Abstract
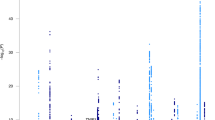
Eleven susceptibility loci for late-onset Alzheimer's disease (LOAD) were identified by previous studies; however, a large portion of the genetic risk for this disease remains unexplained. We conducted a large, two-stage meta-analysis of genome-wide association studies (GWAS) in individuals of European ancestry. In stage 1, we used genotyped and imputed data (7,055,881 SNPs) to perform meta-analysis on 4 previously published GWAS data sets consisting of 17,008 Alzheimer's disease cases and 37,154 controls. In stage 2, 11,632 SNPs were genotyped and tested for association in an independent set of 8,572 Alzheimer's disease cases and 11,312 controls. In addition to the APOE locus (encoding apolipoprotein E), 19 loci reached genome-wide significance (P < 5 × 10−8) in the combined stage 1 and stage 2 analysis, of which 11 are newly associated with Alzheimer's disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Corder, E.H. et al. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer's disease in late onset families. Science 261, 921–923 (1993).
Genin, E. et al. APOE and Alzheimer disease: a major gene with semi-dominant inheritance. Mol. Psychiatry 16, 903–907 (2011).
Lambert, J.-C. et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer's disease. Nat. Genet. 41, 1094–1099 (2009).
Harold, D. et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer's disease. Nat. Genet. 41, 1088–1093 (2009).
Seshadri, S. et al. Genome-wide analysis of genetic loci associated with Alzheimer disease. J. Am. Med. Assoc. 303, 1832–1840 (2010).
Hollingworth, P. et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer's disease. Nat. Genet. 43, 429–435 (2011).
Naj, A.C. et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer's disease. Nat. Genet. 43, 436–441 (2011).
Guerreiro, R. et al. TREM2 variants in Alzheimer's disease. N. Engl. J. Med. 368, 117–127 (2013).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer's disease. N. Engl. J. Med. 368, 107–116 (2013).
Miyashita, A. et al. SORL1 is genetically associated with late-onset Alzheimer's disease in Japanese, Koreans and Caucasians. PLoS ONE 8, e58618 (2013).
Cruchaga, C. et al. GWAS of cerebrospinal fluid tau levels identifies risk variants for Alzheimer's disease. Neuron 78, 256–268 (2013).
Sawcer, S. et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011).
Nalls, M.A. et al. Imputation of sequence variants for identification of genetic risks for Parkinson's disease: a meta-analysis of genome-wide association studies. Lancet 377, 641–649 (2011).
Rogaeva, E. et al. The neuronal sortilin-related receptor SORL1 is genetically associated with Alzheimer disease. Nat. Genet. 39, 168–177 (2007).
Pottier, C. et al. High frequency of potentially pathogenic SORL1 mutations in autosomal dominant early-onset Alzheimer disease. Mol. Psychiatry 17, 875–879 (2012).
Pandey, P. et al. Activation of p38 mitogen-activated protein kinase by PYK2/related adhesion focal tyrosine kinase–dependent mechanism. J. Biol. Chem. 274, 10140–10144 (1999).
Huang, Y. et al. CAKβ/Pyk2 kinase is a signaling link for induction of long-term potentiation in CA1 hippocampus. Neuron 29, 485–496 (2001).
Williamson, R. et al. CRMP2 hyperphosphorylation is characteristic of Alzheimer's disease and not a feature common to other neurodegenerative diseases. J. Alzheimers Dis. 27, 615–625 (2011).
Sulem, P. et al. Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat. Genet. 39, 1443–1452 (2007).
Adeyemo, A. et al. A genome-wide association study of hypertension and blood pressure in African Americans. PLoS Genet. 5, e1000564 (2009).
Larsson, M. et al. GWAS findings for human iris patterns: associations with variants in genes that influence normal neuronal pattern development. Am. J. Hum. Genet. 89, 334–343 (2011).
