Abstract
Xylose is a major constituent of plant lignocellulose, and its fermentation is important for the bioconversion of plant biomass to fuels and chemicals. Pichia stipitis is a well-studied, native xylose-fermenting yeast. The mechanism and regulation of xylose metabolism in P. stipitis have been characterized and genes from P. stipitis have been used to engineer xylose metabolism in Saccharomyces cerevisiae. We have sequenced and assembled the complete genome of P. stipitis. The sequence data have revealed unusual aspects of genome organization, numerous genes for bioconversion, a preliminary insight into regulation of central metabolic pathways and several examples of colocalized genes with related functions. The genome sequence provides insight into how P. stipitis regulates its redox balance while very efficiently fermenting xylose under microaerobic conditions.
Similar content being viewed by others
Main
Xylose is a five-carbon sugar abundant in hardwoods and agricultural residues1, so its fermentation is essential for the economic conversion of lignocellulose to ethanol2. Pichia stipitis Pignal (1967) is a haploid, homothallic, hemiascomycetous yeast3,4 that has the highest native capacity for xylose fermentation of any known microbe5. Fed batch cultures of P. stipitis produce almost 50 g/l of ethanol from xylose6 with yields of 0.35 to 0.44 g/g xylose (Fig. 1)7, and they can ferment hydrolysates at 80% of the maximum theoretical yield8.
P. stipitis Pignal (1967) is closely related to yeast endosymbionts of passalid beetles9 that inhabit and degrade white-rotted hardwood10. It forms yeast-like buds during exponential growth, hat-shaped spores and pseudomycelia (Fig. 2), uses all of the major sugars found in wood11 and transforms low-molecular weight lignin moieties12. P. stipitis genes have been used to engineer xylose metabolism in S. cerevisiae1, but regulation for ethanol production is problematic13. S. cerevisiae regulates fermentation by sensing the presence of glucose, whereas P. stipitis induces fermentative activity in response to oxygen limitation14,15. Increasing the P. stipitis fermentation rate could greatly improve its usefulness in commercial processes. Conversely, by using knowledge of this native xylose-fermenting yeast, researchers could improve xylose metabolism in S. cerevisiae.
We sequenced the P. stipitis genome to better understand its biology, metabolism and regulation. In analyzing the genome we discovered numerous genes for lignocellulose bioconversion (http://www.jgi.doe.gov/pichia).
Note: Supplementary information is available on the Nature Biotechnology website.
Results
The 15.4-Mbp genome of P. stipitis was sequenced using a shotgun approach and finished to high quality (<1 error in 100,000). The eight chromosomes range in size from 3.5 to 0.97 Mbp, as previously reported16. The finished chromosomes have only one gap in the centromere region of chromosome 1. The Joint Genome Institute (JGI) Annotation Pipeline predicted 5,841 genes (Table 1). A majority, 72%, have a single exon. Average gene density is 56%. Average gene, transcript and protein lengths are 1.6 kb, 1.5 kb and 493 amino acids, respectively. Expressed sequence tags (ESTs) support 40% of the predicted genes with 84% showing strong similarity to proteins in other fungi. Best bidirectional BLAST analysis of the gene models against the Debaryomyces hansenii genome identified putative orthologs for 84% of the P. stipitis genes. Additionally, analysis of conservation between the genomes of P. stipitis and D. hansenii at the DNA level using VISTA tools17 provided support for exons in 67.5% of the P. stipitis genes.
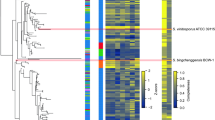
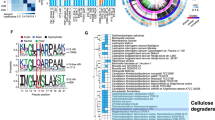
Protein function can be tentatively assigned to about 70% of the genes according to KOG (eukaryotic orthologous groups) classifications (Supplementary Fig. 1 online)18. Protein domains were predicted in 4,083 gene models. These include 1,712 distinct Pfam domains. A PhIGs (phylogenetically inferred groups19, http://phigs.org/) comparison of P. stipitis with eight other yeasts (Fig. 3)20 revealed 25 gene families representing 72 proteins specific to P. stipitis (Supplementary Table 1 online). P. stipitis and D. hansenii share 151 gene families that are not found in the other genomes. The P. stipitis gene set was missing 81 gene families (442 proteins) relative to the other yeast genomes in the analysis.
The most frequent domains include protein kinases, helicases, transporters (sugar and MFS) and domains involved in transcriptional regulation (fungal specific transcription factors, RNA recognition motifs and WD40 domains). A majority is shared with other hemiascomycota. These range from 1,534 common with Schizosaccharomyces pombe to 1,639 with D. hansenii. One of the few P. stipitis–specific domains (Supplementary Table 2 online) belongs to glycosyl hydrolase Family 10, a subgroup of cellulases and xylanases. Several gene families expanded in P. stipitis show some sequence similarity to hyphally regulated cell wall proteins, cell surface flocculins, agglutinin-like proteins and cytochrome p450 nonspecific monooxygenases. Members of these expanded families are poorly conserved and often occur near chromosome termini (within 35,000 bp).
Chromosomal segments that retain the ancestral gene groupings can be identified between P. stipitis and D. hansenii. A total of 263 orthology segments were found, encompassing 4,456 of the genes (10,950,900 bp) in the P. stipitis genome, and 4,689 genes (9,057,788 bp) in the D. hansenii genome. On average, each block in the P. stipitis genome encompasses 16.9 genes and is 41.6 kb in length. The largest of these orthologous chromosomal segments, which is 301.9 kb in length and encompasses 125 genes, is between P. stipitis chromosome 6 and D. hansenii chromosome F (Fig. 4). The rates of genomic rearrangement observed here are consistent with previously reported rates between D. hansenii and Candida albicans21.
