Abstract
Off-target binding of hydrophobic drugs can lead to unwanted side effects, either through specific or non-specific binding to unintended membrane protein targets. However, distinguishing the binding of drugs to membrane proteins from that of detergents, lipids and cofactors is challenging. Here, we use high-resolution mass spectrometry to study the effects of HIV protease inhibitors on the human zinc metalloprotease ZMPSTE24. This intramembrane protease plays a major role in converting prelamin A to mature lamin A. We monitored the proteolysis of farnesylated prelamin A peptide by ZMPSTE24 and unexpectedly found retention of the C-terminal peptide product with the enzyme. We also resolved binding of zinc, lipids and HIV protease inhibitors and showed that drug binding blocked prelamin A peptide cleavage and conferred stability to ZMPSTE24. Our results not only have relevance for the progeria-like side effects of certain HIV protease inhibitor drugs, but also highlight new approaches for documenting off-target drug binding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wolfe, M. S. & Kopan, R. Intramembrane proteolysis: theme and variations. Science 305, 1119–1123 (2004).
Marcoux, J. et al. Mass spectrometry reveals synergistic effects of nucleotides, lipids, and drugs binding to a multidrug resistance efflux pump. Proc. Natl Acad. Sci. USA 110, 9704–9709 (2013).
Hopper, J. T. & Robinson, C. V. Mass spectrometry quantifies protein interactions—from molecular chaperones to membrane porins. Angew. Chem. Int. Ed. 53, 14002–14015 (2014).
Freeman, M. The rhomboid-like superfamily: molecular mechanisms and biological roles. Annu. Rev. Cell Dev. Biol. 30, 235–254 (2014).
Avci, D. & Lemberg, M. K. Clipping or extracting: two ways to membrane protein degradation. Trends Cell Biol. 25, 611–622 (2015).
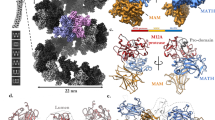
Quigley, A. et al. The structural basis of ZMPSTE24-dependent laminopathies. Science 339, 1604–1607 (2013).
Pryor, E. E. Jr et al. Structure of the integral membrane protein CAAX protease Ste24p. Science 339, 1600–1604 (2013).
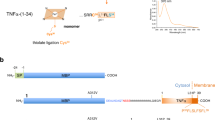
Young, S. G., Fong, L. G. & Michaelis, S. Prelamin A, Zmpste24, misshapen cell nuclei, and progeria—new evidence suggesting that protein farnesylation could be important for disease pathogenesis. J. Lipid Res. 46, 2531–2558 (2005).
Manolaridis, I. et al. Mechanism of farnesylated CAAX protein processing by the intramembrane protease Rce1. Nature 504, 301–305 (2013).
Davies, B. S., Fong, L. G., Yang, S. H., Coffinier, C. & Young, S. G. The posttranslational processing of prelamin A and disease. Annu. Rev. Genom. Hum. Genet. 10, 153–174 (2009).
Eriksson, M. et al. Recurrent de novo point mutations in lamin A cause Hutchinson–Gilford progeria syndrome. Nature 423, 293–298 (2003).
Moulson, C. L. et al. Increased progerin expression associated with unusual LMNA mutations causes severe progeroid syndromes. Hum. Mutat. 28, 882–889 (2007).
Shackleton, S. et al. Compound heterozygous ZMPSTE24 mutations reduce prelamin A processing and result in a severe progeroid phenotype. J. Med. Genet. 42, e36 (2005).
Ahmad, Z., Zackai, E., Medne, L. & Garg, A. Early onset mandibuloacral dysplasia due to compound heterozygous mutations in ZMPSTE24. Am. J. Med. Genet. A 152A, 2703–2710 (2010).
Barrowman, J., Wiley, P. A., Hudon-Miller, S. E., Hrycyna, C. A. & Michaelis, S. Human ZMPSTE24 disease mutations: residual proteolytic activity correlates with disease severity. Hum. Mol. Genet. 21, 4084–4093 (2012).
Coffinier, C. et al. HIV protease inhibitors block the zinc metalloproteinase ZMPSTE24 and lead to an accumulation of prelamin A in cells. Proc. Natl Acad. Sci. USA 104, 13432–13437 (2007).
