Abstract
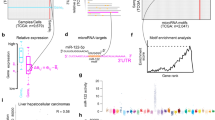
MicroRNAs (miRNAs) belong to a class of noncoding, regulatory RNAs that is involved in oncogenesis and shows remarkable tissue specificity. Their potential for tumor classification suggests they may be used in identifying the tissue in which cancers of unknown primary origin arose, a major clinical problem. We measured miRNA expression levels in 400 paraffin-embedded and fresh-frozen samples from 22 different tumor tissues and metastases. We used miRNA microarray data of 253 samples to construct a transparent classifier based on 48 miRNAs. Two-thirds of samples were classified with high confidence, with accuracy >90%. In an independent blinded test-set of 83 samples, overall high-confidence accuracy reached 89%. Classification accuracy reached 100% for most tissue classes, including 131 metastatic samples. We further validated the utility of the miRNA biomarkers by quantitative RT-PCR using 65 additional blinded test samples. Our findings demonstrate the effectiveness of miRNAs as biomarkers for tracing the tissue of origin of cancers of unknown primary origin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Pimiento, J.M., Teso, D., Malkan, A., Dudrick, S.J. & Palesty, J.A. Cancer of unknown primary origin: a decade of experience in a community-based hospital. Am. J. Surg. 194, 833–7, discussion 837–8 (2007).
Shaw, P.H., Adams, R., Jordan, C. & Crosby, T.D. A clinical review of the investigation and management of carcinoma of unknown primary in a single cancer network. Clin. Oncol. (R. Coll. Radiol.) 19, 87–95 (2007).
Hainsworth, J.D. & Greco, F.A. Treatment of patients with cancer of an unknown primary site. N. Engl. J. Med. 329, 257–263 (1993).
Blaszyk, H., Hartmann, A. & Bjornsson, J. Cancer of unknown primary: clinicopathologic correlations. APMIS 111, 1089–1094 (2003).
Varadhachary, G.R., Abbruzzese, J.L. & Lenzi, R. Diagnostic strategies for unknown primary cancer. Cancer 100, 1776–1785 (2004).
Ramaswamy, S. et al. Multiclass cancer diagnosis using tumor gene expression signatures. Proc. Natl. Acad. Sci. USA 98, 15149–15154 (2001).
Bloom, G. et al. Multi-platform, multi-site, microarray-based human tumor classification. Am. J. Pathol. 164, 9–16 (2004).
Ma, X.J. et al. Molecular classification of human cancers using a 92-gene real-time quantitative polymerase chain reaction assay. Arch. Pathol. Lab. Med. 130, 465–473 (2006).
Talantov, D. et al. A quantitative reverse transcriptase-polymerase chain reaction assay to identify metastatic carcinoma tissue of origin. J. Mol. Diagn. 8, 320–329 (2006).
Tothill, R.W. et al. An expression-based site of origin diagnostic method designed for clinical application to cancer of unknown origin. Cancer Res. 65, 4031–4040 (2005).
Shedden, K.A. et al. Accurate molecular classification of human cancers based on gene expression using a simple classifier with a pathological tree-based framework. Am. J. Pathol. 163, 1985–1995 (2003).
Baskerville, S. & Bartel, D.P. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 11, 241–247 (2005).
Farh, K.K. et al. The widespread impact of mammalian microRNAs on mRNA repression and evolution. Science 310, 1817–1821 (2005).
Landgraf, P. et al. A Mammalian microRNA Expression Atlas Based on Small RNA Library Sequencing. Cell 129, 1401–1414 (2007).
He, L. et al. A microRNA polycistron as a potential human oncogene. Nature 435, 828–833 (2005).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005).
Volinia, S. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 103, 2257–2261 (2006).
Raver-Shapira, N. et al. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol. Cell 26, 731–743 (2007).
Bentwich, I. et al. Identification of hundreds of conserved and nonconserved human microRNAs. Nat. Genet. 37, 766–770 (2005).
Griffiths-Jones, S., Grocock, R.J., van Dongen, S., Bateman, A. & Enright, A.J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 34, D140–D144 (2006).
Xiao, C. et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 131, 146–159 (2007).
Hornstein, E. et al. The microRNA miR-196 acts upstream of Hoxb8 and Shh in limb development. Nature 438, 671–674 (2005).
Lee, Y.S., Kim, H.K., Chung, S., Kim, K.S. & Dutta, A. Depletion of human micro-RNA miR-125b reveals that it is critical for the proliferation of differentiated cells but not for the down-regulation of putative targets during differentiation. J. Biol. Chem. 280, 16635–16641 (2005).