P. stipitis uses the alternative yeast nuclear codon (12) that substitutes serine for leucine when CUG is specified22. A count of CUG usage showed 15,265 occurrences in 4,238 open reading frames (ORFs), or about 72% of all gene models. Nine out of the 21 ORFs having 18 or more CUGs in the gene model occurred at or near a terminus of chromosomes 4, 8, 7 or 1. All gene models having a large number of CUGs in the ORF were large (>2,500 bp), very large (>5,000 bp), repetitive, hypothetical or poorly defined.
P. stipitis possesses genes for a number of transporters that are similar to putative xylose transporters from Debaromyces hansenii (NCBI AAR06925) and Candida intermedia (GXF1, EMBL AJ937350; GXS1, EMBL AJ875406)23. C. intermedia GXF1 has the closest similarity to the previously described, closely related SUT1, SUT2 and SUT3 genes of P. stipitis24 and to the P. stipitis SUT4 gene, which was identified in the present genome sequence (Supplementary Fig. 2 online). Notably, SUT2 and SUT3 are each located very near the ends of their respective chromosomes. Available EST data do not show their expression.
All of the genes for xylose assimilation, the oxidative pentose phosphate pathway (PPP), glycolysis, the tricarboxylic acid cycle (TCA) and ethanol production were present in isoforms similar to those found in other yeasts (Fig. 5). Transcripts of GND1 are strongly induced by growth on xylose under both aerobic and oxygen-limiting conditions (Fig. 5). Transketolase (TKT1) is strongly induced on xylose, and is one of the most abundant transcripts in the cell under those conditions. Transcripts for PGI1, PFK1 and PFK2 were all induced on xylose under oxygen limitation, but were low in number under aerobic conditions (Fig. 5). Glyceraldehyde-3-phosphate dehydrogenase isoform 3 (TDH3), the gateway for glycolysis, was induced by oxygen limitation on both glucose and xylose. Transcripts for PDC1 and ADH1 were low in number on xylose under oxygen-limited conditions. The five NADP(H)-coupled alcohol dehydrogenases (ADH3, 4, 5, 6 and 7) could maintain cofactor balance between NADH and NADPH. Transcripts for mitochondrial isocitrate dehydrogenases (IDH1, IDH2) are elevated on xylose under oxygen-limited conditions, as are those for malate dehydrogenase (MDH1), fumarase (FUM1) and succinic dehydrogenase (SDH1). The transcript for 2-ketoglutarate dehydrogenase (KGD1), which generates NADH in the TCA cycle, was reduced during cultivation on xylose.
An NAD-specific glutamate dehydrogenase (GDH2), a glutamate decarboxylase (GAD2), and two NADP-dependent succinate semialdehyde dehydrogenases (UGA2, UGA22) constitute a bypass to convert α-ketoglutarate into succinate and NADH into NADPH when cells are growing on xylose. The NADH-specific GDH2 is elevated on xylose under oxygen limitation, whereas the NADPH-linked glutamate dehydrogenase 3 (GDH3) is not. The increased level of GDH2 could also account for the decreased level of KGD2 when cells are growing on xylose. Distinctly different sets of genes are strongly induced under oxygen-limited growth on glucose and xylose (Supplementary Table 3 online). On xylose, the transcript for fatty acid synthase 2 (FAS2) and the stearoyl-CoA desaturase, (OLE1) are strongly induced under oxygen limitation.
A Family 10 xylanase, XYN1, was found along with several Family 5 endo-1,4-β-glucanases or cellodextrinases (EGC1, EGC2 and EGC3). EGC2 is strongly expressed in cells growing on xylose. The three exo-1,3-β-glucosidases (EXG1, EXG2, EXG3) could help the beetle host digest fungal hyphae. The Family 17 soluble cell wall glucosidases (SCW4.1, SCW4.2 and SCW11) along with the Family 17 exo-1,3-β-glucanases (BGL2, BOT2), are most likely involved in cell wall expansion and growth. Family 3 β-glucosidases (BGL1-7) can have activity against cellobiose or xylobiose. Of the seven found in P. stipitis, BGL4 is most similar to classical cellobioses and BGL7 is expressed most when cells are growing on xylose (Supplementary Table 3). The Family 2 β-mannosidases (BMS1, MAN2) are probably responsible for the capacity of this yeast to grow on and ferment mannan oligosaccharides, but the endo-1,6-α-mannosidases (DCW1, DFG5) are likely involved in yeast cell wall expansion during growth, since they are present when cells are growing on either glucose or xylose. P. stipitis has four α-glucosidases (MAL6-9) and a Family 31 α-glucosidase/α-xylosidase (YIC1). Of these, only MAL8 was detected when cells were grown on xylose.
Five salicylate hydroxylases (NHG1.1, NHG 1.2, NHG2, NHG3, NHG4) are scattered throughout the genome. Only NHG2 shows conservation relative to D. hansenii. The rest of the genes and their surrounding loci have no identity to proteins found in other yeasts. The genome contains almost 60 ORFs that are identified as chitinases according to KOG classification. Only four (CHT1, CHT2, CHT3, CHT4) are likely to be involved in degradation of insect or fungal cell walls. The remaining models are mucin-like proteins. MUC1 appears four times in nearly identical copies, and segments exist in ∼25 copies, suggesting expansion through frequent duplication.
