Abstract
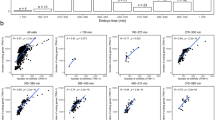
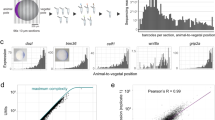
Caenorhabditis elegans is one of the most prominent model systems for embryogenesis, but collecting many precisely staged embryos has been impractical. Thus, early C. elegans embryogenesis has not been amenable to most high-throughput genomics or biochemistry assays. To overcome this problem, we devised a method to collect staged C. elegans embryos by fluorescence-activated cell sorting (eFACS). In a proof-of-principle experiment, we found that a single eFACS run routinely yielded tens of thousands of almost perfectly staged 1-cell stage embryos. As the earliest embryonic events are driven by posttranscriptional regulation, we combined eFACS with second-generation sequencing to profile the embryonic expression of small, noncoding RNAs. We discovered complex and orchestrated changes in the expression between and within almost all classes of small RNAs, including microRNAs and 26G-RNAs, during embryogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Sulston, J.E., Schierenberg, E., White, J.G. & Thomson, J.N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983).
Piano, F. et al. RNAi analysis of genes expressed in the ovary of Caenorhabditis elegans. Curr. Biol. 10, 1619–1622 (2000).
Piano, F. et al. Gene clustering based on RNAi phenotypes of ovary-enriched genes in C. elegans. Curr. Biol. 12, 1959–1964 (2002).
Kamath, R.S. et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421, 231–237 (2003).
Gonczy, P. & Rose, L.S. Asymmetric cell division and axis formation in the embryo. WormBook 2005, 1–20 (2005).
Sonnichsen, B. et al. Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans. Nature 434, 462–469 (2005).
Fernandez, A.G. et al. New genes with roles in the C. elegans embryo revealed using RNAi of ovary-enriched ORFeome clones. Genome Res. 15, 250–259 (2005).
Oegema, K. & Hyman, A.A. Cell division. WormBook 2006, 1–40 (2006).
Stroeher, V.L. et al. DNA-protein interactions in the Caenorhabditis elegans embryo: oocyte and embryonic factors that bind to the promoter of the gut-specific ges-1 gene. Dev. Biol. 163, 367–380 (1994).
Schauer, I.E. & Wood, W.B. Early C. elegans embryos are transcriptionally active. Development 110, 1303–1317 (1990).
Edgar, L.G., Wolf, N. & Wood, W.B. Early transcription in Caenorhabditis elegans embryos. Development 120, 443–451 (1994).
Seydoux, G. & Dunn, M.A. Transcriptionally repressed germ cells lack a subpopulation of phosphorylated RNA polymerase II in early embryos of Caenorhabditis elegans and Drosophila melanogaster. Development 124, 2191–2201 (1997).
Denli, A.M. et al. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004).
Grishok, A. et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106, 23–34 (2001).
Knight, S.W. & Bass, B.L. A role for the RNase III enzyme DCR-1 in RNA interference and germ line development in Caenorhabditis elegans. Science 293, 2269–2271 (2001).
Ambros, V. et al. MicroRNAs and other tiny endogenous RNAs in C. elegans. Curr. Biol. 13, 807–818 (2003).
Baugh, L.R. et al. Composition and dynamics of the Caenorhabditis elegans early embryonic transcriptome. Development 130, 889–900 (2003).
Evans, T.C. & Hunter, C.P. Translational control of maternal RNAs. WormBook 2005, 1–11 (2005).
Ruby, J.G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–1207 (2006).
Lin, R. A gain-of-function mutation in oma-1, a C. elegans gene required for oocyte maturation, results in delayed degradation of maternal proteins and embryonic lethality. Dev. Biol. 258, 226–239 (2003).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Friedlander, M.R. et al. Discovering microRNAs from deep sequencing data using miRDeep. Nat. Biotechnol. 26, 407–415 (2008).
Batista, P.J. et al. PRG-1 and 21U-RNAs interact to form the piRNA complex required for fertility in C. elegans. Mol. Cell 31, 67–78 (2008).
Ghildiyal, M. & Zamore, P.D. Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 10, 94–108 (2009).
