Abstract
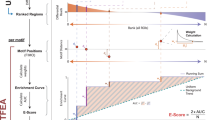
We present here a new computational method for discovering cis-regulatory elements that circumvents the need to cluster genes based on their expression profiles. Based on a model in which upstream motifs contribute additively to the log-expression level of a gene, this method requires a single genome-wide set of expression ratios and the upstream sequence for each gene, and outputs statistically significant motifs. Analysis of publicly available expression data for Saccharomyces cerevisiae reveals several new putative regulatory elements, some of which plausibly control the early, transient induction of genes during sporulation. Known motifs generally have high statistical significance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Cherry, J.M. et al. Genetic and physical maps of Saccharomyces cerevisiae. Nature 387, 67–73 (1997).
Schena, M., Shalon, D., Davis, R.W. & Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Lockhart, D.J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nature Biotechnol. 14, 1675–1680 (1996).
Velculescu, V.E., Zhang, L., Vogelstein, B. & Kinzler, K.W. Serial analysis of gene expression. Science 270, 484–487 (1995).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Roth, F.R., Hughes, J.D., Estep, P.W. & Church, G.M. Finding DNA regulatory motifs within unaligned noncoding sequences clustered by whole-genome mRNA quantitation. Nature Biotechnol. 16, 939–945 (1998).
Lawrence, C.E. et al. Detecting subtle sequence signals: a Gibbs sampling strategy for multiple alignment. Science 262, 208–214 (1993).
Neuwald, A.F., Liu, J.S. & Lawrence, C.E. Gibbs motif sampling: detection of bacterial outer membrane protein repeats. Protein Sci. 4, 1618–1632 (1995).
Van Helden, J., Andre, B. & Collado-Vides, J. Extracting regulatory sites from the upstream region of yeast genes by computational analysis of oligonucleotide frequencies. J. Mol. Biol. 281, 827–842 (1998).
DeRisi, J.L., Iyer, V.R. & Brown, P.O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 287, 680–686 (1997).
Chu, S. et al. The transcriptional program of sporulation in budding yeast. Science 282, 699–705 (1998).
Cho, R.J. et al. A genome-wide transcriptional analysis of the mitotic cell cycle. Mol. Cell 2, 65–73 (1998).
Spellman, P.T. et al. Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol. Biol. Cell. 9, 3273–3297 (1998).
Tavazoie, S., Hughes, J.D., Campbell, M.J., Cho, R.J. & Church, G.M. Systematic determination of genetic network architecture. Nature Genet. 22, 281–285 (1999).
Berg, O.G. & Von Hippel, P.H. Selection of DNA binding sites by regulatory proteins. Statistical-mechanical theory and application to operators and promoters. J. Mol. Biol. 193, 723–750 (1987).
Magasanik, B. Regulation of nitrogen utilisation. in The Molecular and Cellular Biology of the Yeast Saccharomyces: Gene Expression (eds. Jones, E.W., Pringle, J.R. & Broach, J.R.) 283–318 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1992).
Niehrs, C. & Pollet, N. Synexpression groups in eukaryotes. Nature 402, 483–487 (1999).
Ptashne, M. & Gann, A. Imposing specificity by localization: mechanism and evolvability. Curr. Biol. 8, R897 (1998).
Holstege, F.C.P. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998).
Halfter, H., Kavety, B., Vandekerckhove, J., Kiefer, F. & Gallwitz, D. Sequence, expression and mutational analysis of BAF1, a transcriptional activator and ARS1-binding protein of the yeast Saccharomyces cerevisiae. EMBO J. 8, 4265–4272 (1989).
Della Seta, F. et al. The ABF1 factor is the transcriptional activator of the L2 ribosomal 15 protein genes in Saccharomyces cerevisiae. Mol. Cell. Biol. 10, 2437–2441 (1990).
Loots, G.G. et al. Identification of a coordinate regulator of interleukins 4, 13, and 5 by cross-species sequence comparisons. Science 288, 136–140 (2000).
Hardison, R.C., Oeltjen, J. & Miller, W. Long human-mouse sequence alignments reveal novel regulatory elements: a reason to sequence the mouse genome. Genome Res. 7, 959–966 (1997).
Ben-Dor, A., Shamir, R. & Yakhini, Z. Clustering gene expression patterns. J. Comput. Biol. 6, 281–297 (1999).
Acknowledgements
We thank B. Shraiman for suggesting linear multivariate fits to expression data, and L. Grivell, R. Lascaris and H. de Nobel for discussions and critical reading of the manuscript. Support was received from the NSF under grant number DMR 9732083 and from the Keck foundation to H.L.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bussemaker, H., Li, H. & Siggia, E. Regulatory element detection using correlation with expression. Nat Genet 27, 167–171 (2001). https://doi.org/10.1038/84792
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/84792
This article is cited by
-
DeepGAMI: deep biologically guided auxiliary learning for multimodal integration and imputation to improve genotype–phenotype prediction
Genome Medicine (2023)
-
Single-cell sortChIC identifies hierarchical chromatin dynamics during hematopoiesis
Nature Genetics (2023)
-
Bayesian mixture regression analysis for regulation of Pluripotency in ES cells
BMC Bioinformatics (2020)
-
Deciphering eukaryotic gene-regulatory logic with 100 million random promoters
Nature Biotechnology (2020)
-
Insights about collective decision-making at the genetic level
Biophysical Reviews (2020)