A gene for DUR1 (DUR1,2, urea amidolyase) is immediately adjacent to the urea transporter DUR3.1. In addition to DUR3.1 P. stipitis has two other genes for urea transport. DUR3.2 and DUR5.1. These are not found in any of the other yeasts with sequenced genomes. Multiple copies of similar transporters (e.g., DUR4, DUR5.2, DUR5.3, DUR8) are also found in P. stipitis.
β-glucosidases were often adjacent or proximal to genes with related functions (Supplementary Table 4 online). For example, on either side of the β-1,4 endoglucanase EGC2, one finds BGL5 and the probable hexose transporter, HXT2.4. EGC3 is adjacent to HXT2.1. BGL6 is adjacent to EGC1, and BGL3 is adjacent to SUT3, and BGL1 and HXT2.6 are adjacent to SUT2. Both of the putative P. stipitis β-mannosidases (BMS1, MAN2) are adjacent or proximal to putative sugar permeases (LAC3 and LAC2, respectively).
One of the most conspicuous examples of tandem genes with related functions is the MAL3 locus (Fig. 6). This site contains a maltose permease, MAL3, and the α-glucosidase, AGL1. Adjacent to MAL3 is MAL5, which is adjacent to YIC1, an α-glucosidase. Flanking this complex are a fungal transcriptional regulatory protein, SUC1.2, similar to MAL-activator proteins25, and a second putative fungal-specific regulatory protein, SUC1.4. Elsewhere in the genome, on chromosome 6, the α-glucosidase, MAL8, is immediately adjacent to the maltose permease, MAL4.
We identified a number of transposable elements using a composite library of fungal repeats26. The most abundant include long terminal repeat retrotransposons Tdh5, Tdh2, Tse5, pCal, most of which are present in D. hansenii27. Single copies of DNA mediated elements Ty1-I, Mariner-5 and Folyt1 were reported earlier in fungi28. Copies of the retrotransposon Tps5 show one well-defined locus on each chromosome.
Discussion
The CBS 6054 strain was isolated from insect larvae, and other yeast strains closely related to P. stipitis have been isolated from the guts of wood-inhabiting passalid beetles10, suggesting that this family of yeasts has evolved to inhabit an oxygen-limited environment rich in partially digested wood. The presence of numerous genes for endoglucanases and β-glucosidases, along with xylanase, mannanase and chitinase activities indicates that it could metabolize polysaccharides in the beetle gut. Various strains of P. stipitis have been reported to ferment cellobiose to ethanol29. Exo-1,4-cellobiohydrolases, which are responsible in part for the degradation of cellulose, produce cellobiose from cellulose and most endo-1,4-xylanases produce a mixture of xylose, xylobiose and xylotriose. β-glucosidases and β-xylosidase activities are therefore very useful traits when cellulose and hemicellulose saccharification is combined with fermentation.
Genes for xylose assimilation (XYL1, XYL2) were not expressed in the presence of glucose in the medium. GND1 and TKT1 were substantially elevated when growing on xylose, which reflects the increased activity of the PPP for xylose metabolism. PGI1, PFK1 and PFK2 were elevated most with cells growing on xylose under oxygen-limited conditions. Presumably elevated PGI1 is necessary to cycle fructose 6-phosphate (F6P) through the oxidative PPP whereas PFK1 and PFK2 take F6P into glycolysis. GLK1 was elevated in cells growing on xylose aerobically, which could reflect carbon catabolite de-repression.
Excess NADH is generated during growth on xylose1, which necessitates cofactor regeneration. KGD2, which forms NADH in the TCA cycle, was three times higher in cells growing on glucose over those on xylose. Gdh2 consumes NADH while generating NAD+, and leads into a pathway that eventually consumes NADH while generating NADPH. A similar pathway was previously engineered in S. cerevisiae to reduce cofactor imbalances when cells are growing on xylose30, but it appears to exist naturally in P. stipitis.
P. stipitis has a complete mitochondrial respiration system including a SHAM-sensitive terminal alternative oxidase (AOX1 or STO1)31, and NADH dehydrogenase Complex I, both of which are lacking in S. cerevisiae. Without Complex I, S. cerevisiae has less capacity for ATP generation through oxidative phosphorylation. AOX1 is not proton translocating, but it could scavenge for oxygen to balance cofactors. The EST data do not provide evidence of a role for AOX1 in xylose fermentation.
The abundance of genes for NADP(H) oxidoreductase reactions suggests that P. stipitis is capable of various strategies for balancing NAD and NADP-specific cofactors. Not least among these is FAS2, which appears to be highly active when cells are growing under oxygen-limited conditions on xylose. Fas2 synthesizes long chain acyl-CoA precursors of fatty acids that could serve as a reductant sink. Transcripts for fatty acid synthesis including OLE1 and, particularly, FAS2 were elevated in oxygen-limited, xylose-grown cells, indicating that reductant is channeled into lipid synthesis under oxygen limitation.
Colocation of genes having different but related functions (e.g., a permease with a hydrolase for maltose) occurs with high frequency in P. stipitis, but it is not confined to this yeast. For example, the association between urea permease and urea amidolyase is found throughout the sequenced yeast genomes. DUR1,2 is immediately downstream of an ortholog of DUR3 in a wide variety of ascomycetous yeasts including D. hansenii, Candida glabrata, Kluyveromyces lactis, S. cerevisiae and Yarrowia lipolytica, and there are several examples of the MAL3 locus in other yeasts. Other proximal associations, such as those between BGL and SUT genes, appear to be unique to P. stipitis.
