Abstract
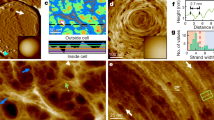
Here we report on in vivo measurement of the mechanical behavior of a cell surface sensor using single-molecule atomic force microscopy. We focus on the yeast wall stress component sensor Wsc1, a plasma membrane protein that is thought to function as a rigid probe of the cell wall status. We first map the distribution of individual histidine-tagged sensors on living yeast cells by scanning the cell surface with atomic force microscopy tips carrying nitrilotriacetate groups. We then show that Wsc1 behaves like a linear nanospring that is capable of resisting high mechanical force and of responding to cell surface stress. Both a genomic pmt4 deletion and the insertion of a stretch of glycines in Wsc1 result in substantial alterations in protein spring properties, supporting the important role of glycosylation at the extracellular serine/threonine-rich region.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Klis, F.M., Boorsma, A. & De Groot, P.W. Cell wall construction in Saccharomyces cerevisiae. Yeast 23, 185–202 (2006).
Levin, D.E. Cell wall integrity signaling in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 69, 262–291 (2005).
Straede, A. & Heinisch, J.J. Functional analyses of the extra- and intracellular domains of the yeast cell wall integrity sensors Mid2 and Wsc1. FEBS Lett. 581, 4495–4500 (2007).
Piao, H.L., Machado, I.M. & Payne, G.S. NPFXD-mediated endocytosis is required for polarity and function of a yeast cell wall stress sensor. Mol. Biol. Cell 18, 57–65 (2007).
Heinisch, J.J. Bakers yeast as a tool for the development of antifungal drugs which target cell integrity—an update. Expert Opin. Drug Discov. 3, 931–943 (2008).
Rodicio, R., Buchwald, U., Schmitz, H.P. & Heinisch, J.J. Dissecting sensor functions in cell wall integrity signaling in Kluyveromyces lactis. Fungal Genet. Biol. 45, 422–435 (2008).
Rief, M., Gautel, M., Oesterhelt, F., Fernandez, J.M. & Gaub, H.E. Reversible unfolding of individual titin immunoglobulin domains by AFM. Science 276, 1109–1112 (1997).
Oberhauser, A.F., Marszalek, P.E., Erickson, H.P. & Fernandez, J.M. The molecular elasticity of the extracellular matrix protein tenascin. Nature 393, 181–185 (1998).
Oesterhelt, F. et al. Unfolding pathways of individual bacteriorhodopsins. Science 288, 143–146 (2000).
Fernandez, J.M. & Li, H. Force-clamp spectroscopy monitors the folding trajectory of a single protein. Science 303, 1674–1678 (2004).
Vogel, V. & Sheetz, M. Local force and geometry sensing regulate cell functions. Nat. Rev. Mol. Cell Biol. 7, 265–275 (2006).
Sotomayor, M. & Schulten, K. Single-molecule experiments in vitro and in silico. Science 316, 1144–1148 (2007).
Müller, D.J. & Dufrêne, Y.F. Atomic force microscopy as a multifunctional molecular toolbox in nanobiotechnology. Nat. Nanotechnol. 3, 261–269 (2008).
Ahimou, F.O., Touhami, A. & Dufrêne, Y.F. Real-time imaging of the surface topography of living yeast cells by atomic force microscopy. Yeast 20, 25–30 (2003).
Koch, Y. & Rademacher, K. Chemical and enzymatic changes in the cell walls of Candida albicans and Saccharomyces cerevisiae by scanning microscopy. Can. J. Microbiol. 26, 965–970 (1980).
Verbelen, C., Gruber, H.J. & Dufrêne, Y.F. The NTA-His6 bond is strong enough for AFM single-molecular recognition studies. J. Mol. Recognit. 20, 490–494 (2007).
Merkel, R., Nassoy, P., Leung, A., Ritchie, K. & Evans, E. Energy landscapes of receptor-ligand bonds explored with dynamic force spectroscopy. Nature 397, 50–53 (1999).
Hinterdorfer, P. & Dufrêne, Y.F. Detection and localization of single molecular recognition events using atomic force microscopy. Nat. Methods 3, 347–355 (2006).
Lata, S., Reichel, A., Brock, R., Tampé, R. & Piehler, J. High-affinity adaptors for switchable recognition of histidine-tagged proteins. J. Am. Chem. Soc. 127, 10205–10215 (2005).
Krieg, M., Helenius, J., Heisenberg, C.-P. & Muller, D.J. A Bond for a lifetime: employing membrane nanotubes from living cells to determine receptor–ligand kinetics. Angew. Chem. Int. Ed. 47, 9775–9777 (2008).
