Abstract
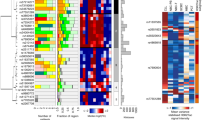
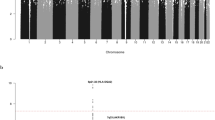
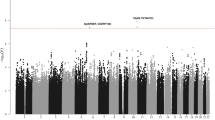
We conducted a genome-wide association study of 299,983 tagging SNPs for chronic lymphocytic leukemia (CLL) and performed validation in two additional series totaling 1,529 cases and 3,115 controls. We identified six previously unreported CLL risk loci at 2q13 (rs17483466; P = 2.36 × 10−10), 2q37.1 (rs13397985, SP140; P = 5.40 × 10−10), 6p25.3 (rs872071, IRF4; P = 1.91 × 10−20), 11q24.1 (rs735665; P = 3.78 × 10−12), 15q23 (rs7176508; P = 4.54 × 10−12) and 19q13.32 (rs11083846, PRKD2; P = 3.96 × 10−9). These data provide the first evidence for the existence of common, low-penetrance susceptibility to a hematological malignancy and new insights into disease causation in CLL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Sellick, G.S., Catovsky, D. & Houlston, R.S. Familial chronic lymphocytic leukemia. Semin. Oncol. 33, 195–201 (2006).
Raval, A. et al. Downregulation of death-associated protein kinase 1 (DAPK1) in chronic lymphocytic leukemia. Cell 129, 879–890 (2007).
Sellick, G.S. et al. A high-density SNP genome-wide linkage search of 206 families identifies susceptibility loci for chronic lymphocytic leukemia. Blood 110, 3326–3333 (2007).
Power, C. & Elliott, J. Cohort profile: 1958 British birth cohort (National Child Development Study). Int. J. Epidemiol. 35, 34–41 (2006).
Pfeifer, D. et al. Genome-wide analysis of DNA copy number changes and LOH in CLL using high-density SNP arrays. Blood 109, 1202–1210 (2007).
Busslinger, M. Transcriptional control of early B cell development. Annu. Rev. Immunol. 22, 55–79 (2004).
Klein, U. & Dalla-Favera, R. Germinal centres: role in B-cell physiology and malignancy. Nat. Rev. Immunol. 8, 22–33 (2008).
Shapiro-Shelef, M. & Calame, K. Regulation of plasma-cell development. Nat. Rev. Immunol. 5, 230–242 (2005).
Bloch, D.B., de la Monte, S.M., Guigaouri, P., Filippov, A. & Bloch, K.D. Identification and characterization of a leukocyte-specific component of the nuclear body. J. Biol. Chem. 271, 29198–29204 (1996).
Dent, A.L. et al. LYSP100-associated nuclear domains (LANDs): description of a new class of subnuclear structures and their relationship to PML nuclear bodies. Blood 88, 1423–1426 (1996).
Ling, P.D. et al. Mediation of Epstein-Barr virus EBNA-LP transcriptional coactivation by Sp100. EMBO J. 24, 3565–3575 (2005).
Madani, N. et al. Implication of the lymphocyte-specific nuclear body protein Sp140 in an innate response to human immunodeficiency virus type 1. J. Virol. 76, 11133–11138 (2002).
Oliver, P.M. et al. Loss of Bim allows precursor B cell survival but not precursor B cell differentiation in the absence of interleukin 7. J. Exp. Med. 200, 1179–1187 (2004).
Kovalevska, L.M. et al. Immunohistochemical studies of protein kinase D (PKD) 2 expression in malignant human lymphomas. Exp. Oncol. 28, 225–230 (2006).
Hamblin, T.J., Davis, Z., Gardiner, A., Oscier, D.G. & Stevenson, F.K. Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood 94, 1848–1854 (1999).
Damle, R.N. et al. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood 94, 1840–1847 (1999).
Weiss, N.S. Geographical variation in the incidence of the leukemias and lymphomas. Natl. Cancer Inst. Monogr. 53, 139–142 (1979).
Catovsky, D. et al. Assessment of fludarabine plus cyclophosphamide for patients with chronic lymphocytic leukaemia (the LRF CLL4 Trial): a randomised controlled trial. Lancet 370, 230–239 (2007).
Sayala, H. et al. Final report of the UKCLL02 trial: A phase II study of subcutaneous alemtuzumab plus fludarabine in patients with fludarabine refractory CLL (on behalf of the NCRI CLL trials sub-group). Blood (ASH Annual Meeting Abstracts) 108, 34 (2006).