Proximal orthologs with strong similarity to EGC2 (DEHA0G07095g), BGL5 (DEHA0G07183g) and HXT2.4, (DHEA0G07117 and DHEA0g07139) are also found in D. hansenii, but their locations and arrangement are different in that yeast, so although the orientation and number of these genes change, functional relationships remain. The closest sequenced relative to P. stipitis, D. hansenii, does not have similar correlations between BGL and SUT genes even though it possesses six putative β-glucosidases and three SUT genes very similar to those found in P. stipitis, which suggests that these proximal relationships evolved through selective pressure in P. stipitis. Two out of the six genes of the MAL3 locus appear to be conserved in C. albicans, and four out of the six are conserved in D. hansenii.
Members of multigene families are found near S. cerevisiae telomeres and are repeated elsewhere in the genome. It has been proposed that the concentration of multigene families in the telomere-adjacent regions reflects a recombination-mediated dispersal mechanism32. In P. stipitis, some genes at chromosome termini have orthologs proximal to functionally related genes at sites deeper within the chromosomes, so a similar mechanism could be working here.
Genes in telomeric regions might be under less selective pressure because of silencing. In S. cerevisiae the COMPASS histone methyltransferase carries out telomeric silencing of gene expression33. The P. stipitis genome contains a COMPASS homolog (SET1), so the same mechanism might be functioning here. Without selective pressure, genes in the telomeric regions could diverge more rapidly. We noted that genes occurring at chromosome termini often had a high frequency of CUG usage, which might indicate genetic drift.
The proximal location of glucosidases to sugar transporters and adjacency of urea amidolyase to urea permease suggest that these loci might be coregulated. In S. cerevisiae, genes for α-glucosidase and maltose permease are adjacent. Each complete MAL locus consists of maltose permease, maltase and a transcription activator. The MAL loci each map to the telomeric region of a different chromosome34.
In eukaryotes, coregulated genes distal from one another can be physically colocalized in nuclear 'transcriptional factories'. Osborne et al. proposed that linked genes are more likely to occupy a transcriptional factory than genes in trans. In the human transcriptional map, genes with increased expression occur in gene-dense regions35. Adjacent eukaryotic genes are more frequently coexpressed than is expected by chance and coexpressed neighboring genes are often functionally related. For example, in Arabidopsis thaliana, 10% of the genes occur in 266 groups of large, coexpressed, chromosomal regions distributed throughout the genome36. One published model37 encapsulates the advantages of proximal colocation of actively transcribed genes: the concentration of RNA polymerase II is 1,000-fold higher in a transcription factory than in the whole nucleus; modifications occurring during transcription leave the promoter open to new transcript initiation; after being released at the termination, promoters in the vicinity of a transcription factory are more likely to encounter machinery for transcriptional initiation again. These factors would all favor survival of strains in which genes with related function and regulatory features would be colocated in the genome.
The P. stipitis genome is endowed with numerous genes and physiological features enabling it to ferment a wide variety of sugars derived from lignocellulose, including a high capacity for using cellobiose and other oligomers. We discerned structural features such as genes with related functions proximal to one another that suggest the combined gene activities enhance survival. The genes can occur separately, but proximal location could affect their mutual function and the probability of co-inheritance. If some gene families persist in multiple copies simply from the advantage of higher transcript levels, then evolution toward higher promoter strength would be sufficient. Their presence in multiple copies suggests multiple functions. If chromosomal colocation affects expression, this would have implications with respect to the design and placement of genes for metabolic pathway engineering.
Methods
Yeast strain.
Pichia stipitis Pignal (1967), synonym Yamadazyma stipitis (Pignal) Bilon-Grand (1989), (NRRL Y-11545 = ATCC 58785 = CBS 6054 = IFO 10063) was obtained as a lyophilized powder. It was revived and streaked on yeast extract, peptone, dextrose (YPD) agar to obtain isolated colonies. A single colony was transferred to 150 ml of YPD broth. To test for contamination, the overnight culture was observed under the microscope and streaked in both YPD and LB plates. For fermentation studies, cells were grown in 125-ml Erlenmeyer flasks containing 50 ml of 1.67 g/l yeast nitrogen base (YNB) with 2.27 g/l urea and 80 g/l xylose. The YNB and urea solutions were filter sterilized in a 20 × solution and added to the sugar, which was sterilized separately by autoclaving. For mRNA preparation, cells were grown in YPD, which was prepared as described38 except that sugars were autoclaved separately from the basal medium. Yeast, peptone, xylose (YPX) was similar to YPD but replaced dextrose with xylose. Preparation of mRNA was by the method previously described22.
DNA preparation.
Yeast genomic DNA was prepared following a published protocol39. Two extra phenol:chloroform/chloroform extractions and ethanol precipitation were carried out. To prevent shredding of the DNA, the sample was not vortexed. The final gDNA concentration was 500 ng/μl as determined by optical density at 260 nm.
cDNA library construction and sequencing.
