Abstract
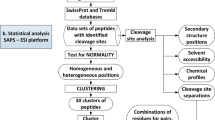
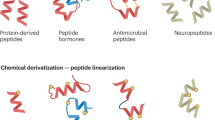
We introduce human proteome–derived, database-searchable peptide libraries for characterizing sequence-specific protein interactions. To identify endoprotease cleavage sites, we used peptides in such libraries with protected primary amines to simultaneously determine sequence preferences on the N-terminal (nonprime P) and C-terminal (prime P′) sides of the scissile bond. Prime-side cleavage products were tagged with biotin, isolated and identified by tandem mass spectrometry, and the corresponding nonprime-side sequences were derived from human proteome databases using bioinformatics. Identification of hundreds to over 1,000 individual cleaved peptides allows the consensus protease cleavage site and subsite cooperativity to be readily determined from P6 to P6′. For the highly specific GluC protease, >95% of the 558 cleavage sites identified displayed the canonical selectivity. For the broad-specificity matrix metalloproteinase 2, >1,200 peptidic cleavage sites were identified. Profiling of HIV protease 1, caspase 3, caspase 7, cathepsins K and G, elastase and thrombin showed that this approach is broadly applicable to all mechanistic classes of endoproteases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Diamond, S.L. Methods for mapping protease specificity. Curr. Opin. Chem. Biol. 11, 4651 (2007).
Schilling, O. & Overall, C.M. Proteomic discovery of protease substrates. Curr. Opin. Chem. Biol. 11, 36–45 (2007).
Overall, C.M. & Kleifeld, O. Validating matrix metalloproteinases as drug targets and antitargets for cancer therapy. Nat. Rev. Cancer 6, 227–239 (2006).
Puente, X.S., Sanchez, L.M., Overall, C.M. & LópezOtín, C. Human and mouse proteases: a comparative genomic approach. Nat. Rev. Genet. 4, 544–558 (2003).
Rawlings, N.D., Morton, F.R., Kok, C.Y., Kong, J. & Barrett, A.J. MEROPS: the peptidase database. Nucleic Acids Res. 36, D320–325 (2008).
Schechter, I. & Berger, A. On the size of the active site in proteases. I. Papain. Biochem. Biophys. Res. Commun. 27, 157–162 (1967).
Schechter, I. Mapping of the active site of proteases in the 1960s and rational design of inhibitors/drugs in the 1990s. Curr. Protein Pept. Sci. 6, 501–512 (2005).
Turk, B. Targeting proteases: successes, failures and future prospects. Nat. Rev. Drug Discov. 5, 785–799 (2006).
Matthews, D.J. & Wells, J.A. Substrate phage: selection of protease substrates by monovalent phage display. Science 260, 1113–1117 (1993).
Boulware, K.T. & Daugherty, P.S. Protease specificity determination by using cellular libraries of peptide substrates (CLiPS). Proc. Natl. Acad. Sci. USA 103, 7583–7588 (2006).
Chen, E.I. et al. A unique substrate recognition profile for matrix metalloproteinase-2. J. Biol. Chem. 277, 4485–4491 (2001).
Kerr, F.K. et al. Elucidation of the substrate specificity of the C1s protease of the classical complement pathway. J. Biol. Chem. 280, 39510–39514 (2005).
Rano, T.A. et al. A combinatorial approach for determining protease specificities: application to interleukin-1β converting enzyme (ICE). Chem. Biol. 4, 149–155 (1997).
Harris, J.L. et al. Rapid and general profiling of protease specificity by using combinatorial fluorogenic substrate libraries. Proc. Natl. Acad. Sci. USA 97, 7754–7759 (2000).
Harris, J. et al. Activity profile of dust mite allergen extract using substrate libraries and functional proteomic microarrays. Chem. Biol. 11, 1361–1372 (2004).
Salisbury, C.M., Maly, D.J. & Ellman, J.A. Peptide microarrays for the determination of protease substrate specificity. J. Am. Chem. Soc. 124, 14868–14870 (2002).
Gosalia, D.N., Salisbury, C.M., Ellman, J.A. & Diamond, S.L. High throughput substrate specificity profiling of serine and cysteine proteases using solution-phase fluorogenic peptide microarrays. Mol. Cell. Proteomics 4, 626–636 (2005).