Sempere, L.F. et al. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 5, R13 (2004).
Ein-Dor, L., Kela, I., Getz, G., Givol, D. & Domany, E. Outcome signature genes in breast cancer: is there a unique set? Bioinformatics 21, 171–178 (2005).
Paik, S. et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J. Clin. Oncol. 24, 3726–3734 (2006).
van de Vijver, M.J. et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl. J. Med. 347, 1999–2009 (2002).
Li, J. et al. Comparison of miRNA expression patterns using total RNA extracted from matched samples of formalin-fixed paraffin-embedded (FFPE) cells and snap frozen cells. BMC Biotechnol. 7, 36 (2007).
Parker, D.C. et al. Potential utility of uroplakin III, thrombomodulin, high molecular weight cytokeratin, and cytokeratin 20 in noninvasive, invasive, and metastatic urothelial (transitional cell) carcinomas. Am. J. Surg. Pathol. 27, 1–10 (2003).
Thomson, J.M., Parker, J., Perou, C.M. & Hammond, S.M. A custom microarray platform for analysis of microRNA gene expression. Nat. Methods 1, 47–53 (2004).
Shi, R. & Chiang, V.L. Facile means for quantifying microRNA expression by real-time PCR. Biotechniques 39, 519–525 (2005).
Thomson, J.M. et al. Extensive post-transcriptional regulation of microRNAs and its implications for cancer. Genes Dev. 20, 2202–2207 (2006).
Hino, K., Fukao, T. & Watanabe, M. Regulatory interaction of HNF1α to microRNA194 gene during intestinal epithelial cell differentiation. Nucleic Acids Symp. Ser. (Oxf.), 415–416 (2007).
van Duin, M. et al. High-resolution array comparative genomic hybridization of chromosome 8q: evaluation of putative progression markers for gastroesophageal junction adenocarcinomas. Cytogenet. Genome Res. 118, 130–137 (2007).
Acknowledgements
We thank Jung-Hwan Yoon of Seoul National University College of Medicine, Seoul, South Korea. N.R. dedicates this work to the memory of Yasha (Yaakov) Rosenfeld.
Author information
Authors and Affiliations
Contributions
R.A., A.A., I. Bentwich, Z.B., D.C., A.C. and I. Barshack directed research; N.R., R.A., E.M., S.R., Y.S., S.G., A.C. and I. Barshack designed experiments; N.S.-V., A.T., M.F., O.K., O.N., D.N., M.P., A.Y., B.S., S.P.-C., E.F. and I. Barshack provided samples and performed pathological analysis; E.M., M.Z., N.S., S.T., D.L. and S.G. performed experiments; N.R., R.A., S.R., Y.G. and E.S. developed algorithms; N.R., S.R., H.B. and Y.G. analyzed data; Y.S., A.L., N.T. and A.B.-A. provided bioinformatic and database support; N.R., R.A., A.C. and I. Barschack wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
All authors affiliated with Rosetta Genomics, except E.S., are full-time employees of Rosetta Genomics Ltd. and hold equity in the company, the value of which may be influenced by this publication. E.S. was engaged as an external consultant to Rosetta Genomics. O.N. is a paid consultant to Rosetta Genomics. All other authors declare that they have no competing interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Table 4 (PDF 515 kb)
Supplementary Table
Supplementary Table 1 (XLS 186 kb)
Supplementary Table
Supplementary Table 2 (XLS 25 kb)
Supplementary Table
Supplementary Table 3 (XLS 21 kb)
Supplementary Table
Supplementary Table 5 (XLS 29 kb)
Rights and permissions
About this article
Cite this article
Rosenfeld, N., Aharonov, R., Meiri, E. et al. MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol 26, 462–469 (2008). https://doi.org/10.1038/nbt1392
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1392
This article is cited by
-
Prognostic value of plasma microRNAs for non-small cell lung cancer based on data mining models
BMC Cancer (2024)
-
Role of microRNA-363 during tumor progression and invasion
Journal of Physiology and Biochemistry (2024)
-
RNA therapy
Experimental & Molecular Medicine (2023)
-
Expression analysis of circulating miR-22, miR-122, miR-217 and miR-367 as promising biomarkers of acute lymphoblastic leukemia
Molecular Biology Reports (2023)
-
A novel microRNA signature for the detection of melanoma by liquid biopsy
Journal of Translational Medicine (2022)