Abstract
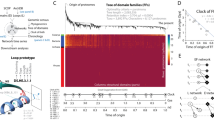
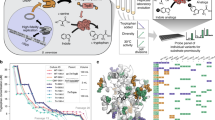
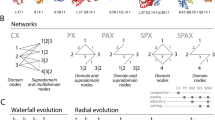
New protein folds have emerged throughout evolution, but it remains unclear how a protein fold can evolve while maintaining its function, particularly when fold changes require several sequential gene rearrangements. Here, we explored hypothetical evolutionary pathways linking different topological families of the DNA-methyltransferase superfamily. These pathways entail successive gene rearrangements through a series of intermediates, all of which should be sufficiently active to maintain the organism's fitness. By means of directed evolution, and starting from HaeIII methyltransferase (M.HaeIII), we selected all the required intermediates along these paths (a duplicated fused gene and duplicates partially truncated at their 5′ or 3′ coding regions) that maintained function in vivo. These intermediates led to new functional genes that resembled natural methyltransferases from three known classes or that belonged to a new class first seen in our evolution experiments and subsequently identified in natural genomes. Our findings show that new protein topologies can evolve gradually through multistep gene rearrangements and provide new insights regarding these processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Ponting, C.P. & Russell, R.B. Swaposins: circular permutations within genes encoding saposin homologues. Trends Biochem. Sci. 20, 179–180 (1995).
Grishin, N.V. Fold change in evolution of protein structures. J. Struct. Biol. 134, 167–185 (2001).
Koonin, E.V., Wolf, Y.I. & Karev, G.P. The structure of the protein universe and genome evolution. Nature 420, 218–223 (2002).
Aravind, L., Mazumder, R., Vasudevan, S. & Koonin, E.V. Trends in protein evolution inferred from sequence and structure analysis. Curr. Opin. Struct. Biol. 12, 392–399 (2002).
Graf, R. & Schachman, H.K. Random circular permutation of genes and expressed polypeptide chains: application of the method to the catalytic chains of aspartate transcarbamoylase. Proc. Natl. Acad. Sci. USA 93, 11591–11596 (1996).
Cunningham, B.A., Hemperly, J.J., Hopp, T.P. & Edelman, G.E. Favin versus concanavalin A: circularly permuted amino acid sequences. Proc. Natl. Acad. Sci. USA 76, 3218–3222 (1979).
Luger, K., Hommel, U., Herold, M., Hofsteenge, J. & Kirschner, K. Correct folding of circularly permuted variants of a beta alpha barrel enzyme in vivo. Science 243, 206–210 (1989).
Jeltsch, A. Circular permutations in the molecular evolution of DNA methyltransferases. J. Mol. Evol. 49, 161–164 (1999).
Bujnicki, J.M. Sequence permutations in the molecular evolution of DNA methyltransferases. BMC Evol. Biol. 2, 3 (2002).
Richardson, J.S. The anatomy and taxonomy of protein structure. Adv. Protein Chem. 34, 167–339 (1981).
Martin, J.L. & McMillan, F.M. SAM (dependent) I AM: the S-adenosylmethionine-dependent methyltransferase fold. Curr. Opin. Struct. Biol. 12, 783–793 (2002).
Malone, T., Blumenthal, R.M. & Cheng, X. Structure-guided analysis reveals nine sequence motifs conserved among DNA amino-methyltransferases, and suggests a catalytic mechanism for these enzymes. J. Mol. Biol. 253, 618–632 (1995).
Klimasauskas, S. et al. Sequence motifs characteristic of DNA[cytosine-N4]methyltransferases: similarity to adenine and cytosine-C5 DNA-methylases. Nucleic Acids Res. 17, 9823–9832 (1989).
Kita, K., Kotani, H., Sugisaki, H. & Takanami, M. The fokI restriction-modification system. I. Organization and nucleotide sequences of the restriction and modification genes. J. Biol. Chem. 264, 5751–5756 (1989).
Sugisaki, H., Kita, K. & Takanami, M. The FokI restriction-modification system. II. Presence of two domains in FokI methylase responsible for modification of different DNA strands. J. Biol. Chem. 264, 5757–5761 (1989).
Leismann, O., Roth, M., Friedrich, T., Wende, W. & Jeltsch, A. The Flavobacterium okeanokoites adenine-N6-specific DNA-methyltransferase M.FokI is a tandem enzyme of two independent domains with very different kinetic properties. Eur. J. Biochem. 251, 899–906 (1998).
Szomolanyi, E., Kiss, A. & Venetianer, P. Cloning the modification methylase gene of Bacillus sphaericus R in Escherichia coli. Gene 10, 219–225 (1980).
