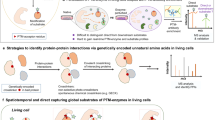
Abstract
Metabolic labeling of proteins with the methionine surrogate azidonorleucine can be targeted exclusively to specified cells through expression of a mutant methionyl-tRNA synthetase (MetRS). In complex cellular mixtures, proteins made in cells that express the mutant synthetase can be tagged with affinity reagents (for detection or enrichment) or fluorescent dyes (for imaging). Proteins made in cells that do not express the mutant synthetase are neither labeled nor detected.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Beynon, R.J. & Pratt, J.M. Mol. Cell. Proteomics 4, 857–872 (2005).
Mann, M. Nat. Rev. Mol. Cell Biol. 7, 952–958 (2006).
Dieterich, D.C., Link, A.J., Graumann, J., Tirrell, D.A. & Schuman, E.M. Proc. Natl. Acad. Sci. USA 103, 9482–9487 (2006).
Dieterich, D.C. et al. Nat. Protoc. 2, 532–540 (2007).
Prescher, J.A. & Bertozzi, C.R. Nat. Chem. Biol. 1, 13–21 (2005).
Beatty, K.E., Xie, F., Wang, Q. & Tirrell, D.A. J. Am. Chem. Soc. 127, 14150–14151 (2005).
Beatty, K.E. et al. Angew. Chem. Int. Edn Engl. 13, 7364–7367 (2006).
Zhang, C.G., Chromy, B.A. & McCutchen-Maloney, S.L. Expert Rev. Proteomics 2, 187–202 (2005).
Macpherson, A.J. & Harris, N.L. Nat. Rev. Immunol. 4, 478–485 (2004).
Ibba, M. & Söll, D. Science 286, 1893–1897 (1999).
Link, A.J. et al. Proc. Natl. Acad. Sci. USA 103, 10180–10185 (2006).
Yoo, T.H. & Tirrell, D.A. Angew. Chem. Int. Ed. 46, 5340–5346 (2006).
Tornøe, C.W., Christensen, C. & Meldal, M. J. Org. Chem. 67, 3057–3064 (2002).
Lewis, W.G. et al. Angew. Chem. Int. Ed. 41, 51053–51057 (2002).
Acknowledgements
We gratefully acknowledge the US National Institutes of Health (R21 DA020589 and R01 GM62523) and the US Army Research Office Institute for Collaborative Biotechnologies for support of this work. We thank K. Boulware for assistance in generating the three-dimensional movie of infected macrophages. E.M.S. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Methods (PDF 2114 kb)
Supplementary Movie 1
Three-dimensional rendering of cell-selective labelling of macrophage infection with E. coli. (MOV 2075 kb)
Rights and permissions
About this article
Cite this article
Ngo, J., Champion, J., Mahdavi, A. et al. Cell-selective metabolic labeling of proteins. Nat Chem Biol 5, 715–717 (2009). https://doi.org/10.1038/nchembio.200
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.200
This article is cited by
-
An integrated workflow for quantitative analysis of the newly synthesized proteome
Nature Communications (2023)
-
Photo-ANA enables profiling of host–bacteria protein interactions during infection
Nature Chemical Biology (2023)
-
Escherichia coli methionine-tRNAi/methionyl tRNA synthetase pairs induced protein initiation of interest (PII) expression
Applied Biological Chemistry (2022)
-
A CRISPR toolbox for generating intersectional genetic mouse models for functional, molecular, and anatomical circuit mapping
BMC Biology (2022)
-
Bioorthogonal chemistry
Nature Reviews Methods Primers (2021)