Kajiho, H. et al. RIN3: a novel Rab5 GEF interacting with amphiphysin II involved in the early endocytic pathway. J. Cell Sci. 116, 4159–4168 (2003).
Chapuis, J. et al. Increased expression of BIN1 mediates Alzheimer genetic risk by modulating tau pathology. Mol. Psychiatry doi:10.1038/mp.2013.1 (12 February 2013).
He, F. et al. Structural insight into the zinc finger CW domain as a histone modification reader. Structure 18, 1127–1139 (2010).
Yokoyama, K. et al. NYAP: a phosphoprotein family that links PI3K to WAVE1 signalling in neurons. EMBO J. 30, 4739–4754 (2011).
Gallo, J.M. & Spickett, C. The role of CELF proteins in neurological disorders. RNA Biol. 7, 474–479 (2010).
Del Villar, K. & Miller, C.A. Down-regulation of DENN/MADD, a TNF receptor binding protein, correlates with neuronal cell death in Alzheimer's disease brain and hippocampal neurons. Proc. Natl. Acad. Sci. USA 101, 4210–4215 (2004).
Duriez, B. et al. A common variant in combination with a nonsense mutation in a member of the thioredoxin family causes primary ciliary dyskinesia. Proc. Natl. Acad. Sci. USA 104, 3336–3341 (2007).
Pluskota, E. et al. The integrin coactivator kindlin-2 plays a critical role in angiogenesis in mice and zebrafish. Blood 117, 4978–4987 (2011).
Shulman, J.M. et al. Functional screening in Drosophila identifies Alzheimer's disease susceptibility genes and implicates Tau-mediated mechanisms. Hum. Mol. Genet. doi:10.1093/hmg/ddt478 (9 October 2013).
Kirsch, K.H., Georgescu, M.M., Ishimaru, S. & Hanafusa, H. CMS: an adapter molecule involved in cytoskeletal rearrangements. Proc. Natl. Acad. Sci. USA 96, 6211–6216 (1999).
Bao, M. et al. CD2AP/SHIP1 complex positively regulates plasmacytoid dendritic cell receptor signaling by inhibiting the E3 ubiquitin ligase Cbl. J. Immunol. 189, 786–792 (2012).
Brauer, H. et al. Leukemia-associated mutations in SHIP1 inhibit its enzymatic activity, interaction with the GM-CSF receptor and Grb2, and its ability to inactivate PI3K/AKT signaling. Cell. Signal. 24, 2095–2101 (2012).
Le Meur, N. et al. MEF2C haploinsufficiency caused by either microdeletion of the 5q14.3 region or mutation is responsible for severe mental retardation with stereotypic movements, epilepsy and/or cerebral malformations. J. Med. Genet. 47, 22–29 (2010).
Akhtar, M.W. et al. In vivo analysis of MEF2 transcription factors in synapse regulation and neuronal survival. PLoS ONE 7, e34863 (2012).
Griciuc, A. et al. Alzheimer's disease risk gene CD33 inhibits microglial uptake of amyloid β. Neuron 78, 631–643 (2013).
Lambert, J.C. & Amouyel, P. Genetics of Alzheimer's disease: new evidences for an old hypothesis? Curr. Opin. Genet. Dev. 21, 295–301 (2011).
Lambert, J.-C. et al. Implication of the immune system in Alzheimer's disease: evidence from genome-wide pathway analysis. J. Alzheimers Dis. 20, 1107–1118 (2010).
Jones, L. et al. Genetic evidence implicates the immune system and cholesterol metabolism in the aetiology of Alzheimer's disease. PLoS ONE 5, e13950 (2010).
Brouwers, N. et al. Alzheimer risk associated with a copy number variation in the complement receptor 1 increasing C3b/C4b binding sites. Mol. Psychiatry 17, 223–233 (2012).
