Abstract
Huntington's disease (HD) is a fatal dominantly inherited neurodegenerative disorder caused by a CAG repeat expansion leading to an elongated polyglutamine stretch in huntingtin1. Mutant huntingtin (mHTT) is ubiquitously expressed in all cells but elicits selective cortical and striatal neurodegeneration in HD2. The mechanistic basis for such selective neuronal vulnerability remains unclear. A necessary step toward resolving this enigma is to define the cell types in which mHTT expression is causally linked to the disease pathogenesis. Using a conditional transgenic mouse model of HD, in which the mice express full-length human mHTT from a bacterial artificial chromosome transgene (BACHD)3, we genetically reduced mHTT expression in neuronal populations in the striatum, cortex or both. We show that reduction of cortical mHTT expression in BACHD mice partially improves motor and psychiatric-like behavioral deficits but does not improve neurodegeneration, whereas reduction of mHTT expression in both neuronal populations consistently ameliorates all behavioral deficits and selective brain atrophy in this HD model. Furthermore, whereas reduction of mHTT expression in cortical or striatal neurons partially ameliorates corticostriatal synaptic deficits, further restoration of striatal synaptic function can be achieved by reduction of mHTT expression in both neuronal cell types. Our study demonstrates distinct but interacting roles of cortical and striatal mHTT in HD pathogenesis and suggests that optimal HD therapeutics may require targeting mHTT in both cortical and striatal neurons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Vonsattel, J.P. & DiFiglia, M. Huntington disease. J. Neuropathol. Exp. Neurol. 57, 369–384 (1998).
Gray, M. et al. Full-length human mutant huntingtin with a stable polyglutamine repeat can elicit progressive and selective neuropathogenesis in BACHD mice. J. Neurosci. 28, 6182–6195 (2008).
Walker, F.O. Huntington's disease. Lancet 369, 218–228 (2007).
Tabrizi, S.J. et al. Potential endpoints for clinical trials in premanifest and early Huntington's disease in the TRACK-HD study: analysis of 24 month observational data. Lancet Neurol. 11, 42–53 (2012).
Kaltenbach, L.S. et al. Huntingtin interacting proteins are genetic modifiers of neurodegeneration. PLoS Genet. 11, e82 (2007).
Shirasaki, D.I. et al. Network organization of the huntingtin proteomic interactome in mammalian brain. Neuron 75, 41–57 (2012).
Ilieva, H., Polymenidou, M. & Cleveland, D.W. Non-cell autonomous toxicity in neurodegenerative disorders: ALS and beyond. J. Cell Biol. 187, 761–772 (2009).
Hadzi, T.C. et al. Assessment of cortical and striatal involvement in 523 Huntington disease brains. Neurology 79, 1708–1715 (2012).
Harper, S.Q. et al. RNA interference improves motor and neuropathological abnormalities in a Huntington's disease mouse model. Proc. Natl. Acad. Sci. USA 102, 5820–5825 (2005).
DiFiglia, M. et al. Therapeutic silencing of mutant huntingtin with siRNA attenuates striatal and cortical neuropathology and behavioral deficits. Proc. Natl. Acad. Sci. USA 104, 17204–17209 (2007).
Kordasiewicz, H.B. et al. Sustained therapeutic reversal of Huntington's disease by transient repression of huntingtin synthesis. Neuron 74, 1031–1044 (2012).
Sah, D.W. & Aronin, N. Oligonucleotide therapeutic approaches for Huntington disease. J. Clin. Invest. 121, 500–507 (2011).
Menalled, L. et al. Systematic behavioral evaluation of Huntington's disease transgenic and knock-in mouse models. Neurobiol. Dis. 35, 319–336 (2009).
Gu, X. et al. Pathological cell-cell interactions elicited by a neuropathogenic form of mutant huntingtin contribute to cortical pathogenesis in HD mice. Neuron 46, 433–444 (2005).
Dang, M.T. et al. Disrupted motor learning and long-term synaptic plasticity in mice lacking NMDAR1 in the striatum. Proc. Natl. Acad. Sci. USA 103, 15254–15259 (2006).
Gorski, J.A. et al. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 22, 6309–6314 (2002).
Iwasato, T. et al. Cortex-restricted disruption of NMDAR1 impairs neuronal patterns in the barrel cortex. Nature 406, 726–731 (2000).
Soriano, P. Generalized lacZ expression with the Rosa26 Cre reporter strain. Nat. Genet. 21, 70–71 (1999).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Schmidt, E.F. et al. Identification of the cortical neurons that mediate antidepressant responses. Cell 149, 1152–1163 (2012).
Matamales, M. et al. Striatal medium-sized spiny neurons: identification by nuclear staining and study of neuronal subpopulations in BAC transgenic mice. PLoS ONE 4, e4770 (2009).
Zoghbi, H.Y. & Warren, S.T. Neurogenetics: advancing the “next-generation” of brain research. Neuron 68, 165–173 (2010).
Raymond, L.A. et al. Pathophysiology of Huntington's disease: time-dependent alterations in synaptic and receptor function. Neuroscience 198, 252–273 (2011).