Allan, J.M. et al. Polymorphism in glutathione S-transferase P1 is associated with susceptibility to chemotherapy-induced leukemia. Proc. Natl. Acad. Sci. USA 98, 11592–11597 (2001).
van Krieken, J.H. et al. Improved reliability of lymphoma diagnostics via PCR-based clonality testing: report of the BIOMED-2 Concerted Action BHM4–CT98–3936. Leukemia 21, 201–206 (2007).
Petitti, D. Meta-analysis Decision Analysis and Cost-Effectiveness Analysis (Oxford Univ. Press, New York, 1994).
Higgins, J.P. & Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
Cox, A. et al. A common coding variant in CASP8 is associated with breast cancer risk. Nat. Genet. 39, 352–358 (2007).
Goldin, L.R., Pfeiffer, R.M., Li, X. & Hemminki, K. Familial risk of lymphoproliferative tumors in families of patients with chronic lymphocytic leukemia: results from the Swedish Family-Cancer Database. Blood 104, 1850–1854 (2004).
Stranger, B.E. et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 1, e78 (2005).
Stranger, B.E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007).
Cuzick, J.A. Wilcoxon-type test for trend. Stat. Med. 4, 87–90 (1985).
Acknowledgements
Leukaemia Research Fund (LRF) provided principal funding for the study (LRF05001 and 06002). Additional funding was provided by CLL Global Research Foundation, Cancer Research UK (C1298/A8362 supported by the Bobby Moore Fund), the Arbib fund and the European Union (CPRB LSHC-CT-2004-503465). D.C.-S. was in receipt of a PhD studentship from the Institute of Cancer Research. We are grateful to M. Yuille for input into our early work on familial CLL. This study made use of genotyping data on the 1958 Birth Cohort. Genotyping data on controls was generated and generously supplied to us by P. Deloukas of the Wellcome Trust Sanger Institute. A full list of the investigators who contributed to the generation of the data are available from http://www.wtccc.org.uk. We are grateful to all investigators who contributed to the study of acute leukemia, from which phase 3 controls were drawn, to members of the ICLLLC (Supplementary Note online) and to investigators who contributed to NSCCG and GELCAPS, from which phase 1 and 2 controls were drawn. Finally, we are grateful to all the study subjects for their participation.
Author information
Authors and Affiliations
Contributions
R.S.H. designed the study with the assistance of G.S.; R.S.H. drafted the manuscript, with substantial contributions from E.W., M.C.D.B. and D.C.-S. M.C.D.B. performed statistical analyses with contributions from E.W. and Y.W. D.C.-S., M.C.D.B. and A.M.P. performed bioinformatics analyses. Phases 1 and 2: P.B. performed laboratory management and oversaw genotyping of cases and controls; D.C.-S., K.S. and J.V. conducted sequencing; D.C.-S. conducted genotyping of cases and controls; R.S.H., G.S., D.C., E.M., C.D. and D.C.-S. developed protocols for recruitment of individuals with CLL and sample acquisition and performed sample collection of cases; members of the ICLLLC consortium carried out ascertainment of familial cases (Supplementary Note). Phase 3: J.M.A. and D.J.A. conceived of the Newcastle-based CLL study. J.M.A. established the study, supervised laboratory management and oversaw genotyping of cases and controls. N.J.S. performed sample management of cases and controls. A.G.H. developed the Newcastle Haematology Biobank, incorporating the Newcastle-based CLL study. T.M.-F., G.H.J., G.S., R.J.H., A.R.P., P.H., D.J.A., J.R.B., G.P., C.P. and C.F. developed protocols for recruitment of individuals with CLL and sample acquisition and performed sample collection of cases. All authors contributed to the final paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–6, Supplementary Note (PDF 5158 kb)
Rights and permissions
About this article
Cite this article
Di Bernardo, M., Crowther-Swanepoel, D., Broderick, P. et al. A genome-wide association study identifies six susceptibility loci for chronic lymphocytic leukemia. Nat Genet 40, 1204–1210 (2008). https://doi.org/10.1038/ng.219
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.219
This article is cited by
-
PASTRY: achieving balanced power for detecting risk and protective minor alleles in meta-analysis of association studies with overlapping subjects
BMC Bioinformatics (2024)
-
Implementation of individualised polygenic risk score analysis: a test case of a family of four
BMC Medical Genomics (2022)
-
The dynamic functions of IRF4 in B cell malignancies
Clinical and Experimental Medicine (2022)
-
Distinct germline genetic susceptibility profiles identified for common non-Hodgkin lymphoma subtypes
Leukemia (2022)
-
IRF4 modulates the response to BCR activation in chronic lymphocytic leukemia regulating IKAROS and SYK
Leukemia (2021)