Bettens, K., Sleegers, K. & Van Broeckhoven, C. Genetic insights in Alzheimer's disease. Lancet Neurol. 12, 92–104 (2013).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Purcell, S. et al. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am. J. Hum. Genet. 81, 559–575 (2007).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies via imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Aulchenko, Y.S., Struchalin, M.V. & van Duijn, C.M. ProbABEL package for genome-wide association analysis of imputed data. BMC Bioinformatics 11, 134 (2010).
Patterson, N., Price, A.L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Bellenguez, C. et al. A robust clustering algorithm for identifying problematic samples in genome-wide association studies. Bioinformatics 28, 134–135 (2012).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Mägi, R. & Morris, A.P. GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11, 288 (2010).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Levin, M.L. The occurrence of lung cancer in man. Acta Unio Int. Contra Cancrum 9, 531–541 (1953).
Acknowledgements
This work was made possible by the generous participation of the control subjects, the patients and their families. iSelect chips were funded by the French National Foundation on Alzheimer's Disease and Related Disorders. Data management involved the Centre National de Génotypage and was supported by the Institut Pasteur de Lille, INSERM, FRC (Fondation pour la Recherche sur le Cerveau) and Rotary. This work has been developed and supported by the LABEX (Laboratory of Excellence Program Investment for the Future) DISTALZ grant (Development of Innovative Strategies for a Transdisciplinary Approach to Alzheimer's Disease). The French National Foundation on Alzheimer's Disease and Related Disorders and the Alzheimer's Association (Chicago, Illinois) grant supported in-person meetings and communication for IGAP, and the Alzheimer's Association (Chicago, Illinois) grant provided some funds to each consortium for analyses.
GERAD was supported by the Wellcome Trust, the MRC, Alzheimer's Research UK (ARUK) and the Welsh government. ADGC and CHARGE were supported by the US National Institutes of Health, National Institute on Aging (NIH-NIA), including grants U01 AG032984 and R01 AG033193 (additional US National Institutes of Health grant numbers are listed in the Supplementary Note). CHARGE was also supported by Erasmus Medical Center and Erasmus University.
Complete acknowledgments are detailed in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
Study concept and design: J.-C.L., C.A.I.-V., D. Harold, A.C.N., A.L.D., J.C.B., A.V.S., M.A.I., H. Schmidt, A.L.F., V.G., O.L.L., D.W.T., D. Blacker, T.H.M., T.B.H., J.I.R., W.A.K., M. Boada, R. Schmidt, R.M., A.H., B.M.P., J.L.H., P.A.H., M.L., M.A.P.-V., L.J.L., L.A.F., C.M.v.D., V.M., S. Seshadri, J.W., G.D.S. and P.A. Acquisition of data: J.-C.L., C.A.I.-V., D. Harold, C. Bellenguez, R. Sims, G.J., B.G.-B., G.R., N.J., V.C., C. Thomas, D.Z., Y.K., A.G., H. Schmidt, M.L.D., M.-T.B., S.-H.C., P.H., V.G., C. Baldwin, C.C., C. Berr, O.L.L., P.L.D.J., D.E., L. Letenneur, G.E., K.S., A.M.G., N.F., M.J.H., M.I.K., E.B.L., A.J.M., C.D., S.T., S. Love, E.R., P.S.G.-H., L.Y., M.M.C., D. Beekly, F.Z., O.V., S.G.Y., W.G., M.J.O., K.M.F., P.V.J., M.C.O., L.B.C., D.A.B., T.B.H., R.F.A.G.d.B., T.J.M., J.I.R., K.M., T.M.F., W.A.K., J.F.P., M.A.N., K.R., J.S.K.K., E.B., M.R., M. Boada, L.