Abstract
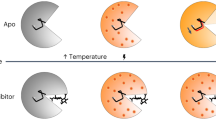
Biomolecules are highly pressure-sensitive, but their dynamics upon return to ambient pressure are often too fast to observe with existing approaches. We describe a sample-efficient method capable of large and very fast pressure drops (<1 nanomole, >2,500 atmospheres and <0.7 microseconds). We validated the method by fluorescence-detected refolding of a genetically engineered lambda repressor mutant from its pressure-denatured state. We resolved barrierless structure formation upon return to ambient pressure; we observed a 2.1 ± 0.7 microsecond refolding time, which is very close to the 'speed limit' for proteins and much faster than the corresponding temperature-jump refolding of the same protein. The ability to experimentally perform a large and very fast pressure drop opens up a new region of the biomolecular energy landscape for atomic-level simulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hartl, F.U. Molecular chaperones in cellular protein folding. Nature 381, 571–580 (1996).
Martin, J.S., Brown, L. & Haberstroh, K. Microfilaments are involved in renal cell responses to sustained hydrostatic pressure. J. Urol. 173, 1410–1417 (2005).
Eigen, M. & Maeyer, L.D. Relaxation methods. In Technique of Organic Chemistry (ed., Weissberger, A.) 895–1054 (Interscience, New York, 1963).
Turner, D.H., Flynn, G.W., Lundberg, S.K., Faller, L.D. & Sutin, N. Dimerization of proflavin by the laser Raman temperature-jump method. Nature 239, 215–217 (1972).
Jennings, P.A. & Wright, P. Formation of a molten globule intermediate early in the kinetic folding pathway of apomyoglobin. Science 262, 892–895 (1993).
Phillips, C.M., Mizutani, Y. & Hochstrasser, R.M. Ultrafast thermally induced unfolding of RNase A. Proc. Natl. Acad. Sci. USA 92, 7292–7296 (1995).
Jacob, M. et al. Microsecond folding of the cold shock protein measured by a pressure-jump technique. Biochemistry 38, 2882–2891 (1999).
Mitra, L. et al. V-i-value analysis: a pressure-based method for mapping the folding transition state ensemble of proteins. J. Am. Chem. Soc. 129, 14108–14109 (2007).
Ballew, R.M., Sabelko, J., Reiner, C. & Gruebele, M. A single-sweep, nanosecond time resolution laser temperature-jump apparatus. Rev. Sci. Instrum. 67, 3694–3699 (1996).
Royer, C.A. The nature of the transition state ensemble and the mechanisms of protein folding: A review. Arch. Biochem. Biophys. 469, 34–45 (2008).
Pearson, D.S., Swartz, D.R. & Geeves, M.A. Fast pressure jumps can perturb calcium and magnesium binding to troponin C F29W. Biochemistry 47, 12146–12158 (2008).
Seddon, J.M. et al. Pressure-jump X-ray studies of liquid crystal transitions in lipids. Philos. Transact. A Math. Phys. Eng. Sci. 364, 2635–2655 (2006).
Winter, R. & Dzwolak, W. Exploring the temperature-pressure configurational landscape of biomolecules: from lipid membranes to proteins. Philos. Transact. A Math. Phys. Eng. Sci. 363, 537–562 (2005).
Kegeles, G. & Ke, C.-H. A light-scattering pressure-jump kinetics apparatus. Anal. Biochem. 68, 138–147 (1975).
Clegg, R.M., Elson, E.L. & Maxfield, B.W. New technique for optical observation of the kinetics of chemical reactions perturbed by small pressure changes. Biopolymers 14, 883–887 (1975).
Knoche, W. & Wiese, G. Improved apparatus for pressure-jump relaxation measurements. Chemical Instrumentation 5, 91–98 (1973).
Magg, C., Kubelka, J., Holtermann, G., Haas, E. & Schmid, F.X. Specificity of the initial collapse in the folding of the cold shock protein. J. Mol. Biol. 360, 1067–1080 (2006).
