Abstract
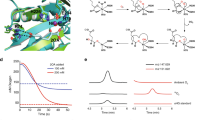
Although the latter portion of lysine biosynthesis, the conversion of α-aminoadipate (AAA) to lysine, in Thermus thermophilus is similar to the latter portion of arginine biosynthesis, enzymes homologous to ArgA and ArgJ are absent from the lysine pathway. Because ArgA and ArgJ are known to modify the amino group of glutamate to avoid intramolecular cyclization of intermediates, their absence suggests that the pathway includes an alternative N-modification system. We reconstituted the conversion of AAA to lysine and found that the amino group of AAA is modified by attachment to the γ-carboxyl group of the C-terminal Glu54 of a small protein, LysW; that the side chain of AAA is converted to the lysyl side chain while still attached to LysW; and that lysine is subsequently liberated from the LysW-lysine fusion. The fact that biosynthetic enzymes recognize the acidic globular domain of LysW indicates that LysW acts as a carrier protein or protein scaffold for the biosynthetic enzymes. This study thus reveals the previously unknown function of a small protein in primary metabolism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Broquist, H.P. Lysine biosynthesis (yeast). Methods Enzymol. 17, 112–129 (1971).
Buchli, R. et al. Cloning and functional expression of a soluble form of kynurenine/α-aminoadipate aminotransferase from rat kidney. J. Biol. Chem. 270, 29330–29335 (1995).
Vogel, H.J. Distribution of lysine pathway among fungi: evolutionary implications. Am. Nat. 98, 446–455 (1964).
Kobashi, N., Nishiyama, M. & Tanokura, M. Aspartate kinase-independent lysine synthesis in an extremely thermophilic bacterium, Thermus thermophilus: lysine is synthesized via α-aminoadipic acid, not via diaminopimeric acid. J. Bacteriol. 181, 1713–1718 (1999).
Nishida, H. et al. A prokaryotic gene cluster involved in synthesis of lysine through the amino adipate pathway: a key to the evolution of amino acid biosynthesis. Genome Res. 9, 1175–1183 (1999).
Wulandari, A.P. et al. Characterization of bacterial homocitrate synthase involved in lysine biosynthesis. FEBS Lett. 522, 35–40 (2002).
Miyazaki, J., Kobashi, N., Nishiyama, M. & Yamane, H. Characterization of homoisocitrate dehydrogenase involved in lysine biosynthesis of an extremely thermophilic bacterium, Thermus thermophilus HB27, and evolutionary implication of β-decarboxylating dehydrogenase. J. Biol. Chem. 278, 1864–1871 (2003).
Jia, Y., Tomita, T., Yamauchi, K., Nishiyama, M. & Palmer, D.R.J. Kinetics and product analysis of the reaction catalyzed by recombinant homoaconitase from Thermus thermophilus. Biochem. J. 396, 479–485 (2006).
Miyazaki, T., Miyazaki, J., Yamane, H. & Nishiyama, M. α-Aminoadipate aminotransferase from an extremely thermophilic bacterium, Thermus thermophilus. Microbiology 150, 2327–2334 (2004).
Miyazaki, J., Kobashi, N., Nishiyama, M. & Yamane, H. Functional and evolutionary relationship between arginine biosynthesis and prokaryotic lysine biosynthesis through α-aminoadipate. J. Bacteriol. 183, 5067–5073 (2001).
Miyazaki, J., Kobashi, N., Fujii, T., Nishiyama, M. & Yamane, H. Characterization of a lysK gene as an argE homolog in Thermus thermophilus HB27. FEBS Lett. 512, 269–274 (2002).
Xu, Y., Labedan, B. & Glansdorff, N. Surprising arginine biosynthesis: a reappraisal of the enzymology and evolution of the pathway in microorganisms. Microbiol. Mol. Biol. Rev. 71, 36–47 (2007).
Leisinger, T. & Haas, D. N-Acetylglutamate synthase of Escherichia coli regulation of synthesis and activity by arginine. J. Biol. Chem. 250, 1690–1693 (1975).
Baetens, M., Legrain, C., Boyen, A. & Glansdorff, N. Genes and enzymes of the acetyl cycle of arginine biosynthesis in the extreme thermophilic bacterium Thermus thermophilus HB27. Microbiology 144, 479–492 (1998).
Martin, P.R. & Mulks, M.H. Sequence analysis and complementation studies of the argJ gene encoding ornithine acetyltransferase from Neisseria gonorrhoeae. J. Bacteriol. 174, 2694–2701 (1992).
Snoke, J.E. Isolation and properties of yeast glutathione synthetase. J. Biol. Chem. 213, 813–824 (1955).
Kang, W.K., Icho, T., Isono, S., Kitakawa, M. & Isono, K. Characterization of the gene rimK responsible for the addition of glutamic acid residues to the C-terminus of ribosomal protein S6 in Escherichia coli K12. Mol. Gen. Genet. 217, 281–288 (1989).
