Abstract
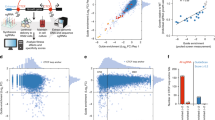
RNA interference (RNAi) has become an important technique for loss-of-gene-function studies in mammalian cells. To achieve reliable results in an RNAi experiment, efficient and specific silencing triggers are required. Here we present genome-wide data sets for the production of endoribonuclease-prepared short interfering RNAs (esiRNAs) for human, mouse and rat. We used an algorithm to predict the optimal region for esiRNA synthesis for every protein-coding gene of these three species. We created a database, RiDDLE, for retrieval of target sequences and primer information. To test this in silico resource experimentally, we generated 16,242 esiRNAs that can be used for RNAi screening in human cells. Comparative analyses with chemically synthesized siRNAs demonstrated a high silencing efficacy of esiRNAs and a 12-fold reduction of downregulated off-target transcripts as detected by microarray analysis. Hence, the presented esiRNA libraries offer an efficient, cost-effective and specific alternative to presently available mammalian RNAi resources.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Elbashir, S.M. et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001).
Hannon, G.J. & Rossi, J.J. Unlocking the potential of the human genome with RNA interference. Nature 431, 371–378 (2004).
Brummelkamp, T.R., Bernards, R. & Agami, R. A system for stable expression of short interfering RNAs in mammalian cells. Science 296, 550–553 (2002).
Paddison, P.J., Caudy, A.A., Bernstein, E., Hannon, G.J. & Conklin, D.S. Short hairpin RNAs (shRNAs) induce sequence-specific silencing in mammalian cells. Genes Dev. 16, 948–958 (2002).
Silva, J.M. et al. Second-generation shRNA libraries covering the mouse and human genomes. Nat. Genet. 37, 1281–1288 (2005).
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006).
Root, D.E., Hacohen, N., Hahn, W.C., Lander, E.S. & Sabatini, D.M. Genome-scale loss-of-function screening with a lentiviral RNAi library. Nat. Methods 3, 715–719 (2006).
Bernards, R., Brummelkamp, T.R. & Beijersbergen, R.L. shRNA libraries and their use in cancer genetics. Nat. Methods 3, 701–706 (2006).
Reynolds, A. et al. Rational siRNA design for RNA interference. Nat. Biotechnol. 22, 326–330 (2004).
Pei, Y. & Tuschl, T. On the art of identifying effective and specific siRNAs. Nat. Methods 3, 670–676 (2006).
Jackson, A.L. et al. Expression profiling reveals off-target gene regulation by RNAi. Nat. Biotechnol. 21, 635–637 (2003).
Jackson, A.L. & Linsley, P.S. Noise amidst the silence: off-target effects of siRNAs? Trends Genet. 20, 521–524 (2004).
Scacheri, P.C. et al. Short interfering RNAs can induce unexpected and divergent changes in the levels of untargeted proteins in mammalian cells. Proc. Natl. Acad. Sci. USA 101, 1892–1897 (2004).
Lin, X. et al. siRNA-mediated off-target gene silencing triggered by a 7 nt complementation. Nucleic Acids Res. 33, 4527–4535 (2005).
Jackson, A.L. et al. Widespread siRNA “off-target” transcript silencing mediated by seed region sequence complementarity. RNA 12, 1179–1187 (2006).
Birmingham, A. et al. 3′ UTR seed matches, but not overall identity, are associated with RNAi off-targets. Nat. Methods 3, 199–204 (2006).
Buchholz, F., Kittler, R., Slabicki, M. & Theis, M. Enzymatically prepared RNAi libraries. Nat. Methods 3, 696–700 (2006).
Kittler, R. et al. An endoribonuclease-prepared siRNA screen in human cells identifies genes essential for cell division. Nature 432, 1036–1040 (2004).
Henschel, A., Buchholz, F. & Habermann, B. DEQOR: a web-based tool for the design and quality control of siRNAs. Nucleic Acids Res. 32, W113–W120 (2004).
Kittler, R., Heninger, A.K., Franke, K., Habermann, B. & Buchholz, F. Production of endoribonuclease-prepared short interfering RNAs for gene silencing in mammalian cells. Nat. Methods 2, 779–784 (2005).
Heale, B.S., Soifer, H.S., Bowers, C. & Rossi, J.J. siRNA target site secondary structure predictions using local stable substructures. Nucleic Acids Res. 33, e30 (2005).