P. stipitis CBS 6054 was grown at 30 °C in 200 ml of either YPD or YPX in either a 2.8 l flask shaken at 300 r.p.m. or a 500 ml flask shaken at 50 r.p.m. Aerobic cultures were inoculated with a low cell density (0.025 mg/ml), shaken at 200 r.p.m. and harvested at a cell density of less than 0.5 mg/ml. Oxygen-limited cultures were inoculated with a high cell density (2.5 mg/ml), shaken at 100 r.p.m. and harvested at 5 mg/ml. Cells were collected by centrifugation at 4 °C and 9,279g. Cells were suspended in water and centrifuged at 835g for 5 min. Cells were then frozen in liquid N2. Poly A+ RNA was isolated from total RNA for all four P. stipitis samples using the Absolutely mRNA Purification kit (Stratagene). cDNA synthesis and cloning was a modified procedure based on the SuperScript plasmid system with Gateway technology for cDNA synthesis and cloning (Invitrogen). We used 1–2 μg of poly A+ RNA, reverse transcriptase SuperScript II (Invitrogen) and oligo dT primer (5′-GACTAGTTCTA GATCGCGAGCGGCCGCCC TTTTTTTTTTTTTTT-3′) to synthesize first-strand cDNA. Second-strand synthesis was performed with Escherichia coli DNA ligase, polymerase I and RNaseH followed by end repair using T4 DNA polymerase. The SalI adaptor (5′-TCGACC CACGCGTCCG and 5′-CGGACGCGTGGG) was ligated to the cDNA, digested with NotI (NEB), and subsequently size selected by gel electrophoresis (1.1% agarose). Size ranges of cDNA were cut out of the gel (Low insert size: 600–1.2 kb; Medium insert size: 1.2 kb–2 kb; High insert size: >2 kb) and directionally ligated into the SalI- and NotI-digested vector pCMVsport6 (Invitrogen). ElectroMAX T1 DH10B cells were transformed by the ligation (Invitrogen).
Library quality was first assessed by PCR amplification of the cDNA inserts of 20 clones with the primers M13-F (5′-GTAAAACGACGGCCAGT-3′) and M13-R (5′-AGGAAACAGCTATGACCAT-3′) to determine insert rate. Clones for each library were inoculated into 384-well plates (Nunc) and grown in LB for 18 h at 37 °C. DNA template for each clone was prepared by rolling circle amplification and sequenced using primers (FW: 5′-ATTTAGGTGACACTA TAGAA-3′ and RV 5′-TAATACGACTCACTATAGGG-3′), using Big Dye chemistry (Applied Biosystems). The average read length and pass rate were 753 (Q20 bases) and 96%, respectively.
EST sequence processing and assembly.
The JGI EST Pipeline begins with the cleanup of DNA sequences derived from the 5′and 3′ end reads from a library of cDNA clones. The Phred software40 is used to call the bases and generate quality scores. Vector, linker, adaptor, poly-A/T and other artifact sequences are removed using the Cross_match software40, and an internally developed short pattern finder. Low-quality regions of the read are identified using internally developed software, which masks regions with a combined quality score of <15. The longest high-quality region of each read is used as the EST. ESTs shorter than 150 bp are removed from the data set. ESTs containing common contaminants such as E. coli, common vectors and sequencing standards are also removed from the data set. EST Clustering is performed ab initio, based on alignments between each pair of trimmed, high-quality ESTs. Pair-wise EST alignments are generated using the Malign software (Chapman J., personal communication, JGI), a modified version of the Smith-Waterman algorithm41,42, which was developed at the JGI for use in whole-genome shotgun assembly. ESTs sharing an alignment of at least 98% identity and 150 bp overlap are assigned to the same cluster. These are relatively strict clustering cutoffs, and are intended to avoid placing divergent members of gene families in the same cluster. However, this could also have the effect of separating splice variants into different clusters. Optionally, ESTs that do not share alignments are assigned to the same cluster, if they are derived from the same cDNA clone. EST cluster consensus sequences were generated by running the Phrap software40 on the ESTs comprising each cluster. All alignments generated by malign are restricted such that they will always extend to within a few bases of the ends of both ESTs. Therefore, each cluster looks more like a 'tiling path' across the gene, which matches well with the genome-based assumptions underlying the Phrap algorithm. Additional improvements were made to the Phrap assemblies by using the 'forcelevel 4' option, which decreases the chances of generating multiple consensus sequences for a single cluster, where the consensus sequences differ only by sequencing errors.
Genome assembly.
The initial data set was derived from four whole-genome shotgun (WGS) libraries: one with an insert size of 3 kb, two with insert sizes of 8 kb, and one with an insert size of 35 kb. The reads were screened for vector using Cross_match, then trimmed for vector and quality. Reads shorter than 100 bases after trimming were then excluded. The data were assembled using release 1.0.1b of Jazz, a WGS assembler developed at the JGI43. A word size of 14 was used for seeding alignments between reads. The unhashability threshold was set to 50, preventing words present in more than 50 copies in the data set from being used to seed alignments. A mismatch penalty of −30.0 was used, which will tend to assemble together sequences that are more than ∼97% identical. The genome size and sequence depth were initially estimated to be 16.5 MB and 9.3, respectively. The assembly contained 394 scaffolds, with 16.4 MB of sequence, of which 4.5% was gap. The scaffold N/L50 was 5/1.46 MB, whereas the contig N/L50 was 21/262 kb. The sequence depth derived from the assembly was 8.77 ± 0.05.
Gap closure and finishing.
