Abstract
Map-based positional cloning of Drosophila melanogaster genes is hampered by both the time-consuming, error-prone nature of traditional methods for genetic mapping and the difficulties in aligning the genetic and cytological maps with the genome sequence. The identification of sequence polymorphisms in the Drosophila genome will make it possible to map mutations directly to the genome sequence with high accuracy and resolution. Here we report the identification of 7,223 single-nucleotide polymorphisms (SNPs) and 1,392 insertions/deletions (InDels) in common laboratory strains of Drosophila. These sequence polymorphisms define a map of 787 autosomal marker loci with a resolution of 114 kb. We have established PCR product–length polymorphism (PLP) or restriction fragment–length polymorphism (RFLP) assays for 215 of these markers. We demonstrate the use of this map by delimiting two mutations to intervals of 169 kb and 307 kb, respectively. Using a local high-density SNP map, we also mapped a third mutation to a resolution of approximately 2 kb, sufficient to localize the mutation within a single gene. These methods should accelerate the rate of positional cloning in Drosophila.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Collins, F.S., Guyer, M.S. & Charkravarti, A. Variations on a theme: cataloging human DNA sequence variation. Science 278, 1580–1581 (1997).
Winzeler, E.A. et al. Direct allelic variation scanning of the yeast genome. Science 281, 1194–1197 (1998).
Cho, R.J. et al. Genome-wide mapping with biallelic markers in Arabidopsis thaliana. Nature Genet. 23, 203–207 (1999).
Wicks, S.R., Yeh, R.T., Gish, W.R., Waterston, R.H. & Plasterk, R.H. Rapid gene mapping in Caenorhabditis elegans using a high density polymorphism map. Nature Genet. 28, 160–164 (2001).
Hoskins, R.A. et al. Single-nucleotide polymorphism markers for genetic mapping in Drosophila melanogaster. Genome Res. 11, 1100–1113 (2001).
Lindblad-Toh, K. et al. Large-scale discovery and genotyping of single-nucleotide polymorphisms in the mouse. Nature Genet. 24, 381–386 (2000).
Kimmerly, W. et al. A P1-based physical map of the Drosophila euchromatic genome. Genome Res. 6, 414–430 (1996).
Teeter, K. et al. Haplotype dimorphism in a SNP collection from Drosophila melanogaster. J. Exp. Zool. 288, 63–75 (2000).
Xu, T. & Rubin, G.M. Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117, 1223–1237 (1993).
Rørth, P. et al. Systematic gain-of-function genetics in Drosophila. Development 125, 1049–1057 (1998).
Xu, T., Wang, W., Zhang, S., Stewart, R.A. & Yu, W. Identifying tumor suppressors in genetic mosaics: the Drosophila lats gene encodes a putative protein kinase. Development 121, 1053–1063 (1995).
Liu, Y. & Montell, D.J. Identification of mutations that cause cell migration defects in mosaic clones. Development 126, 1869–1878 (1999).
Newsome, T.P., Asling, B. & Dickson, B.J. Analysis of Drosophila photoreceptor axon guidance in eye-specific mosaics. Development 127, 851–860 (2000).
Benlali, A., Draskovic, I., Hazelett, D.J. & Treisman, J.E. act up controls actin polymerization to alter cell shape and restrict Hedgehog signaling in the Drosophila eye disc. Cell 101, 271–281 (2000).
Pichaud, F. & Desplan, C. A new visualization approach for identifying mutations that affect differentiation and organization of the Drosophila ommatidia. Development 128, 815–826 (2001).
Marth, G.T. et al. A general approach to single-nucleotide polymorphism discovery. Nature Genet. 23, 452–456 (1999).
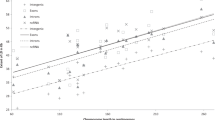
Begun, D.J. & Aquadro, C.F. Levels of naturally occurring DNA polymorphism correlate with recombination rates in D. melanogaster. Nature 356, 519–520 (1992).
Ruan, W., Pang, P. & Rao, Y. The SH2/SH3 adaptor protein dock interacts with the Ste20-like kinase misshapen in controlling growth cone motility. Neuron 24, 595–605 (1999).
Kwok, P.-Y. Methods for genotyping single nucleotide polymorphisms. Annu. Rev. Genomics Hum. Genet. 2, 235–258 (2001).
Adams, M.D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Ewing, B., Hillier, L., Wendl, M.C. & Green, P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 8, 175–185 (1998).
Ewing, B. & Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194 (1998).
Petrov, D.A. & Hartl, D.L. Patterns of nucleotide substitution in Drosophila and mammalian genomes. Proc. Natl Acad. Sci. USA. 96, 1475–1479 (1999).
Acknowledgements
We thank F. Eisenhaber and S. Maurer-Stroh for providing assistance in bioinformatics, A. Graf for technical assistance and M. Hohl and I. Botto for DNA sequencing. We are also grateful to the Berkeley Drosophila Genome Project, Celera Genomics and FlyBase for providing the wealth of resources that made this project possible. This work was funded by Boehringer Ingelheim GmbH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Berger, J., Suzuki, T., Senti, KA. et al. Genetic mapping with SNP markers in Drosophila. Nat Genet 29, 475–481 (2001). https://doi.org/10.1038/ng773
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng773
This article is cited by
-
Genetic background mutations drive neural circuit hyperconnectivity in a fragile X syndrome model
BMC Biology (2020)
-
Coding single nucleotide polymorphisms and SmCPS1 and SmKSL1 subcellular localization are associated with tanshinone biosynthesis in Salvia miltiorrhiza Bunge roots
Acta Physiologiae Plantarum (2018)
-
Genome editing in Drosophila melanogaster: from basic genome engineering to the multipurpose CRISPR-Cas9 system
Science China Life Sciences (2017)
-
MiSNPDb: a web-based genomic resources of tropical ecology fruit mango (Mangifera indica L.) for phylogeography and varietal differentiation
Scientific Reports (2017)
-
Drosophila Strip serves as a platform for early endosome organization during axon elongation
Nature Communications (2014)