Abstract
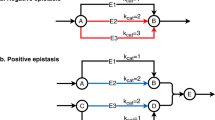
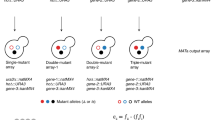
Epistatic interactions, manifested in the effects of mutations on the phenotypes caused by other mutations, may help uncover the functional organization of complex biological networks1,2,3. Here, we studied system-level epistatic interactions by computing growth phenotypes of all single and double knockouts of 890 metabolic genes in Saccharomyces cerevisiae, using the framework of flux balance analysis4. A new scale for epistasis identified a distinctive trimodal distribution of these epistatic effects, allowing gene pairs to be classified as buffering, aggravating or noninteracting2,5. We found that the ensuing epistatic interaction network6 could be organized hierarchically into function-enriched modules that interact with each other 'monochromatically' (i.e., with purely aggravating or purely buffering epistatic links). This property extends the concept of epistasis from single genes to functional units and provides a new definition of biological modularity, which emphasizes interactions between, rather than within, functional modules. Our approach can be used to infer functional gene modules from purely phenotypic epistasis measurements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tong, A.H.Y. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Hartman, J.L., Garvik, B. & Hartwell, L. Cell biology - Principles for the buffering of genetic variation. Science 291, 1001–1004 (2001).
Hartwell, L.H., Hopfield, J.J., Leibler, S. & Murray, A.W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
Famili, I., Forster, J., Nielsen, J. & Palsson, B.O. Saccharomyces cerevisiae phenotypes can be predicted by using constraint-based analysis of a genome-scale reconstructed metabolic network. Proc. Natl. Acad. Sci. USA 100, 13134–13139 (2003).
Phillips, P.C., Otto, S.P. & Whitlock, M.C. Beyond the Average: The Evolutionary Importance of Gene Interactions and Variablity of Epistatic Effects. in Epistasis and the Evolutionary Process (eds. Wolf, J.B., Brodie III, E.D. & Wade, M.J.) 20–38 (Oxford University Press, New York, 2000).
Anholt, R.R. et al. The genetic architecture of odor-guided behavior in Drosophila: epistasis and the transcriptome. Nat. Genet. 35, 180–184 (2003).
Ravasz, E., Somera, A.L., Mongru, D.A., Oltvai, Z.N. & Barabasi, A.L. Hierarchical organization of modularity in metabolic networks. Science 297, 1551–1555 (2002).
Almaas, E., Kovacs, B., Vicsek, T., Oltvai, Z.N. & Barabasi, A.L. Global organization of metabolic fluxes in the bacterium Escherichia coli. Nature 427, 839–843 (2004).
Papin, J.A., Reed, J.L. & Palsson, B.O. Hierarchical thinking in network biology: the unbiased modularization of biochemical networks. Trends Biochem. Sci. 29, 641–647 (2004).
Klamt, S. & Gilles, E.D. Minimal cut sets in biochemical reaction networks. Bioinformatics 20, 226–234 (2004).
Ihmels, J., Levy, R. & Barkai, N. Principles of transcriptional control in the metabolic network of Saccharomyces cerevisiae. Nat. Biotechnol. 22, 86–92 (2004).
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nat. Genet. 34, 166–176 (2003).
Kondrashov, A.S. Deleterious Mutations and the Evolution of Sexual Reproduction. Nature 336, 435–440 (1988).
Elena, S.F. & Lenski, R.E. Test of synergistic interactions among deleterious mutations in bacteria. Nature 390, 395–398 (1997).
Burch, C.L., Turner, P.E. & Hanley, K.A. Patterns of epistasis in RNA viruses: a review of the evidence from vaccine design. J. Evol. Biol. 16, 1223–1235 (2003).
Elena, S.F. Little evidence for synergism among deleterious mutations in a nonsegmented RNA virus. J. Mol. Evol. 49, 703–707 (1999).
Wloch, D.M., Borts, R.H. & Korona, R. Epistatic interactions of spontaneous mutations in haploid strains of the yeast Saccharomyces cerevisiae. J. Evol. Biol. 14, 310–316 (2001).
Lenski, R.E., Ofria, C., Collier, T.C. & Adami, C. Genome complexity, robustness and genetic interactions in digital organisms. Nature 400, 661–664 (1999).
You, L.C. & Yin, J. Dependence of epistasis on environment and mutation severity as revealed by in silico mutagenesis of phage T7. Genetics 160, 1273–1281 (2002).
Kishony, R. & Leibler, S. Environmental stresses can alleviate the average deleterious effect of mutations. J. Biol. 2, 14 (2003).
Remold, S.K. & Lenski, R.E. Pervasive joint influence of epistasis and plasticity on mutational effects in Escherichia coli. Nat. Genet. 36, 423–426 (2004).