To perform finishing, the P. stipitis whole genome shotgun assembly was broken down into scaffold size pieces and each scaffold piece reassembled with Phrap. These scaffold pieces were then finished using our Phred/Phrap/Consed pipeline. Initially all low-quality regions and gaps were targeted with computationally selected sequencing reactions completed with 4:1 BigDye terminator/dGTP chemistry (Applied Biosystems). These automated rounds included resequencing plasmid subclones and walking on plasmid subclones or fosmids using custom primers. After completion of the automated rounds, a trained finisher manually inspected each assembly. Further reactions were than manually selected to complete the genome. These reactions included additional resequencing reactions and custom primer walks on plasmid subclones or fosmids. Again the reactions were completed using 4:1 BigDye terminator: dGTP chemistry. Smaller repeats in the sequence were resolved by transposon-hopping 8-kb plasmid clones. Fosmid clones were shotgun sequenced and finished to fill large gaps, resolve larger repeats or to resolve chromosome duplications and extend into chromosome telomere regions. After completion, each assembly was validated by an independent quality assessment. This examination included a visual examination of subclone paired ends using Orchid (http://www-shgc.stanford.edu/informatics/orchid.html), and visual inspection of high-quality discrepancies and all remaining low-quality areas. All available EST resources were also placed on the assembly to ensure completeness. All finished chromosomes are estimated to have an error rate of less than 1 in 100,000 bp.
Gene prediction and annotation.
The JGI Annotation Pipeline combines a suite of gene prediction and annotation methods. Gene prediction methods used for analysis of the P. stipitis genome include ab initio Fgenesh44, homology-based Fgenesh+ (http://www.softberry.com/) and Genewise45, and an EST-based method estExt (unpublished data). Predictions from each of the methods were taken to produce 'the best' single gene model per every locus. The best model was determined on the basis of similarity to GenBank proteins and EST support. Every predicted gene was annotated using Double Affine Smith-Waterman alignments (http://www.timelogic.com/) with Swissprot and KEGG proteins. Protein domains were predicted using InterProScan46,47 against various domain libraries (Prints, Prosite, PFAM, ProDom, SMART). Individual annotations have then been summarized according to Gene Ontology48, KOGs18 and KEGG metabolic pathways49.
Phylogenetic tree reconstruction of sequenced fungal genomes.
A multiple sequence alignment of 94 single-copy genes present in 26 taxa was constructed using the MUSCLE 3.52 program50, trimmed using Gblocks 0.91b and was used as input for the maximum likelihood tree reconstruction program PHYML (four rate categories, gamma + invariants, 100 bootstrap replicates) resulting in a fully resolved tree with all but one node having bootstrap values of 100. Figure 4 represents the portion of the tree describing relationships between the genomes of interest for this analysis.
Comparative analysis of the six yeast genomes.
Comparisons of the phylic patterns of gene family distributions of P. stipitis and five hemi-ascomycete yeasts (P. stipitis, S. cerevisiae, C. glabrata, K. lactis, D. hansenii and Y. lipolytica) were done using the PhIGs orthology database19. The PhIGs resource generated clusters of genes at each node on the evolutionary tree representing the descendents from a single ancestral gene existing at that node. This allows for the comparisons of the presence/absence patterns of gene families across the six species avoiding confusion from paralogous genes. The set of 3,209 genes determined to be orthologous from the PhIGs19 analysis were used to link regions between the two genomes that represent orthologous chromosomal segments with a minimum of four linking genes that are uninterrupted by other orthology segments in either genome. In this analysis, gene families specific to a single species are defined as those having a minimum of two family members.
Expression analysis.
To enable complete sampling of the expressed genes, we generated four separate EST libraries by growing cells on glucose or xylose under aerobic or oxygen-limited conditions. A set of 19,635 P. stipitis ESTs was sequenced from the four libraries and clustered into 4,085 consensus sequences. We mapped 94% (3,839) of the clusters to the genome and the numbers of hits for each consensus cluster was used to estimate EST frequency under each growth condition. An absolute majority of unplaced ESTs had problems with the sequences so the data indicate completeness and accurateness of genome assembly. Only 44% of the transcripts were represented by more than one EST cluster-hit under any one of the four growth conditions. The cluster-hit enumeration represents only a single biological sample for each of the four conditions, so these observations must be interpreted with care and be limited to the 200–400 most abundant gene models in which at least one transcript was recovered under each of the four conditions. However, the relative abundances of these ESTs under each of the four conditions provided a preliminary expression analysis.
Accession codes.
P. stipitis genome assembly and annotations have been deposited at DDBJ/EMBL/GenBank under the following accession numbers:, chr 1.1 and chr 1.2, AAVQ00000000 and AAVQ01000000; chr_2, CP000496; chr_3, CP000497; chr_4, CP000498; chr_5, CP000499; chr_6, CP000500; chr_7, CP000501; chr_8, CP000502.
Accession codes
Accessions
GenBank/EMBL/DDBJ
References
Jeffries, T.W. Engineering yeasts for xylose metabolism. Curr. Opin. Biotechnol. 17, 320–326 (2006).
Saha, B.C., Dien, B.S. & Bothast, R.J. Fuel ethanol production from corn fiber - Current status and technical prospects. Appl. Biochem. Biotechnol. 70–2, 115–125 (1998).
Kurtzman, C.P. Candida shehatae–genetic diversity and phylogenetic relationships with other xylose-fermenting yeasts. Antonie Van Leeuwenhoek 57, 215–222 (1990).
Melake, T., Passoth, V.V. & Klinner, U. Characterization of the genetic system of the xylose-fermenting yeast Pichia stipitis. Curr. Microbiol. 33, 237–242 (1996).
van Dijken, J.P., van den Bosch, E., Hermans, J.J., de Miranda, L.R. & Scheffers, W.A. Alcoholic fermentation by 'non-fermentative' yeasts. Yeast 2, 123–127 (1986).
du Preez, J.C., van Driessel, B. & Prior, B.A. Ethanol tolerance of Pichia stipitis and Candida shehatae strains in fed-batch cultures at controlled low dissolved-oxygen levels. Appl. Microbiol. Biotechnol. 30, 53–58 (1989).