Ostermeier, M., Shim, J.H. & Benkovic, S.J. A combinatorial approach to hybrid enzymes independent of DNA homology. Nat. Biotechnol. 17, 1205–1209 (1999).
Choe, W., Chandrasegaran, S. & Ostermeier, M. Protein fragment complementation in M.HhaI DNA methyltransferase. Biochem. Biophys. Res. Commun. 334, 1233–1240 (2005).
Reinisch, K.M., Chen, L., Verdine, G.L. & Lipscomb, W.N. The crystal structure of HaeIII methyltransferase convalently complexed to DNA: an extrahelical cytosine and rearranged base pairing. Cell 82, 143–153 (1995).
Wagner, A. Robustness, evolvability, and neutrality. FEBS Lett. 579, 1772–1778 (2005).
Hennecke, J., Sebbel, P. & Glockshuber, R. Random circular permutation of DsbA reveals segments that are essential for protein folding and stability. J. Mol. Biol. 286, 1197–1215 (1999).
Iwakura, M., Nakamura, T., Yamane, C. & Maki, K. Systematic circular permutation of an entire protein reveals essential folding elements. Nat. Struct. Biol. 7, 580–585 (2000).
Chothia, C. The nature of the accessible and buried surfaces in proteins. J. Mol. Biol. 105, 1–12 (1976).
Jeltsch, A. Beyond Watson and Crick: DNA methylation and molecular enzymology of DNA methyltransferases. ChemBioChem 3, 274–293 (2002).
Kusano, K., Naito, T., Handa, N. & Kobayashi, I. Restriction-modification systems as genomic parasites in competition for specific sequences. Proc. Natl. Acad. Sci. USA 92, 11095–11099 (1995).
Roodveldt, C. & Tawfik, D.S. Directed evolution of phosphotriesterase from Pseudomonas diminuta for heterologous expression in Escherichia coli results in stabilization of the metal-free state. Protein Eng. Des. Sel. (2005).
Ostermeier, M., Nixon, A.E., Shim, J.H. & Benkovic, S.J. Combinatorial protein engineering by incremental truncation. Proc. Natl. Acad. Sci. USA 96, 3562–3567 (1999).
Tawfik, D.S. & Griffiths, A.D. Man-made cell-like compartments for molecular evolution. Nat. Biotechnol. 16, 652–656 (1998).
Fraczkiewicz, R. & Braun, W. Exact and efficient analytical calculation of the accessible surface areas and their gradients for macromolecules. J. Comp. Chem. 19, 319–333 (1998).
Acknowledgements
We thank the Minerva Foundation and the Israel Science Foundation for financial support, M. Babor for the software that generated the truncated PDB files and A. Levy for helpful comments on the manuscript. S.G.P. is the recipient of a Dewey D. Stone Postdoctoral Fellowship, L.R. is the recipient of a Feinberg Graduate School Fellowship and D.S.T. is the incumbent of the Elaine Blond Career Development Chair.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Fused Mtase dimers present in natural genomes. (PDF 143 kb)
Supplementary Fig. 2
Strategy used for the creation of truncated libraries using the ITCHY methodology. (PDF 11 kb)
Supplementary Fig. 3
In vivo methyltransferase activity of the disrupted N- and C-terminally truncated selected evolutionary intermediates. (PDF 77 kb)
Supplementary Fig. 4
Selection of Cluster I–derived circular permutants (PDF 39 kb)
Supplementary Fig. 5
In vivo methyltransferase activity of the disrupted Cluster I– and Cluster IV–derived selected circular permutants (PDF 48 kb)
Supplementary Fig. 6
Direct selection of circular permutants. (PDF 43 kb)
Supplementary Fig. 7
In vivo methyltransferase activity of the disrupted directly selected circular permutants. (PDF 62 kb)
Rights and permissions
About this article
Cite this article
Peisajovich, S., Rockah, L. & Tawfik, D. Evolution of new protein topologies through multistep gene rearrangements. Nat Genet 38, 168–174 (2006). https://doi.org/10.1038/ng1717
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1717
This article is cited by
-
A topology framework for macromolecular complexes and condensates
Nano Research (2022)
-
Synthetic evolution
Nature Biotechnology (2019)
-
Cotranslational protein assembly imposes evolutionary constraints on homomeric proteins
Nature Structural & Molecular Biology (2018)
-
Structure–function relationships of family GH70 glucansucrase and 4,6-α-glucanotransferase enzymes, and their evolutionary relationships with family GH13 enzymes
Cellular and Molecular Life Sciences (2016)
-
Restriction endonucleases: natural and directed evolution
Applied Microbiology and Biotechnology (2012)