Abstract
Dichelobacter nodosus causes ovine footrot, a disease that leads to severe economic losses in the wool and meat industries. We sequenced its 1.4-Mb genome, the smallest known genome of an anaerobe. It differs markedly from small genomes of intracellular bacteria, retaining greater biosynthetic capabilities and lacking any evidence of extensive ongoing genome reduction. Comparative genomic microarray studies and bioinformatic analysis suggested that, despite its small size, almost 20% of the genome is derived from lateral gene transfer. Most of these regions seem to be associated with virulence. Metabolic reconstruction indicated unsuspected capabilities, including carbohydrate utilization, electron transfer and several aerobic pathways. Global transcriptional profiling and bioinformatic analysis enabled the prediction of virulence factors and cell surface proteins. Screening of these proteins against ovine antisera identified eight immunogenic proteins that are candidate antigens for a cross-protective vaccine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Thomas, J.H. The pathogenesis of footrot in sheep with reference to proteases of Fusiformis nodosus. Aust. J. Agric. Res. 15, 1001–1016 (1964).
Schwartzkoff, C.L. et al. The effects of antigenic competition on the efficacy of multivalent footrot vaccines. Aust. Vet. J. 70, 123–126 (1993).
Stewart, D.J. Footrot of sheep. in Footrot and Foot Abscess of Ruminants (Egerton, J.R., Yong, W.K. & Riffkin, G.G., eds.) 5–45, (CRC Press, Boca Raton, Florida, USA, 1989).
Rood, J.I., Stewart, D.J., Vaughan, J.A. & Dewhirst, F.E. in Bergey's Manual of Systematic Bacteriology Vol. 2: The Proteobacteria; Part B: The Gammaproteobacteria Edn. 2 (Brenner, D.J., Kreig, N.R. & Staley, J.T., eds.) 124–129 (Springer, New York, 2005).
Calza, L., Manfredi, R. & Chiodo, F. Infective endocarditis: a review of the best treatment options. Expert Opin. Pharmacother. 5, 1899–1916 (2004).
Kirkwood, J.K., Macgregor, S.K., Malnick, H. & Foster, G. Unusual mortality incidents in tit species (family Paridae) associated with the novel bacterium Suttonella ornithocola. Vet. Rec. 158, 203–205 (2006).
Kennan, R.M., Billington, S.J. & Rood, J.I. Electroporation-mediated transformation of the ovine footrot pathogen Dichelobacter nodosus. FEMS Microbiol. Lett. 169, 383–389 (1998).
Kennan, R.M., Dhungyel, O.P., Whittington, R.J., Egerton, J.R. & Rood, J.I. The Type IV fimbrial subunit gene (fimA) of Dichelobacter nodosus is essential for virulence, protease secretion, and natural competence. J. Bacteriol. 183, 4451–4458 (2001).
Giovannoni, S.J. et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science 309, 1242–1245 (2005).
Moran, N.A. Microbial minimalism: genome reduction in bacterial pathogens. Cell 108, 583–586 (2002).
Haring, V. et al. Delineation of the virulence-related locus (vrl) of Dichelobacter nodosus. Microbiology 141, 2081–2089 (1995).
Billington, S.J., Johnston, J.L. & Rood, J.I. Virulence regions and virulence factors of the ovine footrot pathogen, Dichelobacter nodosus. FEMS Microbiol. Lett. 145, 147–156 (1996).
Moses, E.K. et al. A multiple site-specific DNA-inversion model for the control of Omp1 phase and antigenic variation in Dichelobacter nodosus. Mol. Microbiol. 17, 183–196 (1995).
Rood, J.I. Genomic islands of Dichelobacter nodosus. Curr. Top. Microbiol. Immunol. 264, 47–60 (2002).
Cheetham, B.F. & Katz, M.E. A role for bacteriophages in the evolution and transfer of bacterial virulence determinants. Mol. Microbiol. 18, 201–208 (1995).
Parker, D. et al. Regulation of type IV fimbrial biogenesis in Dichelobacter nodosus. J. Bacteriol. 188, 4801–4811 (2006).
Claxton, P.D., Ribeiro, L.A. & Egerton, J.R. Classification of Bacteroides nodosus by agglutination tests. Aust. Vet. J. 60, 331–334 (1983).
Hobbs, M. et al. Organization of the fimbrial gene region of Bacteroides nodosus: class I and class II strains. Mol. Microbiol. 5, 543–560 (1991).