Hahn-Hägerdal, B. & Pamment, N. Microbial pentose metabolism. Appl. Biochem. Biotechnol. 113–16, 1207–1209 (2004).
Nigam, J.N. Ethanol production from wheat straw hemicellulose hydrolysate by Pichia stipitis. J. Biotechnol. 87, 17–27 (2001).
Nardi, J.B. et al. Communities of microbes that inhabit the changing hindgut landscape of a subsocial beetle. Arth. Struct. Dev. 35, 57–68 (2006).
Suh, S.O., Marshall, C.J., McHugh, J.V. & Blackwell, M. Wood ingestion by passalid beetles in the presence of xylose-fermenting gut yeasts. Mol. Ecol. 12, 3137–3145 (2003).
Lee, H., Biely, P., Latta, R.K., Barbosa, M.F.S. & Schneider, H. Utilization of xylan by yeasts and its conversion to ethanol by Pichia stipitis strains. Appl. Environ. Microbiol. 52, 320–324 (1986).
Targonski, Z. Biotransformation of lignin-related aromatic-compounds by Pichia stipitis Pignal. Zentralbl. Mikrobiol 147, 244–249 (1992).
Jin, Y.S., Laplaza, J.M. & Jeffries, T.W. Saccharomyces cerevisiae engineered for xylose metabolism exhibits a respiratory response. Appl. Environ. Microbiol. 70, 6816–6825 (2004).
Passoth, V., Cohn, M., Schafer, B., Hahn-Hägerdal, B. & Klinner, U. Analysis of the hypoxia-induced ADH2 promoter of the respiratory yeast Pichia stipitis reveals a new mechanism for sensing of oxygen limitation in yeast. Yeast 20, 39–51 (2003).
Klinner, U., Fluthgraf, S., Freese, S. & Passoth, V. Aerobic induction of respiro-fermentative growth by decreasing oxygen tensions in the respiratory yeast Pichia stipitis. Appl. Microbiol. Biotechnol. 67, 247–253 (2005).
Passoth, V., Hansen, M., Klinner, U. & Emeis, C.C. The electrophoretic banding pattern of the chromosomes of Pichia stipitis and Candida shehatae. Curr. Genet. 22, 429–431 (1992).
Mayor, C. et al. VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16, 1046–1047 (2000).
Koonin, E.V. et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 5 (2) Art. No. R7 (2004).
Dehal, P.S. & Boore, J.L. A phylogenomic gene cluster resource: the Phylogenetically Inferred Groups (PhIGs) database. BMC Bioinformatics 7 Art. No. 201 (2006).
Dujon, B. et al. Genome evolution in yeasts. Nature 430, 35–44 (2004).
Fischer, G., Rocha, E.P.C., Brunet, F., Vergassola, M. & Dujon, B. Highly variable rates of genome rearrangements between hemiascomycetous yeast Lineages. PLoS Genet. 2, 253–261 (2006).
Laplaza, J.M., Torres, B.R., Jin, Y.S. & Jeffries, T.W. Sh ble and Cre adapted for functional genomics and metabolic engineering of Pichia stipitis. Enzyme Microb. Technol. 38, 741–747 (2006).
Leandro, M.J., Goncalves, P. & Spencer-Martins, I. Two glucose/xylose transporter genes from the yeast Candida intermedia: first molecular characterization of a yeast xylose/H + symporter. Biochem. J. (2006).
Weierstall, T., Hollenberg, C.P. & Boles, E. Cloning and characterization of three genes (SUT1–3) encoding glucose transporters of the yeast Pichia stipitis. Mol. Microbiol. 31, 871–883 (1999).
Chow, T.H., Sollitti, P. & Marmur, J. Structure of the multigene family of MAL loci in Saccharomyces. Mol. Gen. Genet. 217, 60–69 (1989).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Neuveglise, C., Feldmann, H., Bon, E., Gaillardin, C. & Casaregola, S. Genomic evolution of the long terminal repeat retrotransposons in hemiascomycetous yeasts. Genome Res. 12, 930–943 (2002).
Daboussi, M.J. & Capy, P. Transposable elements in filamentous fungi. Annu. Rev. Microbiol. 57, 275–299 (2003).
Parekh, S.R., Parekh, R.S. & Wayman, M. Fermentation of xylose and cellobiose by Pichia stipitis and Brettanomyces clausenii. Appl. Biochem. Biotechnol. 18, 325–338 (1988).
Grotkjaer, T., Christakopoulos, P., Nielsen, J. & Olsson, L. Comparative metabolic network analysis of two xylose fermenting recombinant Saccharomyces cerevisiae strains. Metab. Eng. 7, 437–444 (2005).
Shi, N.Q., Cruz, J., Sherman, F. & Jeffries, T.W. SHAM-sensitive alternative respiration in the xylose-metabolizing yeast Pichia stipitis. Yeast 19, 1203–1220 (2002).
Zakian, V.A. Structure, function, and replication of Saccharomyces cerevisiae telomeres. Annu. Rev. Genet. 30, 141–172 (1996).
Krogan, N.J. et al. COMPASS, a histone H3 (lysine 4) methyltransferase required for telomeric silencing of gene expression. J. Biol. Chem. 277, 10753–10755 (2002).