-S.W., J.-F.D., C. Tzourio, M.M.N., B.M.P., L.J., J.L.H., M.L., L.J.L., L.A.F., A.H., C.M.v.D., S. Seshadri, J.W., G.D.S. and P.A. Sample contribution: A. Ruiz, F. Pasquier, A. Ramirez, O.H., J.D.B., D. Campion, P.K.C., C. Baldwin, T.B., C.C., D. Craig, V.D., J.A.J., S. Lovestone, F.J.M., D.C.R., K.S., A.M.G., N.F., M.G., K. Brown, M.I.K., L.K., P.B.-G., B.M., R.G., A.J.M., D.W., E.R., J.G., P.S.G.-H., J.C., A.L., A. Bayer, M.T., P. Bossù, G.S., P. Proitsi, J.C., S. Sorbi, F.S.-G., N.C.F., J.H., M.C.D.N., P. Bosco, R.C., C. Brayne, D.G., M. Mancuso, F.M., S. Moebus, P.M., M.D.Z., W.M., H. Hampel, A.P., M. Bullido, F. Panza, P.C., B.N., M. Mayhaus, L. Lannfelt, H. Hakonarson, S.P., M.M.C., M.I., V.A., S.G.Y., E.C., C. Razquin, P. Pastor, I.M., O.C., H. Soininen, S. Mead, D.A.B., L.F., C.H., P. Passmore, T.J.M., K. Bettens, A. Brice, D. Hannequin, K.R., M.R., M.H., D.R., C.G. and C.V.B. Data analysis: C.A.I.-V., D. Harold, A.C.N., R. Sims, C. Bellenguez, G.J., A.L.D., J.C.B., G.W.B., B.G.-B., G.R., T.A.T.-W., N.J., A.V.S., V.C., M.A.I., D.Z., Y.K., B.N.V., C.-F.L., A.G., B.K., C. Reitz, J.R.G., O.V., W.A.K., K.L.L., K.L.H.-N., E.R.M., L.-S.W., B.M.P., M.L., V.M. and J.W. Statistical analysis and interpretation: J.-C.L., C.A.I.-V., D. Harold, A.C.N., C. Bellenguez, G.J., A.L.D., J.C.B., G.W.B., T.A.T.-W., A.V.S., V.C., M.A.I., B.N.V., Y.K., C.-F.L., B.K., C. Reitz, A.L.F., N.A., J.R.G., R.F.A.G.d.B., W.A.K., K.L.L., E.R.M., L.-S.W., B.M.P., L.J., J.L.H., P.A.H., M.A.P.-V., L.J.L., L.A.F., C.M.v.D., V.M., S. Seshadri, J.W., G.D.S. and P.A. Drafting of the manuscript: J.-C.L., C.A.I.-V., D. Harold, A.C.N., C. Bellenguez, A.L.D., J.C.B., A.V.S., R.M., B.M.P., J.L.H., M.A.P.-V., L.J.L., L.A.F., C.M.v.D., C.V.B., S. Seshadri, J.W., G.D.S. and P.A.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Full lists of members and affiliations appear in the Supplementary Note
Full lists of members and affiliations appear in the Supplementary Note
Full lists of members and affiliations appear in the Supplementary Note
Full lists of members and affiliations appear in the Supplementary Note
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–18, Supplementary Tables 1–10 and Supplementary Note (PDF 3193 kb)
Rights and permissions
About this article
Cite this article
Lambert, JC., Ibrahim-Verbaas, C., Harold, D. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat Genet 45, 1452–1458 (2013). https://doi.org/10.1038/ng.2802
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2802
This article is cited by
-
iPSC-derived PSEN2 (N141I) astrocytes and microglia exhibit a primed inflammatory phenotype
Journal of Neuroinflammation (2024)
-
Unraveling the intercellular communication disruption and key pathways in Alzheimer’s disease: an integrative study of single-nucleus transcriptomes and genetic association
Alzheimer's Research & Therapy (2024)
-
Revealing third-order interactions through the integration of machine learning and entropy methods in genomic studies
BioData Mining (2024)
-
Regulation of human microglial gene expression and function via RNAase-H active antisense oligonucleotides in vivo in Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
Hotspots and trends of microglia in Alzheimer's disease: a bibliometric analysis during 2000–2022
European Journal of Medical Research (2024)