Iwasaki, T. et al. Rational design of a mononuclear metal site into the archaeal Rieske-type protein scaffold. J. Biol. Chem. 280, 9129–9134 (2005).
Kurihara, S. et al. A novel putrescine utilization pathway involves γ-glutamylated intermediates of Escherichia coli K-12. J. Biol. Chem. 280, 4602–4608 (2005).
Majerus, P.W., Alberts, A.W. & Vagelos, P.R. The acyl carrier protein of fatty acid synthesis: purification, physical properties, and substrate binding site. Proc. Natl. Acad. Sci. USA 51, 1231–1238 (1964).
Stachelhaus, T., Huser, A. & Marahiel, M.A. Biochemical characterization of peptidyl carrier protein (PCP), the thiolation domain of multifunctional peptide synthetases. Chem. Biol. 3, 913–921 (1996).
Lynen, F. On the structure of fatty acid synthetase of yeast. Eur. J. Biochem. 112, 431–442 (1980).
Lomakin, I.B., Xiong, Y. & Steitz, T.A. The crystal structure of yeast fatty acid synthase, a cellular machine with eight active sites working together. Cell 129, 319–332 (2007).
Jenni, S. et al. Structure of fungal fatty acid synthase and implications for iterative substrate shuttling. Science 316, 254–261 (2007).
Zhao, Q. et al. Characterization of the azinomycin B biosynthetic gene cluster revealing a different iterative type I polyketide synthase for naphthoate biosynthesis. Chem. Biol. 15, 693–705 (2008).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Gouet, P., Courcelle, E., Stuart, D.I. & Metoz, F. ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999).
Henne, A. et al. The genome sequence of the extreme thermophile Thermus thermophilus. Nat. Biotechnol. 22, 547–553 (2004).
White, O. et al. Genome sequence of the radioresistant bacterium Deinococcus radiodurans R1. Science 286, 1571–1577 (1999).
Kawarabayasi, Y. et al. Complete genome sequence of an aerobic thermoacidophilic crenarchaeon, Sulfolobus tokodaii strain7. DNA Res. 8, 123–140 (2001).
She, Q. et al. The complete genome of the crenarchaeon Sulfolobus solfataricus P2. Proc. Natl. Acad. Sci. USA 98, 7835–7840 (2001).
Kawarabayasi, Y. et al. Complete sequence and gene organization of the genome of a hyper-thermophilic archaebacterium, Pyrococcus horikoshii OT3. DNA Res. 5, 55–76 (1998).
Robb, F.T. et al. Genomic sequence of hyperthermophile, Pyrococcus furiosus: implications for physiology and enzymology. Methods Enzymol. 330, 134–157 (2001).
Kawarabayasi, Y. et al. Complete genome sequence of an aerobic hyper-thermophilic crenarchaeon, Aeropyrum pernix K1. DNA Res. 6, 83–101, 145–152 (1999).
Bolhuis, H. et al. The genome of the square archaeon Haloquadratum walsbyi: life at the limits of water activity. BMC Genomics 7, 169 (2006).
Baliga, N.S. et al. Genome sequence of Haloarcula marismortui: a halophilic archaeon from the Dead Sea. Genome Res. 14, 2221–2234 (2004).
Acknowledgements
We thank H. Satsu for technical help with amino acid analysis. This work was supported in part by a grant-in-aid for scientific research from the Ministry of Education, Culture, Sports, Science, and Technology of Japan, from the Nagase Science and Technology Foundation, from the Asahi Glass Foundation and from the Charitable Trust Araki Medical and Biochemistry Memorial Research Promotion Fund.
Author information
Authors and Affiliations
Contributions
Research planning and supervision were by T.T., T.K. and M.N.; biochemical experiments were by A.H.; gene knockout and replacement of T. thermophilus were by A.S.; LC-MS/MS and MALDI-TOF MS were by H.T., R.M., T.F. and C.N.; in silico modeling was by H.K. and T.T.; and manuscript writing was by A.H., T.T. and M.N.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 4638 kb)
Rights and permissions
About this article
Cite this article
Horie, A., Tomita, T., Saiki, A. et al. Discovery of proteinaceous N-modification in lysine biosynthesis of Thermus thermophilus. Nat Chem Biol 5, 673–679 (2009). https://doi.org/10.1038/nchembio.198
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.198
This article is cited by
-
Insight into de-regulation of amino acid feedback inhibition: a focus on structure analysis method
Microbial Cell Factories (2023)
-
Amino-group carrier-protein-mediated secondary metabolite biosynthesis in Streptomyces
Nature Chemical Biology (2016)
-
Identification of Nα-acetyl-α-lysine as a probable thermolyte and its accumulation mechanism in Salinicoccus halodurans H3B36
Scientific Reports (2015)
-
Genome-wide comprehensive analysis of transcriptional regulation by ArgR in Thermus thermophilus
Extremophiles (2014)
-
Lysine and arginine biosyntheses mediated by a common carrier protein in Sulfolobus
Nature Chemical Biology (2013)