Winssinger, N. et al. PNA-encoded protease substrate microarrays. Chem. Biol. 11, 1351–1360 (2004).
Turk, B.E., Huang, L.L., Piro, E.T. & Cantley, L.C. Determination of protease cleavage site motifs using mixture-based oriented peptide libraries. Nat. Biotechnol. 19, 661–667 (2001).
Gorodkin, J., Heyer, L.J., Brunak, S. & Stormo, G.D. Displaying the information contents of structural RNA alignments: the structure logos. Comput. Appl. Biosci. 13, 583–586 (1997).
Schneider, T.D. & Stephens, R.M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Kersey, P.J. et al. The International Protein Index: an integrated database for proteomics experiments. Proteomics 4, 1985–1988 (2004).
Drapeau, G.R., Boily, Y. & Houmard, J. Purification and properties of an extracellular protease of Staphylococcus aureus. J. Biol. Chem. 247, 6720–6726 (1972).
Netzel-Arnett, S. et al. Comparative sequence specificities of human 72 and 92-kDa gelatinases (type IV collagenases) and PUMP (matrilysin). Biochemistry 32, 6427–6432 (1993).
Nagase, H. & Fields, G.B. Human matrix metalloproteinase specificity studies using collagen sequence-based synthetic peptides. Biopolymers 40, 399–416 (1996).
Gomis-Ruth, F.X. Structural aspects of the metzincin clan of metalloendopeptidases. Mol. Biotechnol. 24, 157–202 (2003).
Maskos, K. Crystal structures of MMPs in complex with physiological and pharmacological inhibitors. Biochimie 87, 249–263 (2005).
Thornberry, N.A. et al. A combinatorial approach defines specificities of members of the caspase family and granzyme B. Functional relationships established for key mediators of apoptosis. J. Biol. Chem. 272, 17907–17911 (1997).
Stephens, A.W., Siddiqui, A. & Hirs, C.H. Site-directed mutagenesis of the reactive center (serine 394) of antithrombin III. J. Biol. Chem. 263, 15849–15852 (1988).
Lecaille, F., Bromme, D. & Lalmanach, G. Biochemical properties and regulation of cathepsin K activity. Biochimie 90, 208–226 (2008).
Fosang, A.J. et al. The interglobular domain of cartilage aggrecan is cleaved by PUMP, gelatinases, and cathepsin B. J. Biol. Chem. 267, 19470–19474 (1992).
Stetler-Stevenson, W.G., Krutzsch, H.C., Wacher, M.P., Margulies, I.M. & Liotta, L.A. The activation of human type IV collagenase proenzyme. Sequence identification of the major conversion product following organomercurial activation. J. Biol. Chem. 264, 1353–1356 (1989).
Van Damme, P. et al. Caspase–specific and nonspecific in vivo protein processing during Fas-induced apoptosis. Nat. Methods 2, 771–777 (2005).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
López–Otín, C. & Overall, C.M. Protease degradomics: a new challenge for proteomics. Nat. Rev. Mol. Cell Biol. 3, 509–519 (2002).
Tam, E.M., Morrison, C.J., Wu, Y.I., Stack, M.S. & Overall, C.M. Membrane protease proteomics: isotope-coded affinity tag MS identification of undescribed MT1-matrix metalloproteinase substrates. Proc. Natl. Acad. Sci. USA 101, 6917–6922 (2004).
Dean, R.A. & Overall, C.M. Proteomics discovery of metalloproteinase substrates in the cellular context by iTRAQ™ labeling reveals a diverse MMP–2 substrate degradome. Mol. Cell. Proteomics 6, 611–623 (2007).
McDonald, L., Robertson, D.H., Hurst, J.L. & Beynon, R.J. Positional proteomics: selective recovery and analysis of N-terminal proteolytic peptides. Nat. Methods 2, 955–957 (2005).
Enoksson, M. et al. Identification of proteolytic cleavage sites by quantitative proteomics. J. Proteome Res. 6, 2850–2858 (2007).
Overall, C.M., McQuibban, G.A. & Clark-Lewis, I. Discovery of chemokine substrates for matrix metalloproteinases by exosite scanning: a new tool for degradomics. Biol. Chem. 383, 1059–1066 (2002).
