Abstract
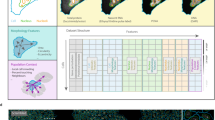
Cell division is fundamental for all organisms. Here we report a genome-scale RNA-mediated interference screen in HeLa cells designed to identify human genes that are important for cell division. We have used a library of endoribonuclease-prepared short interfering RNAs for gene silencing and have used DNA content analysis to identify genes that induced cell cycle arrest or altered ploidy on silencing. Validation and secondary assays were performed to generate a nine-parameter loss-of-function phenoprint for each of the genes. These phenotypic signatures allowed the assignment of genes to specific functional classes by combining hierarchical clustering, cross-species analysis and proteomic data mining. We highlight the richness of our dataset by ascribing novel functions to genes in mitosis and cytokinesis. In particular, we identify two evolutionarily conserved transcriptional regulatory networks that govern cytokinesis. Our work provides an experimental framework from which the systematic analysis of novel genes necessary for cell division in human cells can begin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Hannon, G. J. RNA interference. Nature 418, 244–251 (2002).
Echeverri, C. J. & Perrimon, N. High-throughput RNAi screening in cultured cells: a user's guide. Nature Rev. Genet. 7, 373–384 (2006).
Bjorklund, M. et al. Identification of pathways regulating cell size and cell-cycle progression by RNAi. Nature 439, 1009–1013 (2006).
Eggert, U. S. et al. Parallel chemical genetic and genome-wide RNAi screens identify cytokinesis inhibitors and targets. PLoS Biol. 2, e379 (2004).
Bettencourt-Dias, M. et al. Genome-wide survey of protein kinases required for cell cycle progression. Nature 432, 980–987 (2004).
Gonczy, P. et al. Functional genomic analysis of cell division in C. elegans using RNAi of genes on chromosome III. Nature 408, 331–336 (2000).
Sönnichsen, B. et al. Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans. Nature 434, 462–469 (2005).
Kastan, M. B. & Bartek, J. Cell-cycle checkpoints and cancer. Nature 432, 316–323 (2004).
Massagué, J. G1 cell-cycle control and cancer. Nature 432, 298–306 (2004).
Rajagopalan, H. & Lengauer, C. Aneuploidy and cancer. Nature 432, 338–341 (2004).
Buchholz, F., Kittler, R., Slabicki, M. & Theis, M. Enzymatically prepared RNAi libraries. Nature Methods 3, 696–700 (2006).
Kittler, R. et al. An endoribonuclease-prepared siRNA screen in human cells identifies genes essential for cell division. Nature 432, 1036–1040 (2004).
Kittler, R. et al. Genome-wide resources of endoribonuclease-prepared short interfering RNAs for specific loss-of-function studies. Nature Methods 4, 337–344 (2007).
Jackson, A. L. et al. Expression profiling reveals off-target gene regulation by RNAi. Nature Biotechnol. 21, 635–637 (2003).
Birmingham, A. et al. 3′ UTR seed matches, but not overall identity, are associated with RNAi off-targets. Nature Methods 3, 199–204 (2006).
Jackson, A. L. et al. Widespread siRNA 'off-target' transcript silencing mediated by seed region sequence complementarity. RNA 12, 1179–1187 (2006).
Nelson, D. M. et al. Coupling of DNA synthesis and histone synthesis in S phase independent of cyclin/cdk2 activity. Mol. Cell. Biol. 22, 7459–7472 (2002).
Ye, X. et al. Defective S phase chromatin assembly causes DNA damage, activation of the S phase checkpoint, and S phase arrest. Mol. Cell 11, 341–351 (2003).
Shiomi, Y. et al. ATP-dependent structural change of the eukaryotic clamp-loader protein, replication factor C. Proc. Natl Acad. Sci. USA 97, 14127–14132 (2000).
Storchova, Z. & Pellman, D. From polyploidy to aneuploidy, genome instability and cancer. Nature Rev. Mol. Cell Biol. 5, 45–54 (2004).
Takimoto, M. et al. Frequent expression of new cancer/testis gene D40/AF15q14 in lung cancers of smokers. Br. J. Cancer 86, 1757–1762 (2002).
Chinwalla, V. et al. A t(11;15) fuses MLL to two different genes, AF15q14 and a novel gene MPFYVE on chromosome 15. Oncogene 22, 1400–1410 (2003).
Kuefer, M. U. et al. Characterization of the MLL partner gene AF15q14 involved in t(11;15)(q23;q14). Oncogene 22, 1418–1424 (2003).
Cheeseman, I. M. et al. A conserved protein network controls assembly of the outer kinetochore and its ability to sustain tension. Genes Dev. 18, 2255–2268 (2004).
Obuse, C. et al. A conserved Mis12 centromere complex is linked to heterochromatic HP1 and outer kinetochore protein Zwint-1. Nature Cell Biol. 6, 1135–1141 (2004).
Desai, A. et al. KNL-1 directs assembly of the microtubule-binding interface of the kinetochore in C. elegans. Genes Dev. 17, 2421–2435 (2003).
Mukherji, M. et al. Genome-wide functional analysis of human cell-cycle regulators. Proc. Natl Acad. Sci. USA 103, 14819–14824 (2006).
Echard, A., Hickson, G. R., Foley, E. & O'Farrell, P. H. Terminal cytokinesis events uncovered after an RNAi screen. Curr. Biol. 14, 1685–1693 (2004).