Coffinier, C. et al. A potent HIV protease inhibitor, darunavir, does not inhibit ZMPSTE24 or lead to an accumulation of farnesyl–prelamin A in cells. J. Biol. Chem. 283, 9797–9804 (2008).
Hudon, S. E. et al. HIV-protease inhibitors block the enzymatic activity of purified Ste24p. Biochem. Biophys. Res. Commun. 374, 365–368 (2008).
Barrowman, J. & Michaelis, S. ZMPSTE24, an integral membrane zinc metalloprotease with a connection to progeroid disorders. Biol. Chem. 390, 761–773 (2009).
Perrin, S. et al. HIV protease inhibitors do not cause the accumulation of prelamin A in PBMCs from patients receiving first line therapy: the ANRS EP45 ‘aging’ study. PLoS ONE 7, e53035 (2012).
Agarwal, A. K., Fryns, J. P., Auchus, R. J. & Garg, A. Zinc metalloproteinase, ZMPSTE24, is mutated in mandibuloacral dysplasia. Hum. Mol. Genet. 12, 1995–2001 (2003).
Torres, R. A. & Lewis, W. Aging and HIV/AIDS: pathogenetic role of therapeutic side effects. Lab. Invest. 94, 120–128 (2014).
Rose, R. J., Damoc, E., Denisov, E., Makarov, A. & Heck, A. J. High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies. Nat. Methods 9, 1084–1086 (2012).
Gault, J. et al. High-resolution mass spectrometry of small molecules bound to membrane proteins. Nat. Methods 13, 333–336 (2016).
Belov, M. E. et al. From protein complexes to subunit backbone fragments: a multi-stage approach to native mass spectrometry. Anal. Chem. 85, 11163–11173 (2013).
Erba, E. B. & Zenobi, R. Mass spectrometric studies of dissociation constants of noncovalent complexes. Annu. Rep. Prog. Chem. C 107, 199–228 (2011).
Clemmer, D. E., Hudgins, R. R. & Jarrold, M. F. Naked protein conformations: cytochrome c in the gas phase. J. Am. Chem. Soc. 117, 10141–10142 (1995).
Gill, A. C., Jennings, K. R., Wyttenbach, T. & Bowers, M. T. Conformations of biopolymers in the gas phase: a new mass spectrometric method. Int. J. Mass Spectrom. 195–196, 685–697 (2000).
Ruotolo, B. T. et al. Ion mobility-mass spectrometry reveals long-lived, unfolded intermediates in the dissociation of protein complexes. Angew Chem. Int. Ed. 46, 8001–8004 (2007).
Laganowsky, A. et al. Membrane proteins bind lipids selectively to modulate their structure and function. Nature 510, 172–175 (2014).
Allison, T. M. et al. Quantifying the stabilizing effects of protein–ligand interactions in the gas phase. Nat. Commun. 6, 8551 (2015).
Silvius, J. R. & l'Heureux, F. Fluorimetric evaluation of the affinities of isoprenylated peptides for lipid bilayers. Biochemistry 33, 3014–3022 (1994).
Coffinier, C. et al. Direct synthesis of lamin A, bypassing prelamin A processing, causes misshapen nuclei in fibroblasts but no detectable pathology in mice. J. Biol. Chem. 285, 20818–20826 (2010).
Toth, J. I. et al. Blocking protein farnesyltransferase improves nuclear shape in fibroblasts from humans with progeroid syndromes. Proc. Natl Acad. Sci. USA 102, 12873–12878 (2005).
Yang, S. H. et al. Severe hepatocellular disease in mice lacking one or both CaaX prenyltransferases. J. Lipid Res. 53, 77–86 (2012).
Lu, Z. Second generation HIV protease inhibitors against resistant virus. Exp. Opin. Drug Disc. 3, 775–786 (2008).
Boffito, M. et al. Protein binding in antiretroviral therapies. AIDS Res. Hum. Retroviruses 19, 825–835 (2003).
Solas, C. et al. Discrepancies between protease inhibitor concentrations and viral load in reservoirs and sanctuary sites in human immunodeficiency virus-infected patients. Antimicrob. Agents Chemother. 47, 238–243 (2003).
Caron, M. et al. Human lipodystrophies linked to mutations in A-type lamins and to HIV protease inhibitor therapy are both associated with prelamin A accumulation, oxidative stress and premature cellular senescence. Cell Death Differ. 14, 1759–1767 (2007).