Sandkvist, M. Biology of type II secretion. Mol. Microbiol. 40, 271–283 (2001).
Riffkin, M.C., Wang, L.F., Kortt, A.A. & Stewart, D.J. A single amino-acid change between the antigenically different extracellular serine proteases V2 and B2 from Dichelobacter nodosus. Gene 167, 279–283 (1995).
Frey, J. & Kuhnert, P. RTX toxins in Pasteurellaceae. Int. J. Med. Microbiol. 292, 149–158 (2002).
Suzuki, T. & Sasakawa, C. Molecular basis of the intracellular spreading of Shigella. Infect. Immun. 69, 5959–5966 (2001).
Pizza, M. et al. Identification of vaccine candidates against serogroup B meningococcus by whole-genome sequencing. Science 287, 1816–1820 (2000).
Hess, J. et al. Listeria monocytogenes p60 supports host cell invasion by and in vivo survival of attenuated Salmonella typhimurium. Infect. Immun. 63, 2047–2053 (1995).
Adler, B. et al. Candidate vaccine antigens and genes in Pasteurella multocida. J. Biotechnol. 73, 83–90 (1999).
Yorgey, P., Rahme, L.G., Tan, M.W. & Ausubel, F.M. The roles of mucD and alginate in the virulence of Pseudomonas aeruginosa in plants, nematodes and mice. Mol. Microbiol. 41, 1063–1076 (2001).
Konstantinidis, K.T. & Tiedje, J.M. Trends between gene content and genome size in prokaryotic species with larger genomes. Proc. Natl. Acad. Sci. USA 101, 3160–3165 (2004).
Stover, C.K. et al. Complete genome sequence of Pseudomonas aeruginosa PA01, an opportunistic pathogen. Nature 406, 959–964 (2000).
Perez-Rueda, E. & Collado-Vides, J. The repertoire of DNA-binding transcriptional regulators in Escherichia coli K-12. Nucleic Acids Res. 28, 1838–1847 (2000).
Egerton, J.R. & Burrell, D.H. Prophylactic and therapeutic vaccination against ovine foot-rot. Aust. Vet. J. 46, 517–522 (1970).
Kennan, R.M., Dhungyel, O.P., Whittington, R.J., Egerton, J.R. & Rood, J.I. Transformation-mediated serogroup conversion of Dichelobacter nodosus. Vet. Microbiol. 92, 169–178 (2003).
Zhou, H. & Hickford, J.G. Extensive diversity in New Zealand Dichelobacter nodosus strains from infected sheep and goats. Vet. Microbiol. 71, 113–123 (2000).
Stewart, D.J., Clark, B.L., Emery, D.L., Peterson, J.E. & Fahey, K.J. A Bacteroides nodosus immunogen, distinct from the pilus, which induces cross-protective immunity in sheep vaccinated against footrot. Aust. Vet. J. 60, 83–85 (1983).
Stewart, D.J. et al. The protection given by pilus and whole cell vaccines of Bacteroides nodosus strain 198 against ovine foot-rot induced by strains of different serogroups. Aust. Vet. J. 62, 153–159 (1985).
Parker, D., Kennan, R.M., Myers, G.S., Paulsen, I. & Rood, J.I. Identification of a Dichelobacter nodosus ferric uptake regulator and determination of its regulatory targets. J. Bacteriol. 187, 366–375 (2005).
Cianciotto, N.P. & Fields, B.S. Legionella pneumophila mip gene potentiates intracellular infection of protozoa and human macrophages. Proc. Natl. Acad. Sci. USA 89, 5188–5191 (1992).
Leuzzi, R. et al. Ng-MIP, a surface-exposed lipoprotein of Neisseria gonorrhoeae, has a peptidyl-prolyl cis/trans isomerase (PPIase) activity and is involved in persistence in macrophages. Mol. Microbiol. 58, 669–681 (2005).
Fraser, C.M. et al. Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi. Nature 390, 580–586 (1997).
Tettelin, H., Radune, D., Kasif, S., Khouri, H. & Salzberg, S.L. Optimized multiplex PCR: efficiently closing a whole-genome shotgun sequencing project. Genomics 62, 500–507 (1999).
Bulach, D.M. et al. Genome reduction in Leptospira borgpetersenii reflects limited transmission potential. Proc. Natl. Acad. Sci. USA 103, 14560–14565 (2006).
