) Phase II trial of the histone deacetylase inhibitor pivaloyloxymethyl butyrate (Pivanex, AN-9) in advanced non-small cell lung cancer. Lung Cancer 45: 381–386
Atmaca A et al. (2004) A dose escalating phase I study with valproic acid (VPA) in patients with advanced cancer. ASCO Annual Meeting Proceedings (Post-Meeting Edition) [abstract #3169] J Clin Oncol 22 (Suppl)
Lucas DM et al. (2004) The histone deacetylase inhibitor MS-275 induces caspase-dependent apoptosis in B-cell chronic lymphocytic leukemia cells. Leukemia 18: 1207–1214
Kimmel KA et al. (2001) A Phase I dose-finding study of CI-994 in combination with capecitabine in patients with advanced solid tumors. [abstract #345] Proc Am Soc Clin Oncol 87: a345
Wolffe AP and Pruss D (1996) Deviant nucleosomes: the functional specialization of chromatin. Trends Genet 12: 58–62
Luger K et al. (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389: 251–260
Jenuwein T and Allis CD (2001) Translating the histone code. Science 293: 1074–1080
Spotswood HT and Turner BM (2002) An increasingly complex code. J Clin Invest 110: 577–582
Fischle W et al. (2003) Binary switches and modification cassettes in histone biology and beyond. Nature 425: 475–479
Gui CY et al. (2004) Histone deacetylase (HDAC) inhibitor activation of p21WAF1 involves changes in promoter-associated proteins, including HDAC1. Proc Natl Acad Sci U S A 101: 1241–1246
Arts J et al. (2003) Histone deacetylase inhibitors: from chromatin remodeling to experimental cancer therapeutics. Curr Med Chem 10: 2343–2350
Glaser KB et al. (2003) Role of class I and class II histone deacetylases in carcinoma cells using siRNA. Biochem Biophys Res Commun 310: 529–536
Cress WD and Seto E (2000) Histone deacetylases, transcriptional control, and cancer. J Cell Physiol 184: 1–16
de Ruijter AJ et al. (2003) Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem J 370 (Part 3): 737–749
Jones PA and Baylin SB (2002) The fundamental role of epigenetic events in cancer. Nat Rev Genet 3: 415–428
Johnstone RW and Licht JD (2003) Histone deacetylase inhibitors in cancer therapy: is transcription the primary target? Cancer Cell 4: 13–18
He LZ et al. (2001) Histone deacetylase inhibitors induce remission in transgenic models of therapy-resistant acute promyelocytic leukemia. J Clin Invest 108: 1321–1330
Murata T et al. (2001) Defect of histone acetyltransferase activity of the nuclear transcriptional coactivator CBP in Rubinstein–Taybi syndrome. Hum Mol Genet 10: 1071–1076
Finnin MS (1999) Structures of a histone deacetylase homologue bound to the TSA and SAHA inhibitors. Nature 401: 188–193
Gray SG et al. (2004) Microarray profiling of the effects of histone deacetylase inhibitors on gene expression in cancer cell lines. Int J Oncol 24: 773–795
Richon VM et al. (2000) Histone deacetylase inhibitor selectively induces p21WAF1 expression and gene-associated histone acetylation. Proc Natl Acad Sci U S A 97: 10014–10019
Mitsiades CS et al. (2004) Transcriptional signature of histone deacetylase inhibition in multiple myeloma: biological and clinical implications. Proc Natl Acad Sci U S A 101: 540–545
Butler LM et al. (2002) The histone deacetylase inhibitor SAHA arrests cancer cell growth, up-regulates thioredoxin-binding protein-2, and down-regulates thioredoxin. Proc Natl Acad Sci U S A 99: 11700–11705
Warrener R et al. (2003) Tumor cell-selective cytotoxicity by targeting cell cycle checkpoints. FASEB J 17: 1550–1552
Qiu L et al. (1999) Anti-tumour activity in vitro and in vivo of selective differentiating agents containing hydroxamate. Br J Cancer 80: 1252–1258
Polevoda B and Sherman F (2002) The diversity of acetylated proteins. Genome Biol 3: reviews 0006
Kouzarides T (2000) Acetylation: a regulatory modification to rival phosphorylation? EMBO J 19: 1176–1179
Pei XY et al. (2004) Synergistic induction of oxidative injury and apoptosis in human multiple myeloma cells by the proteasome inhibitor bortezomib and histone deacetylase inhibitors. Clin Cancer Res 10: 3839–3852
Rahmani M et al. (2003) Coadministration of the heat shock protein 90 antagonist 17-allylamino- 17-demethoxygeldanamycin with suberoylanilide hydroxamic acid or sodium butyrate synergistically induces apoptosis in human leukemia cells. Cancer Res 63: 8420–8427