Schubert, S., Grunweller, A., Erdmann, V.A. & Kurreck, J. Local RNA target structure influences siRNA efficacy: systematic analysis of intentionally designed binding regions. J. Mol. Biol. 348, 883–893 (2005).
Kulkarni, M.M. et al. Evidence of off-target effects associated with long dsRNAs in Drosophila melanogaster cell-based assays. Nat. Methods 3, 833–838 (2006).
Ma, Y., Creanga, A., Lum, L. & Beachy, P.A. Prevalence of off-target effects in Drosophila RNA interference screens. Nature 443, 359–363 (2006).
Myers, J.W. et al. Minimizing off-target effects by using diced siRNAs for RNA interference. Journal of RNAi and Gene Silencing 2, 181–194 (2006).
Manber, U. & Myers, G. Suffix arrays: a new method for on-line string searches. SIAM Journal on Computing 22, 935–948 (1993).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386 (2000).
Hughes, T.R. et al. Expression profiling using microarrays fabricated by an ink-jet oligonucleotide synthesizer. Nat. Biotechnol. 19, 342–347 (2001).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Marchler-Bauer, A. et al. CDD: a database of conserved domain alignments with links to domain three-dimensional structure. Nucleic Acids Res. 30, 281–283 (2002).
Acknowledgements
We thank Eberhard Krausz (Technology Development Studio, Max Planck Institute for Molecular Cell Biology and Genetics) for providing robotics support, and the Rosetta Gene Expression Laboratory for microarray hybridizations. This work was supported by the EU grants “FunGenES” (LSHG-CT-2003-503494), “Mitocheck” (LSHG-CT-2004-503464), by BMBF grant PTJ-BIO/0313130, the NGFN2 grant SMP-RNAi (01GR0402) and the Max Planck Society.
Author information
Authors and Affiliations
Contributions
R.K., A.K.H., M.S., M.T., G.P., K.F. and A.C. generated the human esiRNA library; V.S. and B.H. performed the in silico analyses; H.G. and J.W. performed automation; K.K. generated LIMS; E.R. and B.K. generated esiRNAs; C.F. performed QPCRs; C.S. and B.S. performed and analyzed QPCRs; J.G., J.S., J.B. and A.L.J. performed and analyzed microarray studies; P.S.L. analyzed microarray data; R.K. and F.B. designed and analyzed the experiments; R.K., V.S., A.L.J., B.H. and F.B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
V.S. and B.H. work for Scionics Computer Innovation.
E.R. and B.K. work for the RZPD.
C.F., C.S. and B.S. work for Cenix Bioscience.
J.G., J.S., J.B., P.S.L. and A.L.J. work for Rosetta Inpharmatics.
Supplementary information
Supplementary Fig. 1
Correlation of silencing efficacy between esiRNAs and siRNAs. (PDF 452 kb)
Supplementary Fig. 2
No induction of interferon response genes by esiRNA and siRNA. (PDF 600 kb)
Supplementary Fig. 3
Comparative analysis of off-target regulation. (PDF 319 kb)
Supplementary Table 1
Primer sequences used to generate esiRNAs. (PDF 13 kb)
Supplementary Table 2
Sequences of employed siRNAs. (PDF 11 kb)
Supplementary Table 3
Primers employed for QPCRs. (PDF 10 kb)
Rights and permissions
About this article
Cite this article
Kittler, R., Surendranath, V., Heninger, AK. et al. Genome-wide resources of endoribonuclease-prepared short interfering RNAs for specific loss-of-function studies. Nat Methods 4, 337–344 (2007). https://doi.org/10.1038/nmeth1025
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1025
This article is cited by
-
Loss of USP28 and SPINT2 expression promotes cancer cell survival after whole genome doubling
Cellular Oncology (2022)
-
Resolution of R-loops by INO80 promotes DNA replication and maintains cancer cell proliferation and viability
Nature Communications (2020)
-
In-cell identification and measurement of RNA-protein interactions
Nature Communications (2019)
-
The lincRNA MIRAT binds to IQGAP1 and modulates the MAPK pathway in NRAS mutant melanoma
Scientific Reports (2018)
-
Evaluation and control of miRNA-like off-target repression for RNA interference
Cellular and Molecular Life Sciences (2018)