Jepsen, K. & Rosenfeld, M. G. Biological roles and mechanistic actions of co-repressor complexes. J. Cell Sci. 115, 689–698 (2002).
Pijnappel, W. W. et al. The S. cerevisiae SET3 complex includes two histone deacetylases, Hos2 and Hst1, and is a meiotic-specific repressor of the sporulation gene program. Genes Dev. 15, 2991–3004 (2001).
Kittler, R. et al. RNA interference rescue by bacterial artificial chromosome transgenesis in mammalian tissue culture cells. Proc. Natl Acad. Sci. USA 102, 2396–2401 (2005).
Lewis, P. W. et al. Identification of a Drosophila Myb-E2F2/RBF transcriptional repressor complex. Genes Dev. 18, 2929–2940 (2004).
Korenjak, M. et al. Native E2F/RBF complexes contain Myb-interacting proteins and repress transcription of developmentally controlled E2F target genes. Cell 119, 181–193 (2004).
Harrison, M. M., Ceol, C. J., Lu, X. & Horvitz, H. R. Some C. elegans class B synthetic multivulva proteins encode a conserved LIN-35 Rb-containing complex distinct from a NuRD-like complex. Proc. Natl Acad. Sci. USA 103, 16782–16787 (2006).
Dimova, D. K., Stevaux, O., Frolov, M. V. & Dyson, N. J. Cell cycle-dependent and cell cycle-independent control of transcription by the Drosophila E2F/RB pathway. Genes Dev. 17, 2308–2320 (2003).
Ren, B. et al. E2F integrates cell cycle progression with DNA repair, replication, and G2/M checkpoints. Genes Dev. 16, 245–256 (2002).
Hauser, B. A., He, J. Q., Park, S. O. & Gasser, C. S. TSO1 is a novel protein that modulates cytokinesis and cell expansion in Arabidopsis. Development 127, 2219–2226 (2000).
Beall, E. L. et al. Role for a Drosophila Myb-containing protein complex in site-specific DNA replication. Nature 420, 833–837 (2002).
Litovchick, L. et al. Evolutionarily conserved multisubunit RBL2/p130 and E2F4 protein complex represses human cell cycle-dependent genes in quiescence. Mol. Cell 26, 539–551 (2007).
Echeverri, C. J. et al. Minimizing the risk of reporting false positives in large-scale RNAi screens. Nature Methods 3, 777–779 (2006).
Gunsalus, K. C. et al. Predictive models of molecular machines involved in Caenorhabditis elegans early embryogenesis. Nature 436, 861–865 (2005).
Rual, J. F. et al. Towards a proteome-scale map of the human protein–protein interaction network. Nature 437, 1173–1178 (2005).
Kittler, R., Heninger, A. K., Franke, K., Habermann, B. & Buchholz, F. Production of endoribonuclease-prepared short interfering RNAs for gene silencing in mammalian cells. Nature Methods 2, 779–784 (2005).
Cheeseman, I. M. & Desai, A. A combined approach for the localization and tandem affinity purification of protein complexes from metazoans. Sci STKE 2005, pl1 (2005).
Acknowledgements
We thank S. Payne for support with laser scanning cytometry; E. Krausz for support with automated transfection; D. Pinchev for help with the CASC5 experiments; E. Tanaka, F. Stewart and W. Zachariae for critical reading of an earlier version of the manuscript; P. Goodwin and C. Brown for development of the DeltaVision/CellWorX screening platform; and all members of the Buchholz laboratory and our colleagues in Mitocheck and SMP-RNAi for discussions. This study was funded by the Max Planck Society, the Bundesministerium für Bildung und Forschung grant SMP-RNAi under the framework of NGFN-2 (01GR0402) and the EU-FP6 grant Mitocheck (LSHG-CT-2004-503464). During the course of this work R.K. and L.P. were supported by postdoctoral fellowships of the Human Frontier Science Program, and L.P. was supported by the Samuel Lunenfeld Research Institute.
Author information
Authors and Affiliations
Contributions
R.K and L.P. contributed equally to this work. R.K., L.P. and F.B. performed project planning, experimental work, data analysis and wrote the manuscript. A.-K.H., M.S., M.T., L.M., I.P., S.L., H.G., K.K., J.W., V.S., C.R., W.B., A.L.J. and B.H. performed experimental work and data analysis. A.A.H. performed data analysis.
Corresponding author
Supplementary information
Supplementary Information
Supplementary Methods (PDF 335 kb)
Rights and permissions
About this article
Cite this article
Kittler, R., Pelletier, L., Heninger, AK. et al. Genome-scale RNAi profiling of cell division in human tissue culture cells. Nat Cell Biol 9, 1401–1412 (2007). https://doi.org/10.1038/ncb1659
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1659
This article is cited by
-
Biochemistry—Not Oncogenes—May Demystify and Defeat Cancer
Oncology and Therapy (2023)
-
Single object profiles regression analysis (SOPRA): a novel method for analyzing high-content cell-based screens
BMC Bioinformatics (2022)
-
Biallelic variants in COPB1 cause a novel, severe intellectual disability syndrome with cataracts and variable microcephaly
Genome Medicine (2021)
-
Secreted midbody remnants are a class of extracellular vesicles molecularly distinct from exosomes and microparticles
Communications Biology (2021)
-
A novel lncRNA PTTG3P/miR-132/212-3p/FoxM1 feedback loop facilitates tumorigenesis and metastasis of pancreatic cancer
Cell Death Discovery (2020)