Loo, J. A., DeJohn, D. E., Du, P., Stevenson, T. I. & Ogorzalek Loo, R. R. Application of mass spectrometry for target identification and characterization. Med. Res. Rev. 19, 307–319 (1999).
Harvey, S. R. et al. Small-molecule inhibition of c-MYC:MAX leucine zipper formation is revealed by ion mobility mass spectrometry. J. Am. Chem. Soc. 134, 19384–19392 (2012).
Pacholarz, K. J., Garlish, R. A., Taylor, R. J. & Barran, P. E. Mass spectrometry based tools to investigate protein–ligand interactions for drug discovery. Chem. Soc. Rev. 41, 4335–4355 (2012).
Maple, H. J. et al. Application of the Exactive Plus EMR for automated protein–ligand screening by non-covalent mass spectrometry. Rapid Commun. Mass Spectrom. 28, 1561–1568 (2014).
Jacobs, A. D. et al. Resolution of stepwise cooperativities of copper binding by the homotetrameric copper-sensitive operon repressor (CsoR): impact on structure and stability. Angew Chem. Int. Ed. 54, 12795–12799 (2015).
Kell, D. B., Dobson, P. D., Bilsland, E. & Oliver, S. G. The promiscuous binding of pharmaceutical drugs and their transporter-mediated uptake into cells: what we (need to) know and how we can do so. Drug Discov. Today 18, 218–239 (2013).
Acknowledgements
This work is supported by programme grants from the Medical Research Council (98101), the National Institutes of Health (NIH) and an ERC Advanced Investigator Award IMPRESS (26851). C.V.R. has a Wellcome Trust Investigator Award. E.P.C. and A.Q. are funded by the Structural Genomics Consortium, a registered charity (no. 1097737) that receives funds from AbbVie, Bayer, Boehringer Ingelheim, the Canada Foundation for Innovation, the Canadian Institutes for Health Research, Genome Canada, GlaxoSmithKline, Janssen, Lilly Canada, the Novartis Research Foundation, the Ontario Ministry of Economic Development and Innovation, Pfizer, Takeda and the Wellcome Trust (092809/Z/10/Z). E.P.C. and A.Q. are also funded by a Medical Research Council grant number MR/L017458/1. S.Mi. was funded for this work by NIH grant R01 GM041223. S.G.Y. was supported by NIH grants AG035626-10 and HL126551. The authors thank S. Wang and B. Rotty for purification of three of the protein samples used in this study. J.G. is a Junior Research Fellow at Queen's College, Oxford.
Author information
Authors and Affiliations
Contributions
S.Me. and C.V.R. designed the research with assistance from A.Q. and E.P.C. A.Q. and E.P.C. purified ZMPSTE24. S.Me. and J.G. carried out the mass spectrometry experiments. J.M. initiated preliminary mass spectrometry experiment. S.Mi. helped with new reagents. S.G.Y. performed the western blot analysis. S.Me. and C.V.R. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
C.V.R. is a consultant to Omass Technologies Ltd, a spinout company from the Department of Chemistry at the University of Oxford.
Supplementary information
Rights and permissions
About this article
Cite this article
Mehmood, S., Marcoux, J., Gault, J. et al. Mass spectrometry captures off-target drug binding and provides mechanistic insights into the human metalloprotease ZMPSTE24. Nature Chem 8, 1152–1158 (2016). https://doi.org/10.1038/nchem.2591
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2591
This article is cited by
-
A DFT/TD-DFT study of [Amprenavir + C60] PET nanocomplex: feasibility of C60 fullerene application as a nanocarrier
Journal of the Iranian Chemical Society (2022)
-
Studying biomolecular folding and binding using temperature-jump mass spectrometry
Nature Communications (2020)
-
Molecular basis for chirality-regulated Aβ self-assembly and receptor recognition revealed by ion mobility-mass spectrometry
Nature Communications (2019)
-
Nanosecond photochemically promoted click chemistry for enhanced neuropeptide visualization and rapid protein labeling
Nature Communications (2019)
-
A Semi-Empirical Framework for Interpreting Traveling Wave Ion Mobility Arrival Time Distributions
Journal of the American Society for Mass Spectrometry (2019)