Abstract
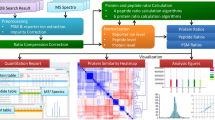
To take advantage of the potential quantitative benefits offered by tandem mass spectrometry, we have modified the method in which tandem mass spectrum data are acquired in 'shotgun' proteomic analyses. The proposed method is not data dependent and is based on the sequential isolation and fragmentation of precursor windows (of 10 m/z) within the ion trap until a desired mass range has been covered. We compared the quantitative figures of merit for this method to those for existing strategies by performing an analysis of the soluble fraction of whole-cell lysates from yeast metabolically labeled in vivo with 15N. To automate this analysis, we modified software (RelEx) previously written in the Yates lab to generate chromatograms directly from tandem mass spectra. These chromatograms showed improvements in signal-to-noise ratio of approximately three- to fivefold over corresponding chromatograms generated from mass spectrometry scans. In addition, to demonstrate the utility of the data-independent acquisition strategy coupled with chromatogram reconstruction from tandem mass spectra, we measured protein expression levels in two developmental stages of Caenorhabditis elegans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Griffin, T.J. et al. Abundance ratio-dependent proteomic analysis by mass spectrometry. Anal. Chem. 75, 867–874 (2003).
Wu, C., MacCoss, M. & Yates, J.R. III . Metabolic labeling of mammalian organisms with stable isotopes for quantitative proteomic analysis. Anal. Chem. in the press (2004).
Han, D.K., Eng, J., Zhou, H. & Aebersold, R. Quantitative profiling of differentation-induced microsomal proteins using isotope-coded affinity tags and mass spectrometry. Nat. Biotechnol. 19, 946–951 (2001).
Gygi, S.P., Rist, B., Griffin, T.J., Eng, J. & Aebersold, R. Proteome analysis of low-abundance proteins using multidimensional chromatography and isotope-coded affinity tags. J. Proteome Res. 1, 47–54 (2002).
Washburn, M.P., Ulaszek, R., Deciu, C., Schieltz, D.M. & Yates, J.R. III. Analysis of quantitative proteomic data generated via multidimensional protein identification technology. Anal. Chem. 74, 1650–1657 (2002).
Arnott, D. et al. Selective detection of membrane proteins without antibodies. Mol. Cell. Proteomics 1, 148–156 (2002).
Ong, S.-E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Berger, S.J. et al. High-throughput global peptide proteomic analysis by combining stable isotope amino acid labeling and data-dependent multiplexed-MS/MS. Anal. Chem. 74, 4994–5000 (2002).
Owens, K.G. Application of correlation analysis techniques to mass spectral data. Appl. Spectrosc. Rev. 27, 1–49 (1992).
Quackenbush, J. Microarray data normalization and transformation. Nat. Genet. 32, 496–501 (2002).
Yang, I.V. et al. Within the fold: assessing differential expression measures and reproducibility in microarray assays. Genome Biol. 3, 0062 (2002).
Ong, S.-E., Kratchmarova, I. & Mann, M. Properties of 13C-substituted arginine in stable isotope labeling by amino acids in cell culture (SILAC). J. Proteome Res. 2, 173–181 (2003).
MacCoss, M.J., Wu, C.C. & Yates, J.R. III. A correlation algorithm for the automated analysis of quantitative 'shotgun' proteomics data. Anal. Chem. 75, 6912–6921 (2003).
Fenyö, D. & Beavis, R.C. A method for assessing the statistical significance of mass spectrometry–based protein identifications using general scoring schemes. Anal. Chem. 75, 768–774 (2003).
Anderson, D.C., Li, W., Payan, D.G. & Noble, W.S. A new algorithm for the evaluation of shotgun peptide sequencing in proteomics: support vector machine classification of peptide MS/MS spectra and SEQUEST scores. J. Proteome Res. 2, 137–146 (2003).
Olsen, J.V., Ong, S.-E. & Mann, M. Trypsin cleaves exclusively C-terminal to arginine and lysine residues. Mol. Cell. Proteom. (2004).
Clauser, K.R., Baker, P. & Burlingame, A.L. Role of accurate mass measurement (10 ppm) in protein identification strategies employing MS or MS/MS and database searching. Anal. Chem. 71, 2871–2882 (1999).
Purvine, S., Eppel, J.-T., Yi, E.C. & Goodlett, D.R. Shotgun collision–induced dissociation of peptides using a time of flight mass analyzer. Proteomics 3, 847–850 (2003).
Acknowledgements
The authors would especially like to thank M. MacCoss for his insightful comments about the manuscript and detailed knowledge of RelEx. They thank H. McDonald and C. Wu for discussion and critical reading of the manuscript, and H.R. Horvitz, A.P. Page and J. Ahnn for the SQV-4, CYP-5, and CRT-1 antibodies. J.D.V. and J.W. are supported by National Research Service Award (NIH) fellowships. J.R.Y. is supported by National Institutes of Health grants RR11823, R01MH067880 and R21067598-01.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Example tandem mass spectra obtained using different isolation widths (a) 1, (b) 20, and (c) 100 m/z for a 1 pmol/μl infusion of Glu-fibrinopeptide B. (PDF 46 kb)
Supplementary Fig. 2
Relex screenshots detailing the 6 reconstructed chromatograms from MS scans used to quantify RPL19A in the 1:1 yeast whole cell lysate standard mixture. (PDF 76 kb)
Supplementary Fig. 3
Relex screenshots detailing the 6 reconstructed chromatograms from MS scans used to quantify RPL19A in the 10:1 yeast whole cell lysate standard mixture. (PDF 73 kb)
Supplementary Fig. 4
Relex screenshots detailing the 10 reconstructed chromatograms from tandem mass spectra used to quantify RPL19A in the 1:1 yeast whole cell lysate standard mixture. (PDF 95 kb)
Supplementary Fig. 5
Relex screenshots detailing the 7 reconstructed chromatograms from tandem mass spectra used to quantify RPL19A in the 10:1 yeast whole cell lysate standard mixture. (PDF 60 kb)
Supplementary Table 1
Signal to noise enhancements we observed for chromatograms extracted from both MS scans and tandem mass spectra for a collection of peptides identified in a whole cell yeast lysate. (PDF 17 kb)
Supplementary Table 2
Averages and standard deviations of protein ratios identified from unlabeled and metabolically 15N labeled peptides from whole cell yeast lysates mixed in known ratios (PDF 19 kb)
Supplementary Table 3
Relex output file showing the normalized, average ion current ratios for 573 proteins identified and quantified from tandem mass spectra. Individual peptide ratio measurements are shown as well. (PDF 289 kb)
Rights and permissions
About this article
Cite this article
Venable, J., Dong, MQ., Wohlschlegel, J. et al. Automated approach for quantitative analysis of complex peptide mixtures from tandem mass spectra. Nat Methods 1, 39–45 (2004). https://doi.org/10.1038/nmeth705
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth705
This article is cited by
-
DeepFLR facilitates false localization rate control in phosphoproteomics
Nature Communications (2023)
-
Sampling the proteome by emerging single-molecule and mass spectrometry methods
Nature Methods (2023)
-
Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics
Nature Communications (2023)
-
Prediction of peptide mass spectral libraries with machine learning
Nature Biotechnology (2023)
-
Deep and fast label-free Dynamic Organellar Mapping
Nature Communications (2023)