Abstract
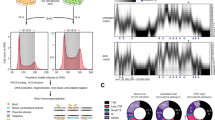
Eukaryotic chromosomes are packaged in nuclei by many orders of folding. Little is known about how higher-order chromatin packaging might affect gene expression. SATB1 is a cell-type specific nuclear protein that recruits chromatin-remodeling factors and regulates numerous genes during thymocyte differentiation. Here we show that in thymocyte nuclei, SATB1 has a cage-like 'network' distribution circumscribing heterochromatin and selectively tethers specialized DNA sequences onto its network. This was shown by fluorescence in situ hybridization on wild-type and Satb1-null thymocytes using in vivo SATB1-bound sequences as probes. Many gene loci, including that of Myc and a brain-specific gene, are anchored by the SATB1 network at specific genomic sites, and this phenomenon is precisely correlated with proper regulation of distant genes. Histone-modification analyses across a gene-enriched genomic region of 70 kb showed that acetylation of histone H3 at Lys9 and Lys14 peaks at the SATB1-binding site and extends over a region of roughly 10 kb covering genes regulated by SATB1. By contrast, in Satb1-null thymocytes, this site is marked by methylation at H3 Lys9. We propose SATB1 as a new type of gene regulator with a novel nuclear architecture, providing sites for tissue-specific organization of DNA sequences and regulating region-specific histone modification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Roth, S.Y., Denu, J.M. & Allis, C.D. Histone acetyltransferases. Annu. Rev. Biochem. 70, 81–120 (2001).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Kouzarides, T. Histone methylation in transcriptional control. Curr. Opin. Genet. Dev. 12, 198–209 (2002).
Varga-Weisz, P. ATP-dependent chromatin remodeling factors: nucleosome shufflers with many missions. Oncogene 20, 3076–3085 (2001).
Kingston, R.E. & Narlikar, G.J. ATP-dependent remodeling and acetylation as regulators of chromatin fluidity. Genes Dev. 13, 2339–2352 (1999).
Fry, C.J. & Peterson, C.L. Chromatin remodeling enzymes: who's on first? Curr. Biol. 11, R185–R197 (2001).
Chan, H.M. & La Thangue, N.B. p300/CBP proteins: HATs for transcriptional bridges and scaffolds. J. Cell Sci. 114, 2363–2373 (2001).
Bulger, M., Sawado, T., Schubeler, D. & Groudine, M. ChIPs of the β-globin locus: unraveling gene regulation within an active domain. Curr. Opin. Genet. Dev. 12, 170–177 (2002).
Noma, K., Allis, C.D. & Grewal, S.I. Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science 293, 1150–1155 (2001).
Schubeler, D. et al. Nuclear localization and histone acetylation: a pathway for chromatin opening and transcriptional activation of the human β-globin locus. Genes Dev. 14, 940–950 (2000).
Litt, M.D., Simpson, M., Recillas-Targa, F., Prioleau, M.N. & Felsenfeld, G. Transitions in histone acetylation reveal boundaries of three separately regulated neighboring loci. EMBO J. 20, 2224–2235 (2001).
Litt, M.D., Simpson, M., Gaszner, M., Allis, C.D. & Felsenfeld, G. Correlation between histone lysine methylation and developmental changes at the chicken β-globin locus. Science 293, 2453–2455 (2001).
Francastel, C., Schubeler, D., Martin, D.I. & Groudine, M. Nuclear compartmentalization and gene activity. Nat. Rev. Mol. Cell Biol. 1, 137–143 (2000).
Dickinson, L.A., Joh, T., Kohwi, Y. & Kohwi-Shigematsu, T. A tissue-specific MAR/SAR DNA-binding protein with unusual binding site recognition. Cell 70, 631–645 (1992).
Kohwi-Shigematsu, T. & Kohwi, Y. Torsional stress stabilizes extended base unpairing in suppressor sites flanking immunoglobulin heavy chain enhancer. Biochemistry 29, 9551–9560 (1990).
Bode, J. et al. Biological significance of unwinding capability of nuclear matrix-associating DNAs. Science 255, 195–197 (1992).