Kelly WK et al. (2003) Phase I clinical trial of histone deacetylase inhibitor: suberoylanilide hydroxamic acid administered intravenously. Clin Cancer Res 9: 3578–3588
Duvic M et al. (2003) Phase II trial of oral suberoylanilide hydroxamic acid (SAHA) for cutaneous T-cell lymphoma (CTCL) and peripheral T-cell lymphoma (PTCL). [abstract #625] Proc Am Soc Hematol
Marks PA and Kelly WK Histone deacetylase inhibitors: novel targeted anti-cancer agents. In DNA Methylation Epigenetics and Metastasis. (Ed. Esteller M) Boston, Dordrecht, London: Kluwer Academic Publisher, in press
Ungerstedt JS et al. (2005) Role of thioredoxin in the response of normal and transformed cells to histone deacetylase inhibitor. Proc Natl Acad Sci U S A 102: 673–678
Peart MJ et al. (2003) Novel mechanisms of apoptosis induced by histone deacetylase inhibitors. Cancer Res 63: 4460–4471
Butler LM et al. (2000) Suberoylanilide hydroxamic acid, an inhibitor of histone deacetylase, suppresses the growth of prostate cancer cells in vitro and in vivo. Cancer Res 60: 5165–5170
Kelly WK et al. Phase I study of the oral histone deacetylase inhibitor: suberoylanilide hydroxamic acid (SAHA), in patients with advanced cancer. J Clin Oncol, in press
Piekarz RL et al. (2001) Inhibitor of histone deacetylation, depsipeptide (FR901228), in the treatment of peripheral and cutaneous T-cell lymphoma: a case report. Blood 98: 2865–2868
Piekarz RL et al. (2004) T-cell lymphoma as a model for the use of histone deacetylase inhibitors in cancer therapy: impact of depsipeptide on molecular markers, therapeutic targets, and mechanisms of resistance. Blood 103: 4636–4643
Acknowledgements
The studies reviewed in this chapter representing research in the authors' laboratories were supported, in part, by grants from: the National Cancer Institute (P30-CA-08748-40); Robert J and Helen C Kleberg Foundation; DeWitt Wallace Fund for the Memorial Sloan-Kettering Cancer Center; Susan and Jack Rudin Foundation; and The David H Koch Prostate Cancer Research Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Memorial Sloan-Kettering Cancer Center and Columbia University jointly hold patents on hydroxamic acid based polar compounds, including SAHA, which are exclusively licensed to Aton Pharma, Inc., a Biotechnology Company acquired by Merck, Inc. Paul A Marks was a founder of Aton and both institutions and the founder had an equity position in Aton Pharma, Inc. Paul A Marks is a scientific consultant to Merck.
Glossary
- PML-RARα
-
A specific chromosomal translocation, t(15;17), results in the fusion of the promyelocytic leukemia gene (PML) and the retinoic acid receptor gene (RARα)
- CBFA2T1
-
Core-binding factor 2 translocated to 1, also known as AML1/ETO
- RUBINSTEIN–TAYBI SYNDROME
-
A rare genetic disorder that causes developmental abnormalities of many organs, caused by mutations in the gene encoding CREB binding protein (CBP)
- CDKN1A
-
Gene encoding the cyclin-dependent kinase inhibitor-1A protein, which is an important inhibitor of cellular proliferation in response to DNA damage
- TNFSF10
-
Tumor necrosis factor (ligand) superfamily member 10 (also known as TRAIL or APO-2L)
Rights and permissions
About this article
Cite this article
Kelly, W., Marks, P. Drug Insight: histone deacetylase inhibitors—development of the new targeted anticancer agent suberoylanilide hydroxamic acid. Nat Rev Clin Oncol 2, 150–157 (2005). https://doi.org/10.1038/ncponc0106
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ncponc0106
This article is cited by
-
Suppression of histone deacetylases by SAHA relieves bone cancer pain in rats via inhibiting activation of glial cells in spinal dorsal horn and dorsal root ganglia
Journal of Neuroinflammation (2020)
-
Histone deacetylase inhibitor induces cell apoptosis and cycle arrest in lung cancer cells via mitochondrial injury and p53 up-acetylation
Cell Biology and Toxicology (2016)
-
Novel analogs targeting histone deacetylase suppress aggressive thyroid cancer cell growth and induce re-differentiation
Cancer Gene Therapy (2015)
-
Target deconvolution of bioactive small molecules: the heart of chemical biology and drug discovery
Archives of Pharmacal Research (2015)
-
Phase I study of combination of vorinostat, carboplatin, and gemcitabine in women with recurrent, platinum-sensitive epithelial ovarian, fallopian tube, or peritoneal cancer
Cancer Chemotherapy and Pharmacology (2015)