Willer, T., Valero, M.C., Tanner, W., Cruces, J. & Strahl, S. O-mannosyl glycans: from yeast to novel associations with human disease. Curr. Opin. Struct. Biol. 13, 621–630 (2003).
Jentoft, N. Why are proteins O-glycosylated? Trends Biochem. Sci. 15, 291–294 (1990).
Lodder, A.L., Lee, T.K. & Ballester, R. Characterization of the Wsc1 protein, a putative receptor in the stress response of Saccharomyces cerevisiae. Genetics 152, 1487–1499 (1999).
Rajavel, M., Philip, B., Buehrer, B.M., Errede, B. & Levin, D.E. Mid2 is a putative sensor for cell integrity signaling in Saccharomyces cerevisiae. Mol. Cell. Biol. 19, 3969–3976 (1999).
Philip, B. & Levin, D.E. Wsc1 and Mid2 are cell surface sensors for cell wall integrity signaling that act through Rom2, a guanine nucleotide exchange factor for Rho1. Mol. Cell. Biol. 21, 271–280 (2001).
Lommel, M., Bagnat, M. & Strahl, S. Abberant processing of the WSC family and Mid2p cell surface sensors results in cell death of Saccharomyces cerevisiae O-mannosylation mutants. Mol. Cell. Biol. 24, 46–57 (2004).
Powell, C.D., Quain, D.E. & Smart, K.A. Chitin scar breaks in aged Saccharomyces cerevisiae. Microbiology 149, 3129–3137 (2003).
Lee, G. et al. Nanospring behaviour of ankyrin repeats. Nature 440, 246–249 (2006).
Schlierf, M. & Rief, M. Temperature softening of a protein in single-molecule experiments. J. Mol. Biol. 354, 497–503 (2005).
Law, R. et al. Pathway shifts and thermal softening in temperature-coupled forced unfolding of spectrin domains. Biophys. J. 85, 3286–3293 (2003).
Dubreuil, R.R. Functional links between membrane transport and the spectrin cytoskeleton. J. Membr. Biol. 211, 151–161 (2006).
Julien, M.A., Wang, P., Haller, C.A., Wen, J. & Chaikof, E.L. Mechanical strain regulates syndecan-4 expression and shedding in smooth muscle cells through differential activation of MAP kinase signaling pathways. Am. J. Physiol. Cell Physiol. 292, C517–C525 (2007).
Becker, N. et al. Molecular nanosprings in spider capture-like threads. Nat. Mater. 2, 278–283 (2003).
Straede, A., Corran, A., Bundy, J. & Heinisch, J.J. The effect of tea tree oil and antifungal agents on a reporter for yeast cell integrity signalling. Yeast 24, 321–334 (2007).
Arvanitidis, A. & Heinisch, J.J. Studies on the function of yeast phosphofructokinase subunits by in vitro mutagenesis. J. Biol. Chem. 269, 8911–8918 (1994).
Gietz, R.D. & Sugino, A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 74, 527–534 (1988).
Acknowledgements
This work was supported by the Belgian National Foundation for Scientific Research (FNRS), the Université catholique de Louvain (Fonds Spéciaux de Recherche), the Région wallonne, the Federal Office for Scientific, Technical and Cultural Affairs (Interuniversity Poles of Attraction Programme), and the Research Department of the Communauté française de Belgique (Concerted Research Action). Work at the University of Osnabrück was funded by the Deutsche Forschungsgemeinschaft within the framework of the SFB431.
Author information
Authors and Affiliations
Contributions
V.D., D.A., S.W., B.H., J.J.H. and Y.F.D. designed the experiments, analyzed the data and wrote the article. S.W., B.H. and J.J.H. performed the genetic manipulations. V.D. and D.A. collected the AFM data.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 254 kb)
Rights and permissions
About this article
Cite this article
Dupres, V., Alsteens, D., Wilk, S. et al. The yeast Wsc1 cell surface sensor behaves like a nanospring in vivo. Nat Chem Biol 5, 857–862 (2009). https://doi.org/10.1038/nchembio.220
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.220
This article is cited by
-
Force spectroscopy of single cells using atomic force microscopy
Nature Reviews Methods Primers (2021)
-
Plant cell wall integrity maintenance in model plants and crop species-relevant cell wall components and underlying guiding principles
Cellular and Molecular Life Sciences (2020)
-
Not just the wall: the other ways to turn the yeast CWI pathway on
International Microbiology (2020)
-
Atomic force microscopy-based characterization and design of biointerfaces
Nature Reviews Materials (2017)
-
Optical and force nanoscopy in microbiology
Nature Microbiology (2016)