Delcher, A.L., Phillippy, A., Carlton, J. & Salzberg, S.L. Fast algorithms for large-scale genome alignment and comparison. Nucleic Acids Res. 30, 2478–2483 (2002).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Howe, K., Bateman, A. & Durbin, R. QuickTree: building huge Neighbour-Joining trees of protein sequences. Bioinformatics 18, 1546–1547 (2002).
Thomas, J.H. A simple medium for the isolation and cultivation of Fusiformis nodosus. Aust. Vet. J. 34, 411–413 (1958).
Gardy, J.L. et al. PSORT-B: Improving protein subcellular localization prediction for Gram-negative bacteria. Nucleic Acids Res. 31, 3613–3617 (2003).
Nielsen, H., Engelbrecht, J., Brunak, S. & von Heijne, G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 10, 1–6 (1997).
Juncker, A.S. et al. Prediction of lipoprotein signal peptides in Gram-negative bacteria. Protein Sci. 12, 1652–1662 (2003).
Cabrita, L.D., Dai, W. & Bottomley, S.P. A family of E. coli expression vectors for laboratory scale and high throughput soluble protein production. BMC Biotechnol. 6, 12 (2006).
Acknowledgements
The research was supported by an Initiative for Future Agriculture and Food Systems Grant No. 2001-52100 11445 from the USDA Cooperative State Research, Education, and Extension Service, and by grants from the Australian Research Council. D.P was the recipient of an Australian Postgraduate Award and a Monash Faculty of Medicine, Nursing, and Health Sciences Postgraduate Excellence Award. We thank I. McPherson for technical assistance, C. Whitchurch for helpful discussions, B. Cheetham for providing unpublished information and helpful discussions, and the TIGR faculty, sequencing facility and informatics group for expert advice and assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Distribution of database hits to γ-proteobacteria, β-proteobacteria and other phylogenetic groups. (PDF 1132 kb)
Supplementary Fig. 2
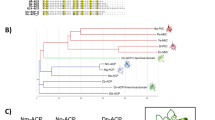
Bootstrapped maximum parsimony tree of representative sequenced species from the β- and γ-proteobacteria. (PDF 73 kb)
Supplementary Fig. 3
Graphical representation of genes displaying variability using comparative genomic hybridization. (PDF 5083 kb)
Supplementary Fig. 4
Comparative genomic locations of type IV fimbrial biogenesis genes in D. nodosus and P. aeruginosa. (PDF 83 kb)
Supplementary Fig. 5
Graphical representation of the D. nodosus outer membrane protein locus. (PDF 71 kb)
Supplementary Fig. 6
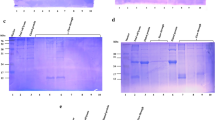
Immunoblots demonstrating recognition of D. nodosus proteins by pooled sera from experimentally infected sheet. (PDF 316 kb)
Supplementary Table 1
Characteristics of strains used in CGH analysis. (PDF 66 kb)
Supplementary Table 2
Comparison of selected metabolic capabilities between organisms with small genome sizes. (PDF 93 kb)
Supplementary Table 3
Differentially expressed genes of D. nodosus when grown on hoof agar. (PDF 155 kb)
Supplementary Table 4
Oligonucleotide primers used for QRT-PCR. (PDF 83 kb)
Supplementary Table 5
Statistical comparison of normal and atypical regions of nucleotide composition within the D. nodosus genome. (PDF 83 kb)
Rights and permissions
About this article
Cite this article
Myers, G., Parker, D., Al-Hasani, K. et al. Genome sequence and identification of candidate vaccine antigens from the animal pathogen Dichelobacter nodosus. Nat Biotechnol 25, 569–575 (2007). https://doi.org/10.1038/nbt1302
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1302
This article is cited by
-
Serogroups of Dichelobacter nodosus, the cause of footrot in sheep, are randomly distributed across England
Scientific Reports (2020)
-
Sites of persistence of Fusobacterium necrophorum and Dichelobacter nodosus: a paradigm shift in understanding the epidemiology of footrot in sheep
Scientific Reports (2019)
-
Characterization of two putative Dichelobacter nodosus footrot vaccine antigens identifies the first lysozyme inhibitor in the genus
Scientific Reports (2019)
-
Cloning, Expression, and Functional Characterization of Serine Protease Aprv2 from Virulent Isolate Dichelobacter nodosus of Indian Origin
Applied Biochemistry and Biotechnology (2016)
-
A recently introduced Dichelobacter nodosus strain caused an outbreak of footrot in Norway
Acta Veterinaria Scandinavica (2014)