Duan, Y. & Kollman, P.A. Pathways to a protein folding intermediate observed in a 1-microsecond simulation in aqueous solution. Science 282, 740–744 (1998).
Snow, C.D., Nguyen, H., Pande, V. & Gruebele, M. Absolute comparison of simulated and experimental protein folding dynamics. Nature 420, 102–106 (2002).
Paschek, D., Hempel, S. & Garcia, A.E. Computing the stability diagram ofthe Trp-cage miniprotein. Proc. Natl. Acad. Sci. USA 105, 17754–17759 (2008).
Kubelka, J., Hofrichter, J. & Eaton, W.A. The protein folding 'speed limit'. Curr. Opin. Struct. Biol. 14, 76–88 (2004).
Yang, W.Y. & Gruebele, M. Folding at the speed limit. Nature 423, 193–197 (2003).
Freddolino, P.L., Liu, F., Gruebele, M. & Schulten, K. Ten-microsecond molecular dynamics simulation of a fast-folding WW domain. Biophys. J. 94, L75–L77 (2008).
Snow, C.D., Sorin, E.J., Rhee, Y.M. & Pande, V.S. How well can simulation predict protein folding kinetics and thermodynamics. Annu. Rev. Biophys. Biomol. Struct. 34, 43–69 (2005).
Pitera, J.W., Swope, W.C. & Abraham, F.F. Observation of noncooperative folding thermodynamics in simulations of 1BBL. Biophys. J. 94, 4837–4846 (2008).
Yang, W.Y. & Gruebele, M. Rate-temperature relationships in lambda repressor fragment 6–85 folding. Biochemistry 43, 13018–13025 (2004).
Dumont, C. et al. Solvent-tuning collapse and helix formation time scales of lambda6–85. Protein Sci. 15, 2596–2604 (2006).
Yang, W.Y. & Gruebele, M. Kinetic equivalence of the heat and cold structural transitions of lambda6–85. Phil. Trans. R. Soc. Lond. B 43, 13018–13025 (2005).
Bryngelson, J.D., Onuchic, J.N., Socci, N.D. & Wolynes, P.G. Funnels, pathways, and the energy landscape of protein folding: a synthesis. Proteins Struct. Funct. Genet. 21, 167–195 (1995).
Camilloni, C. et al. Urea and guanidinium chloride denature protein L in different ways in molecular dynamics simulations. Biophys. J. 94, 4654–4661 (2008).
Acknowledgements
We acknowledge funding from the National Science Foundation (MCB 0613643), assistance with steady-state data collection by S. Ates, help with the prototype setup by J. Ervin and J. Lee and helpful discussions with H. Bohr. T. Oas (Duke University) provided the pseudo-wild-type lambda repressor fragment plasmid as a gift.
Author information
Authors and Affiliations
Contributions
C.D. performed the experiments and did data analysis. T.E. designed and made experimental components. M.G. designed the experiment, performed data analysis and wrote the article.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–2 (PDF 871 kb)
Rights and permissions
About this article
Cite this article
Dumont, C., Emilsson, T. & Gruebele, M. Reaching the protein folding speed limit with large, sub-microsecond pressure jumps. Nat Methods 6, 515–519 (2009). https://doi.org/10.1038/nmeth.1336
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1336
This article is cited by
-
Effect of X-ray free-electron laser-induced shockwaves on haemoglobin microcrystals delivered in a liquid jet
Nature Communications (2021)
-
Directly monitor protein rearrangement on a nanosecond-to-millisecond time-scale
Scientific Reports (2017)
-
Ultrafast structural molecular dynamics investigated with 2D infrared spectroscopy methods
Topics in Current Chemistry (2017)
-
Fast-folding proteins under stress
Cellular and Molecular Life Sciences (2015)
-
Downhill protein folding under pressure
Nature Methods (2009)