Vidgren, V., Ruohonen, L. & Londesborough, J. Characterization and functional analysis of the MAL and MPH loci for maltose utilization in some ale and lager yeast strains. Appl. Environ. Microbiol. 71, 7846–7857 (2005).
Osborne, C.S. et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat. Genet. 36, 1065–1071 (2004).
Zhan, S., Horrocks, J. & Lukens, L.N. Islands of co-expressed neighbouring genes in Arabidopsis thaliana suggest higher-order chromosome domains. Plant J. 45, 347–357 (2006).
Bartlett, O. et al. Specialized transcription factories. in Transcription vol. 73, 67–75, (Portland Press Ltd., London, 2006).
Kaiser, C., Michaelis, S. & Mitchell, A. . Methods in Yeast Genetics (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1994).
Burke, D., Dawson, D. & Stearns, T. Methods in Yeast Genetics: a Cold Spring Harbor Laboratory Course Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 2000).
Ewing, B. & Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194 (1998).
Smith, T.F. & Waterman, M.S. Overlapping genes and information theory. J. Theor. Biol. 91, 379–380 (1981).
Smith, T.F. & Waterman, M.S. Identification of common molecular subsequences. J. Mol. Biol. 147, 195–197 (1981).
Aparicio, S. et al. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297, 1301–1310 (2002).
Salamov, A.A. & Solovyev, V.V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000).
Birney, E. & Durbin, R. Using GeneWise in the Drosophila annotation experiment. Genome Res. 10, 547–548 (2000).
Zdobnov, E.M. & Apweiler, R. InterProScan–an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847–848 (2001).
Mulder, N.J. et al. InterPro, progress and status in 2005. Nucleic Acids Res. 33, D201–D205 (2005).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, D277–D280 (2004).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Acknowledgements
This work was performed under the auspices of the US Department of Energy's Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Livermore National Laboratory under Contract No. W-7405-Eng-48; Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231; Los Alamos National Laboratory under contract No. W-7405-ENG-36; Stanford University under contract No. DE-FC02-99ER62873, and by the US, Forest Service, Forest Products Laboratory. The authors are grateful to C.P. Kurtzman of the USDA ARS Culture Collection (NRRL) for providing the P. stipitis stock culture, to W. Huang, G. Werner and his group of the JGI for engineering support of annotation, to A. Polyakov and I. Dubchak of the JGI for VISTA analysis, to A. Darling for advice and support in MAUVE analysis, W. R. Kenealy, T. A. Kuster and Mark Davis of the USDA Forest Products Laboratory for carrying out continuous culture studies, providing photomicrographs and analyzing fermentation products, Samuel Pitluck and Kemin Zhou of JGI for assistance with the GenBank submission and to James Cregg, and Lisbeth Olsson and Jennifer Headman Van Vleet for critical readings of early drafts.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
After initiating the genomic sequencing study but prior to submitting the manuscript for publication, on December 15, 2006, we entered into a contract with Xylose Technologies Inc. (XTI, a subsidiary of Xethanol Corp.) to develop improved strains of Pichia stipitis for xylitol production. We currently receive funding from that source, and we are currently negotiating for a renewal of that contract, which will extend our studies to improved strains for ethanol production from xylose. Publication of this manuscript could have both positive and negative effects on that research contract and on the company. Negative effects could result by making a great deal of very useful information available to competitors. Positive effects could result from a perception that having the genetic sequence will enhance strain development.
Supplementary information
Supplementary Fig. 1
Graphical representation of gene models as classified according to KOG categories and the results as tallied on the JGI Pichia stipitis website. (PDF 465 kb)
Supplementary Fig. 2
Phylogenetic tree of selected sugar transporters from Pichia stipitis compared to putative sugar transporters from other yeast sources. (PDF 1065 kb)
Supplementary Table 1
Functional domain analysis and comparison to protein domains in eight other fungi. (DOC 65 kb)
Supplementary Table 2
Phyletic patterns of five yeast protein families. (DOC 56 kb)
Supplementary Table 3
Relative expression of most abundant P. stipitis transcripts under four growth conditions. (DOC 406 kb)
Supplementary Table 4
Proximal genes with related functions. (DOC 89 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike license (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar license.
About this article
Cite this article
Jeffries, T., Grigoriev, I., Grimwood, J. et al. Genome sequence of the lignocellulose-bioconverting and xylose-fermenting yeast Pichia stipitis. Nat Biotechnol 25, 319–326 (2007). https://doi.org/10.1038/nbt1290
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1290
This article is cited by
-
Xylose and shikimate transporters facilitates microbial consortium as a chassis for benzylisoquinoline alkaloid production
Nature Communications (2023)
-
Understanding the Dynamics of the Saccharomyces cerevisiae and Scheffersomyces stipitis Abundance in Co-culturing Process for Bioethanol Production from Corn Stover
Waste and Biomass Valorization (2023)
-
Dealing with Environmental Fluctuations: Diversity of Potassium Uptake Systems Across the Three Domains of Life
Journal of Plant Growth Regulation (2023)
-
Identification of traits to improve co-assimilation of glucose and xylose by adaptive evolution of Spathaspora passalidarum and Scheffersomyces stipitis yeasts
Applied Microbiology and Biotechnology (2023)
-
Xylose Metabolism in Bioethanol Production: Saccharomyces cerevisiae vs Non-Saccharomyces Yeasts
BioEnergy Research (2022)