Kauffman, K.J., Prakash, P. & Edwards, J.S. Advances in flux balance analysis. Curr. Opin. Biotechnol. 14, 491–496 (2003).
Forster, J., Famili, I., Fu, P., Palsson, B.O. & Nielsen, J. Genome-scale reconstruction of the Saccharomyces cerevisiae metabolic network. Genome Res. 13, 244–253 (2003).
Papp, B., Pal, C. & Hurst, L.D. Metabolic network analysis of the causes and evolution of enzyme dispensability in yeast. Nature 429, 661–664 (2004).
Fong, S.S. & Palsson, B.O. Metabolic gene-deletion strains of Escherichia coli evolve to computationally predicted growth phenotypes. Nat. Genet. 36, 1056–1058 (2004).
Dykhuizen, D.E., Dean, A.M. & Hartl, D.L. Metabolic flux and fitness. Genetics 115, 25–31 (1987).
Segrè, D., Vitkup, D. & Church, G.M. Analysis of optimality in natural and perturbed metabolic networks. Proc. Natl. Acad. Sci. USA 99, 15112–15117 (2002).
Milo, R. et al. Network motifs: simple building blocks of complex networks. Science 298, 824–827 (2002).
Kaufman, A., Kupiec, M. & Ruppin, E. Multi-knockout genetic network analysis: the Rad6 example. in 2004 IEEE Computational Systems Bioinformatics Conference 332–340 (IEEE, Stanford, California, 2004).
Salthe, S.N. Evolving Hierarchical Systems. (Columbia University Press, New York, 1985).
Acknowledgements
We thank L. Garwin, A. Murray, D. Fisher and D. Hartl for feedback and advice; A. Murray for the idea of complete buffering as indicative of biological modularity; Z. Kang for computational help; and J. Aach, U. Alon, N. Berger, A. Dudley, M. Elowitz, D. Fraenkel, G. Getz, M. Hegreness, P. Ma, B. Palsson D. Peer, O. Rando, A. Regev, U. Sauer, N. Shoresh, M. Turelli, D. Weinreich and M. Wright for comments. D.S. and G.C. acknowledge support from the Defense Advanced Research Projects Agency, the Genomes to Life program of the US Department of Energy and the Pharmaceutical Research and Manufacturers of America Foundation. R.K. acknowledges support from the Bauer Center for Genomics Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
The epistasis scale captures the strength of the epistasis effect relative to natural extreme values. (PDF 4 kb)
Supplementary Fig. 2
Analysis of the effects of double deletions of metabolic enzyme genes in simple metabolic networks demonstrating examples of multiplicative, aggravating and buffering gene deletion interactions. (PDF 27 kb)
Supplementary Fig. 3
Sensitivity analysis with respect to free parameters and physiological conditions. (PDF 3 kb)
Supplementary Fig. 4
Sub-classification of buffering interactions into directional and non-directional links is related to the underlying biochemical network. (PDF 6 kb)
Supplementary Fig. 5
Changes in monochromatically interacting epistasis modules following variation of oxygen uptake rate. (PDF 7 kb)
Supplementary Fig. 6
Examples of monochromatic clustering of three toy networks using the Prism algorithm. (PDF 11 kb)
Supplementary Fig. 7
Randomization algorithms and statistical tests for monochromaticity in the epistasis network. (PDF 7 kb)
Supplementary Table 1
List of free parameters. (PDF 179 kb)
Supplementary Methods
The yeast flux balance model. (PDF 32 kb)
Supplementary Note
Discussion of Prism modules and predicted interactions. (PDF 46 kb)
Supplementary Video 1
Schematic demonstration of monochromatic classification. A network of two types of connections, such as buffering (green) and aggravating (red) epistasis, is sorted into module of genes that interact with one another in a purely monochromatic way. (AVI 27 kb)
Rights and permissions
About this article
Cite this article
Segrè, D., DeLuna, A., Church, G. et al. Modular epistasis in yeast metabolism. Nat Genet 37, 77–83 (2005). https://doi.org/10.1038/ng1489
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1489
This article is cited by
-
Scalable genetic screening for regulatory circuits using compressed Perturb-seq
Nature Biotechnology (2023)
-
Hsp90 inhibition sensitizes DLBCL cells to cisplatin
Cancer Chemotherapy and Pharmacology (2022)
-
Non-additive microbial community responses to environmental complexity
Nature Communications (2021)
-
Mechanisms of drug interactions between translation-inhibiting antibiotics
Nature Communications (2020)
-
Interaction between MyD88, TIRAP and IL1RL1 against Helicobacter pylori infection
Scientific Reports (2020)