McQuibban, G.A. et al. Inflammation dampened by gelatinase A cleavage of monocyte chemoattractant protein-3. Science 289, 1202–1206 (2000).
Boyd, S.E., Pike, R.N., Rudy, G.B., Whisstock, J.C. & Garcia de la Banda, M. PoPS: a computational tool for modeling and predicting protease specificity. J. Bioinform. Comput. Biol. 3, 551–585 (2005).
Dean, R.A. et al. Identification of candidate angiogenic inhibitors processed by matrix metalloproteinase 2 (MMP-2) in cell-based proteomic screens: disruption of vascular endothelial growth factor (VEGF)/heparin affin regulatory peptide (pleiotrophin) and VEGF/connective tissue growth factor angiogenic inhibitory complexes by MMP2 proteolysis. Mol. Cell. Biol. 27, 8454–8465 (2007).
Bigg, H.F. et al. Tissue inhibitor of metalloproteinases-4 inhibits but does not support the activation of gelatinase A via efficient inhibition of membrane type 1-matrix metalloproteinase. Cancer Res. 61, 3610–3618 (2001).
Stricklin, G.P., Jeffrey, J.J., Roswit, W.T. & Eisen, A.Z. Human skin fibroblast procollagenase: mechanisms of activation by organomercurials and trypsin. Biochemistry 22, 61–68 (1983).
Keller, A., Nesvizhskii, A.I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Elias, J.E., Haas, W., Faherty, B.K. & Gygi, S.P. Comparative evaluation of mass spectrometry platforms used in largescale proteomics investigations. Nat. Methods 2, 667–675 (2005).
Stephan, C. et al. Automated reprocessing pipeline for searching heterogeneous mass spectrometric data of the HUPO Brain Proteome Project pilot phase. Proteomics 6, 5015–5029 (2006).
Martens, L. et al. PRIDE: the proteomics identifications database. Proteomics 5, 3537–3545 (2005).
Acknowledgements
The authors thank S. Perry, S. He and W. Chen (UBC) for mass spectrometer operation, U. auf dem Keller (UBC) for scientific discussion, S. Boyd (Monash University, Melbourne, Australia) for assistance in building the MMP-2 PoPS model and L. Martens (European Bioinformatics Institute) for assistance with data submission to PRIDE. O.S. was supported by the Deutsche Forschungsgemeinschaft and the Michael Smith Foundation for Health Research. C.M.O. is supported by a Canada Research Chair in Metalloproteinase Proteomics and Systems Biology with research grants from the Canadian Institutes of Health Research, the National Cancer Institute of Canada (with funds raised by the Canadian Cancer Association), and the Canadian Breast Cancer Research Alliance Special Program Grant on Metastasis as well as with a center grant from the Michael Smith Research Foundation.
Author information
Authors and Affiliations
Contributions
O.S. designed and implemented the chemistry in the workflow, performed all analyses, wrote the Perl scripts and performed the bioinformatics and wrote and edited the paper. C.M.O. conceived, designed and oversaw the development of PICS, performed proof of concept experiments, wrote and edited the paper and provided financial support for the project.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–24, Figures 1–10 (PDF 5997 kb)
Rights and permissions
About this article
Cite this article
Schilling, O., Overall, C. Proteome-derived, database-searchable peptide libraries for identifying protease cleavage sites. Nat Biotechnol 26, 685–694 (2008). https://doi.org/10.1038/nbt1408
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1408
This article is cited by
-
Proteomic data and structure analysis combined reveal interplay of structural rigidity and flexibility on selectivity of cysteine cathepsins
Communications Biology (2023)
-
Mapping specificity, cleavage entropy, allosteric changes and substrates of blood proteases in a high-throughput screen
Nature Communications (2021)
-
Bile and urine peptide marker profiles: access keys to molecular pathways and biological processes in cholangiocarcinoma
Journal of Biomedical Science (2020)
-
Multiobjective evolutionary-based multi-kernel learner for realizing transfer learning in the prediction of HIV-1 protease cleavage sites
Soft Computing (2020)
-
Neutrophilic proteolysis in the cystic fibrosis lung correlates with a pathogenic microbiome
Microbiome (2019)