Cockerill, P.N., Yuen, M.H. & Garrard, W.T. The enhancer of the immunoglobulin heavy chain locus is flanked by presumptive chromosomal loop anchorage elements. J. Biol. Chem. 262, 5394–5397 (1987).
Forrester, W.C., van Genderen, C., Jenuwein, T. & Grosschedl, R. Dependence of enhancer-mediated transcription of the immunoglobulin mu gene on nuclear matrix attachment regions. Science 265, 1221–1225 (1994).
Jenuwein, T. et al. Extension of chromatin accessibility by nuclear matrix attachment regions. Nature 385, 269–272 (1997).
Lichtenstein, M., Keini, G., Cedar, H. & Bergman, Y. B cell-specific demethylation: a novel role for the intronic kappa chain enhancer sequence. Cell 76, 913–923 (1994).
Kirillov, A. et al. A role for nuclear NFκB in B-cell-specific demethylation of the Igκ locus. Nat. Genet. 13, 435–441 (1996).
Fernandez, L.A., Winkler, M. & Grosschedl, R. Matrix attachment region–dependent function of the immunoglobulin mu enhancer involves histone acetylation at a distance without changes in enhancer occupancy. Mol. Cell. Biol. 21, 196–208 (2001).
Nakagomi, K., Kohwi, Y., Dickinson, L.A. & Kohwi-Shigematsu, T. A novel DNA-binding motif in the nuclear matrix attachment DNA-binding protein SATB1. Mol. Cell. Biol. 14, 1852–1860 (1994).
Dickinson, L.A., Dickinson, C.D. & Kohwi-Shigematsu, T. An atypical homeodomain in SATB1 promotes specific recognition of the key structural element in a matrix attachment region. J. Biol. Chem. 272, 11463–11470 (1997).
Galande, S., Dickinson, L.A., Mian, I.S., Sikorska, M. & Kohwi-Shigematsu, T. SATB1 cleavage by caspase 6 disrupts PDZ domain–mediated dimerization, causing detachment from chromatin early in T-cell apoptosis. Mol. Cell. Biol. 21, 5591–5604 (2001).
Hawkins, S.M., Kohwi-Shigematsu, T. & Skalnik, D.G. The matrix attachment region–binding protein SATB1 interacts with multiple elements within the gp91phox promoter and is down-regulated during myeloid differentiation. J. Biol. Chem. 276, 44472–44480 (2001).
Alvarez, J.D. et al. The MAR-binding protein SATB1 orchestrates temporal and spatial expression of multiple genes during T-cell development. Genes Dev. 14, 521–535 (2000).
Dickinson, L.A. & Kohwi-Shigematsu, T. Nucleolin is a matrix attachment region DNA-binding protein that specifically recognizes a region with high base-unpairing potential. Mol. Cell. Biol. 15, 456–465 (1995).
Galande, S. & Kohwi-Shigematsu, T. Poly(ADP-ribose) polymerase and Ku autoantigen form a complex and synergistically bind to matrix attachment sequences. J. Biol. Chem. 274, 20521–20528 (1999).
Liu, W.M., Guerra-Vladusic, F.K., Kurakata, S., Lupu, R. & Kohwi-Shigematsu, T. HMG-I(Y) recognizes base-unpairing regions of matrix attachment sequences and its increased expression is directly linked to metastatic breast cancer phenotype. Cancer Res. 59, 5695–5703 (1999).
Herrscher, R.F. et al. The immunoglobulin heavy-chain matrix-associating regions are bound by Bright: a B cell-specific trans-activator that describes a new DNA-binding protein family. Genes Dev. 9, 3067–3082 (1995).
de Belle, I., Cai, S. & Kohwi-Shigematsu, T. The genomic sequences bound to special AT-rich sequence-binding protein 1 (SATB1) in vivo in Jurkat T cells are tightly associated with the nuclear matrix at the bases of the chromatin loops. J. Cell Biol. 141, 335–348 (1998).
Yasui, D., Miyano, M., Cai, S., Varga-Weisz, P. & Kohwi-Shigematsu, T. SATB1 targets chromatin remodelling to regulate genes over long distances. Nature 419, 641–645 (2002).
Kohwi-Shigematsu, T. & Kohwi, Y. Detection of non-B-DNA structures at specific sites in supercoiled plasmid DNA and chromatin with haloacetaldehyde and diethyl pyrocarbonate. Methods Enzymol. 212, 155–180 (1992).
