Abstract
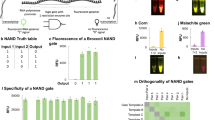
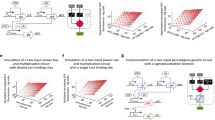
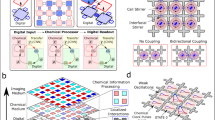
Autonomous programmable computing devices made of biomolecules could interact with a biological environment and be used in future biological and medical applications1,2,3,4,5,6,7. Biomolecular implementations of finite automata8,9 and logic gates4,10,11,12,13 have already been developed14,15,16,17,18. Here, we report an autonomous programmable molecular system based on the manipulation of DNA strands that is capable of performing simple logical deductions. Using molecular representations of facts such as Man(Socrates) and rules such as Mortal(X) ← Man(X) (Every Man is Mortal), the system can answer molecular queries such as Mortal(Socrates)? (Is Socrates Mortal?) and Mortal(X)? (Who is Mortal?). This biomolecular computing system compares favourably with previous approaches in terms of expressive power, performance and precision2,4,8,9,11,12,19. A compiler translates facts, rules and queries into their molecular representations and subsequently operates a robotic system that assembles the logical deductions and delivers the result. This prototype is the first simple programming language with a molecular-scale implementation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
24 August 2009
In the version of this Letter initially published online, the 'Query' that appeared in lines 1, 3, 5 and 7 of Fig. 3b was incorrect. This has been corrected for all versions of the Letter.
References
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Benenson, Y., Gil, B., Ben-Dor, U., Adar, R. & Shapiro, E. An autonomous molecular computer for logical control of gene expression. Nature 429, 423–429 (2004).
Basu, S., Gerchman, Y., Collins, C. H., Arnold, F. H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Rinaudo, K. et al. A universal RNAi-based logic evaluator that operates in mammalian cells. Nature Biotechnol. 25, 795–801 (2007).
Deans, T. L., Cantor, C. R. & Collins, J. J. A tunable genetic switch based on RNAi and repressor proteins for regulating gene expression in mammalian cells. Cell 130, 363–372 (2007).
Kobayashi, H. et al. Programmable cells: interfacing natural and engineered gene networks. Proc. Natl Acad. Sci. USA 101, 8414–8419 (2004).
Benenson, Y. et al. Programmable and autonomous computing machine made of biomolecules. Nature 414, 430–434 (2001).
Stojanovic, M. N. & Stefanovic, D. A deoxyribozyme-based molecular automaton. Nature Biotechnol. 21, 1069–1074 (2003).
Breaker, R. R. Engineered allosteric ribozymes as biosensor components. Curr. Opin. Biotechnol. 31–39 (2002).
Macdonald, J. et al. Medium scale integration of molecular logic gates in an automaton. Nano Lett. 6, 2598–2603 (2006).
Zhang, D. Y., Turberfield, A. J., Yurke, B. & Winfree, E. Engineering entropy-driven reactions and networks catalyzed by DNA. Science 318, 1121–1125 (2007).
Isaacs, F. J. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nature Biotechnol. 22, 841–847 (2004).
Kobayashi, S., Yokomori, T., Sampei, G. & Mizobuchi, K. DNA implementation of simple horn clause computation. IEEE International Conference on Evolutionary Computation, 213–217 (Indianapolis, USA, 1997).
Kobayashi, S. Horn clause computation with DNA molecules. J. Comb. Optim. 3, 277–299 (1999).
Siewicz, P., Janczak, T., Mulawaka, J. J. & Plucienniczak, A. The inference via DNA computing. Congress on Evolutionary Computation, 988–993 (Washington, USA, 1999).
Siewicz, P., Janczak, T., Mulawaka, J. J. & Plucienniczak, A. The inference based on molecular computing. Int. J. Cybern. Syst. 31/3, 283–315 (2000).
Uejima, H., Hagiya, M. & Kobayashi, S. Horn clause computation by self-assembly of DNA molecules. Lecture Notes in Computer Science, DNA Computing 7, 308–320 (2002).
Adar, R. et al. Stochastic computing with biomolecular automata. Proc. Natl Acad. Sci. USA 101, 9960–9965 (2004).
Sterling, L. & Shapiro, E. The Art of Prolog: Advanced Programming Techniques (MIT Press, 1986, 1994).
Lloyd, J. W. Foundations of Logic Programming (Springer-Verlag, 1987).
Bratko, I. Prolog Programming for Artificial Intelligence (Addison-Wesley, 2001).
Covington, M. A. Natural Language Processing for Prolog Programmers (Prentice Hall, 1994).
Gazdar, G. & Mellish, C. S. Natural Language Processing in Prolog: an Introduction to Computational Linguistics (Addison-Wesley, 1989).
Shapiro, E. Concurrent Prolog: Collected Papers, Vols 1 and 2 (MIT Press, 1987).
Tarnland, S. A. Horn clause computability. J. Comb. Optim. 17, 215–226 (1977).
Shapiro, E. Alternation and the computational complexity of logic programs. J. Logic Programming 1, 19–33 (1984).
Benenson, Y., Adar, R., Paz-Elizur, T., Livneh, Z. & Shapiro, E. DNA molecule provides a computing machine with both data and fuel. Proc. Natl Acad. Sci. USA 100, 2191–2196 (2003).
Maslow, A. A theory of human motivation. Psychol. Rev. 50, 370–396 (1943).
Winfree, E. Biochemical logic: submerged circuits of floating DNA. ENGenious, 52–54 (2007).
Acknowledgements
We thank R. Adar, M. Kahan and B. Gil for their assistance and advice and A. Mishali and G. Brodsky for the preparation of the figures. This research was supported by The Israel Science Foundation grant no. 285/02, a research grant from the Clore Center for Biological Physics, a research grant from the Louis Chor Memorial Trust, a research grant from the Estate of Funnie Sherr, the Cymerman-Jubskind Prize, the Estate of Karl Felix Jakubskind and by the European Union FP7-ERC-AdG. S.K. is supported by the Yeshaya Horowitz association through the Center for Complexity Science. E.S. is the Incumbent of the Harry Weinrebe Professorial Chair of Computer Science and Biology and of the France Telecom–Orange Excellence Chair for Interdisciplinary Studies of the Paris ‘Centre de Recherche Interdisciplinaire’ (FTO/CRI).
Author information
Authors and Affiliations
Contributions
E.S. led the project. T.R. conceived the biomolecular design and the compiler and designed and performed the experiments. T.R. and S.K. wrote the analysis tools and automated the experimental protocols. T.R. and E.S. wrote the paper.
Corresponding author
Supplementary information
Supplementary information
Supplementary information (PDF 1419 kb)
Rights and permissions
About this article
Cite this article
Ran, T., Kaplan, S. & Shapiro, E. Molecular implementation of simple logic programs. Nature Nanotech 4, 642–648 (2009). https://doi.org/10.1038/nnano.2009.203
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2009.203
This article is cited by
-
Probabilistic reasoning with a Bayesian DNA device based on strand displacement
Natural Computing (2014)
-
Programmable chemical controllers made from DNA
Nature Nanotechnology (2013)
-
Biomolecular computing systems: principles, progress and potential
Nature Reviews Genetics (2012)