Nance, J., Munro, E.M. & Priess, J.R. C. elegans PAR-3 and PAR-6 are required for apicobasal asymmetries associated with cell adhesion and gastrulation. Development 130, 5339–5350 (2003).
Miska, E.A. et al. Most Caenorhabditis elegans microRNAs are individually not essential for development or viability. PLoS Genet. 3, e215 (2007).
Sutovsky, P. Ubiquitin-dependent proteolysis in mammalian spermatogenesis, fertilization, and sperm quality control: killing three birds with one stone. Microsc. Res. Tech. 61, 88–102 (2003).
Aravin, A.A. et al. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316, 744–747 (2007).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007).
Stiernagle, T. Maintenance of C. elegans. WormBook 2006, 1–11 (2006).
Hoffman, R.A. & Wood, J.C.S. Characterization of flow cytometer instrument sensitivity. Curr. Protoc. Cytometry 1.20, 1.20.21–21.20.18 (2007).
Griffiths-Jones, S., Saini, H.K., van Dongen, S. & Enright, A.J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–D158 (2008).
Jurka, J. Repbase update: a database and an electronic journal of repetitive elements. Trends Genet. 16, 418–420 (2000).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Karolchik, D. et al. The UCSC Genome Browser Database: 2008 update. Nucleic Acids Res. 36, D773–D779 (2008).
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001).
Acknowledgements
We thank R. Lin (University of Texas) for providing us with the TX189(P(oma-1)∷oma-1∷GFP) strain. All other strains used in this project were provided by the Caenorhabditis Genetic Center, which is funded by the US National Center for Research Resources. M.S. acknowledges part-time funding from the Berlin Institute for Medical Systems Biology, funded by Bundesministerium für Bildung und Forschung, and New York University PhD exchange program, and a travel grand from Boehringer Ingelheim Fonds. J.M. thanks the Deutsche forschungsgemeinschaft for a fellowship in the International Research Training Group Genomics and Systems Biology of Molecular Networks (GRK 1360). F.P. and N.R. acknowledge partial funding from US National Human Genome Research Institute (ModEncode U01 HG004276) and US National Institutes of Health (R01HD046236). We thank S. Lebedeva for help with sequencing runs.
Author information
Authors and Affiliations
Contributions
F.P. and N.R. conceived, designed and supervised the study. M.S. designed and performed the experiments. J.M. designed and performed computational studies with the exception of predicting new miRNAs, which was done by M.R.F.; T.C. contributed to initial eFACS experiments; H.-P.R. helped with flow cytometer settings and runs; W.C. and N.L. contributed to library preparations and sequencing; M.S., J.M., F.P. and N.R. analyzed the data. M.S. and N.R. wrote the paper, and J.M. and F.P. edited it.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–3,5,7–9 (PDF 2957 kb)
Supplementary Table 4
miRNA predictions using miRDeep. The table depicts 19 predicted miRNAs from our datasets including star and precursor sequences. (XLS 46 kb)
Supplementary Table 6
Expression of 21U-RNAs. We retrieved 15,703 21U-RNA sequences from the supplementary material of Batista et al, 2009, and mapped them to the genome. The subset mapping to unique positions in the genome was quantified by reporting weighted mappings of reads that overlap with each 21U-RNA. (XLS 1488 kb)
Rights and permissions
About this article
Cite this article
Stoeckius, M., Maaskola, J., Colombo, T. et al. Large-scale sorting of C. elegans embryos reveals the dynamics of small RNA expression. Nat Methods 6, 745–751 (2009). https://doi.org/10.1038/nmeth.1370
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1370
This article is cited by
-
Germline inherited small RNAs facilitate the clearance of untranslated maternal mRNAs in C. elegans embryos
Nature Communications (2021)
-
Composition of Caenorhabditis elegans extracellular vesicles suggests roles in metabolism, immunity, and aging
GeroScience (2020)
-
Comparative RNA-Seq analysis reveals pervasive tissue-specific alternative polyadenylation in Caenorhabditis elegans intestine and muscles
BMC Biology (2015)
-
High-throughput fluorescence-based isolation of live C. elegans larvae
Nature Protocols (2012)
-
The early bird catches the worm: new technologies for the Caenorhabditis elegans toolkit
Nature Reviews Genetics (2011)