Goto, S. & Hirano, A. Synaptophysin expression in the striatum in Huntington's disease. ActaNeuropathol. 80, 88–91 (1990).
Cepeda, C. et al. Transient and progressive electrophysiological alterations in the corticostriatal pathway in a mouse model of Huntington's disease. J. Neurosci. 23, 961–969 (2003).
Kuhn, A. et al. Mutant huntingtin's effects on striatal gene expression in mice recapitulate changes observed in human Huntington's disease brain and do not differ with mutant huntingtin length or wild-type huntingtin dosage. Hum. Mol. Genet. 16, 1845–1861 (2007).
Becanovic, K. et al. Transcriptional changes in Huntington disease identified using genome-wide expression profiling and cross-platform analysis. Hum. Mol. Genet. 19, 1438–1452 (2010).
Wyszynski, M. et al. Competitive binding of α-actinin and calmodulin to the NMDA receptor. Nature 385, 439–442 (1997).
Dunah, A.W., Wyszynski, M., Martin, D.M., Sheng, M. & Standaert, D.G. α-actinin-2 in rat striatum: localization and interaction with NMDA glutamate receptor subunits. Brain Res. Mol. Brain Res. 79, 77–87 (2000).
Kennedy, M.B. The postsynaptic density at glutamatergic synapses. Trends Neurosci. 20, 264–268 (1997).
Fan, J., Cowan, C.M., Zhang, L.Y., Hayden, M.R. & Raymond, L.A. Interaction of postsynaptic density protein-95 with NMDA receptors influences excitotoxicity in the yeast artificial chromosome mouse model of Huntington's disease. J. Neurosci. 29, 10928–10938 (2009).
Crook, Z.R. & Housman, D. Huntington's disease: can mice lead the way to treatment? Neuron 69, 423–435 (2011).
Yang, X.W., Model, P. & Heintz, N. Homologous recombination based modification in Escherichia coli and germline transmission in transgenic mice of a bacterial artificial chromosome. Nat. Biotechnol. 15, 859–865 (1997).
Lobo, M.K., Karsten, S.L., Gray, M., Geschwind, D.H. & Yang, X.W. FACS-array profiling of striatal projection neuron subtypes in juvenile and adult mouse brains. Nat. Neurosci. 9, 443–452 (2006).
Gu, X. et al. Serines 13 and 16 are critical determinants of full-length human mutant huntingtin induced disease pathogenesis in HD mice. Neuron 64, 828–840 (2009).
Acknowledgements
X.W.Y. is supported by the US National Institutes of Health (NIH) NINDS grants R01 NS049501 and R01 NS074312, the Hereditary Disease Foundation, CHDI Foundation, Inc., Neuroscience of Brain Disorders Award from The McKnight Foundation and the David Weil Fund to the Semel Institute at University of California, Los Angeles (UCLA). X.W.Y. also receives support as the Carol Moss Spivak Scholar chair in Neuroscience from UCLA Brain Research Institute. M.S.L. is supported by NIH NINDS grant R01 NS41574. M.S.L. and C.C. are also supported by NIH grant P30 HD004612.
Author information
Authors and Affiliations
Contributions
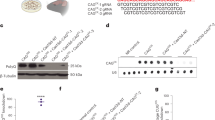
X.W.Y. provided the conceptual framework for the study, X.W.Y., N.W. and M.G. designed the experiments and discussed the results and X.W.Y. and N.W. wrote the manuscript. N.W. performed experiments shown in Figures 1b–f, 2a–g and 3a–d, Table 1, Supplementary Table 1 and Supplementary Figures 1–5. M.G. performed the experiment shown in Figure 1e, and many of the original behavioral and neuropathological studies evaluating the phenotypes of BE mice. X.-H.L. contributed to Figure 2h,i. E.G. and D.S. contributed to Figure 1e. X.G. and J.P.C. contributed to the generation of Emx1-Cre; Rgs9-Cre mice. C.C., S.M.H. and M.S.L. performed experiments shown in Figure 3e,f and Supplementary Figure 6 . H.D. helped in neuroanatomical and image analyses. Y.L. provided Rgs9-Cre mice and consulted on the use of the mice.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 and Supplementary Figures 1–6 (PDF 704 kb)
Rights and permissions
About this article
Cite this article
Wang, N., Gray, M., Lu, XH. et al. Neuronal targets for reducing mutant huntingtin expression to ameliorate disease in a mouse model of Huntington's disease. Nat Med 20, 536–541 (2014). https://doi.org/10.1038/nm.3514
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3514
This article is cited by
-
A clinical study and future prospects for bioactive compounds and semi-synthetic molecules in the therapies for Huntington's disease
Molecular Neurobiology (2024)
-
Microglia and complement mediate early corticostriatal synapse loss and cognitive dysfunction in Huntington’s disease
Nature Medicine (2023)
-
High-throughput virtual screening of potential inhibitors of GPR52 using docking and biased sampling method for Huntington’s disease therapy
Molecular Diversity (2023)
-
Huntington’s Disease: New Frontiers in Therapeutics
Current Neurology and Neuroscience Reports (2021)
-
Deficits in coordinated neuronal activity and network topology are striatal hallmarks in Huntington’s disease
BMC Biology (2020)