Kohwi-Shigematsu, T., deBelle, I., Dickinson, L.A., Galande, S. & Kohwi, Y. Identification of base-unpairing region-binding proteins and characterization of their in vivo binding sequences. Methods Cell Biol. 53, 323–354 (1998).
Cai, S. & Kohwi-Shigematsu, T. Intranuclear relocalization of matrix binding sites during T cell activation detected by amplified fluorescence in situ hybridization. Methods 19, 394–402 (1999).
Riegel, J.S., Richie, E.R. & Allison, J.P. Nuclear events after activation of CD4+8+ thymocytes. J. Immunol. 144, 3611–3618 (1990).
Eisenman, R.N. Deconstructing myc. Genes Dev. 15, 2023–2030 (2001).
Douglas, N.C., Jacobs, H., Bothwell, A.L. & Hayday, A.C. Defining the specific physiological requirements for c-Myc in T cell development. Nat. Immunol. 2, 307–315 (2001).
Sanders, J., Maassen, J.A. & Moller, W. Elongation factor-1 messenger-RNA levels in cultured cells are high compared to tissue and are not drastically affected further by oncogenic transformation. Nucleic Acids Res. 20, 5907–5910 (1992).
Kohwi-Shigematsu, T., Maass, K. & Bode, J. A thymocyte factor SATB1 suppresses transcription of stably integrated matrix-attachment region-linked reporter genes. Biochemistry 36, 12005–12010 (1997).
Liu, J. et al. The matrix attachment region–binding protein SATB1 participates in negative regulation of tissue-specific gene expression. Mol. Cell. Biol. 17, 5275–5287 (1997).
Bell, A.C., West, A.G. & Felsenfeld, G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98, 387–396 (1999).
Brown, K.E. et al. Association of transcriptionally silent genes with Ikaros complexes at centromeric heterochromatin. Cell 91, 845–854 (1997).
Gerasimova, T.I., Byrd, K. & Corces, V.G. A chromatin insulator determines the nuclear localization of DNA. Mol. Cell 6, 1025–1035 (2000).
Duncan, R. et al. A sequence-specific, single-strand binding protein activates the far upstream element of c-myc and defines a new DNA-binding motif. Genes Dev. 8, 465–480 (1994).
He, L. et al. Loss of FBP function arrests cellular proliferation and extinguishes c-myc expression. EMBO J. 19, 1034–1044 (2000).
Kohwi, Y. & Panchenko, Y. Transcription-dependent recombination induced by triple-helix formation. Genes Dev. 7, 1766–1778 (1993).
Nielsen, S.J. et al. Rb targets histone H3 methylation and HP1 to promoters. Nature 412, 561–565 (2001).
Wreggett, K.A. et al. A mammalian homologue of Drosophila heterochromatin protein 1 (HP1) is a component of constitutive heterochromatin. Cytogenet. Cell Genet. 66, 99–103 (1994).
Acknowledgements
We thank A. Dernburg for help in obtaining the DeltaVision image, P. Varga-Weisz for valuable discussion and J.-F. Cheng and M. Kohwi for critically reading the manuscript. This work was supported by US National Institutes of Health grants to T.K.-S.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Cai, S., Han, HJ. & Kohwi-Shigematsu, T. Tissue-specific nuclear architecture and gene expession regulated by SATB1. Nat Genet 34, 42–51 (2003). https://doi.org/10.1038/ng1146
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1146
This article is cited by
-
A dual function for the chromatin organizer Special A-T rich Binding Protein 1 in B-lineage cells
Cellular & Molecular Immunology (2023)
-
MicroRNA-495: a therapeutic and diagnostic tumor marker
Journal of Molecular Histology (2023)
-
Expression of SATB1 and SATB2 in the brain of bony fishes: what fish reveal about evolution
Brain Structure and Function (2023)
-
Evaluation of 41 single nucleotide polymorphisms in canine diffuse large B-cell lymphomas using MassARRAY
Scientific Reports (2022)
-
Global chromatin organizer SATB1 acts as a context-dependent regulator of the Wnt/Wg target genes
Scientific Reports (2021)