Abstract
Direct physical information that describes where transcription factors, nucleosomes, modified histones, RNA polymerase II and other key proteins interact with the genome provides an invaluable mechanistic foundation for understanding complex programs of gene regulation. We present a method, joint binding deconvolution (JBD), which uses additional easily obtainable experimental data about chromatin immunoprecipitation (ChIP) to improve the spatial resolution of the transcription factor binding locations inferred from ChIP followed by DNA microarray hybridization (ChIP-Chip) data. Based on this probabilistic model of binding data, we further pursue improved spatial resolution by using sequence information. We produce positional priors that link ChIP-Chip data to sequence data by guiding motif discovery to inferred protein-DNA binding sites. We present results on the yeast transcription factors Gcn4 and Mig2 to demonstrate JBD's spatial resolution capabilities and show that positional priors allow computational discovery of the Mig2 motif when a standard approach fails.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000).
Lieb, J., Liu, X., Botstein, D. & Brown, P. Promoter-specific binding of Rap1 revealed by genome-wide maps of protein-DNA association. Nat. Genet. 28, 327–324 (2001).
Iyer, V. et al. Genomic binding sites of the yeast cell-cycle transcription factors SBF and MBF. Nature 409, 533–538 (2001).
Simon, I. et al. Serial regulation of transcriptional regulators in the yeast cell cycle. Cell 106, 697–708 (2001).
Lee, T. et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298, 799–804 (2002).
Horak, C. et al. GATA-1 binding sites mapped in the betaglobin locus by using mammalian ChIP-chip analysis. Proc. Natl. Acad. Sci. USA 99, 2924–2929 (2002).
Weinmann, A., Yan, P., Oberley, M., Huang, T. & Farnham, P. Isolating human transcription factor targets by coupling chromatin immunoprecipitation and CpG island microarray analysis. Genes Dev. 16, 235–244 (2002).
Li, Z. et al. A global transcriptional regulatory role for c-Myc in Burkitts lymphoma cells. Proc. Natl. Acad. Sci. USA 100, 8164–8169 (2003).
Wells, J., Yan, P., Cechvala, M., Huang, T. & Farnham, P. Identification of novel pRb binding sites using CpG microarrays suggests that E2F recruits pRb to specific genomic sites during S phase. Oncogene 22, 1445–1460 (2003).
Harbison, C.T. et al. Transcriptional regulatory code of a eukaryotic genome. Nature 431, 99–104 (2004).
Cawley, S. et al. Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116, 499–509 (2004).
Robert, F. et al. Global position and recruitment of HATs and HDACs in the yeast genome. Molecular Cell 16, 119–209 (2004).
Pokholok, D.K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005).
Wyrick, J. et al. Genome-wide distribution of ORC and MCM proteins in S. cerevisiae: high-resolution mapping of replication origins. Science 294, 2357–2360 (2001).
Gerton, J. et al. Inaugural article: global mapping of meiotic recombination hotspots and coldspots in the yeast Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 97, 11383–11390 (2000).
Bernstein, B.E. et al. Methylation of histone H3 Lys 4 in coding regions of active genes. Proc. Natl. Acad. Sci. USA 99, 8695–8700 (2002).
Ng, H., Robert, F., Young, R. & Struhl, K. Regulated recruitment of the ATP-dependent chromatin remodeling complex RSC in response to transcriptional repression and activation. Genes Dev. 16, 806–819 (2002).
Robyr, D. et al. Microarray deacetylation maps determine genome-wide functions for yeast histone deacetylases. Cell 109, 437–446 (2002).
Nagy, P., Cleary, M., Brown, P. & Lieb, J. Genomewide demarcation of RNA polymerase II transcription units revealed by physical fractionation of chromatin. Proc. Natl. Acad. Sci. USA 100, 6364–6369 (2003).
Kurdistani, S.K., Tavazoie, S. & Grunstein, M. Mapping global histone acetylation patterns to gene expression. Cell 117, 721–733 (2004).
Bernstein, B.E. et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120, 169–181 (2005).
Yuan, G. et al. Genome-scale identification of nucleosome positions in S. cerevisiae. Science 309, 626–630 (2005).
Marion, R.M. et al. Sfp1 is a stress- and nutrient-sensitive regulator of ribosomal protein gene expression. Proc. Natl. Acad. Sci. USA 101, 14315–14322 (2004).
Li, X. & Wong, W. Sampling motifs on phylogenetic trees. Proc. Natl. Acad. Sci. USA 102, 9481–9486 (2005).
Hartemink, A.J., Gifford, D.K., Jaakkola, T.S. & Young, R.A. Combining location and expression data for principled discovery of genetic regulatory network models. Proceedings of Pacific Symposium on Biocomputing, (Lihue, Hawaii, January 3–7, 2002) 7, 437–449 (2002).
Bar-Joseph, Z. et al. Computational discovery of gene modules and regulatory networks. Nat. Biotechnol. 21, 1337–1342 (2003).
Luscombe, N. et al. Genomic analysis of regulatory network dynamics reveals large topological changes. Nature 431, 308–312 (2004).
Buck, M.J., Nobel, A.B. & Lieb, J.D. Chipotle: a user-friendly tool for the analysis of chip-chip data. Genome Biol. 6, R97 (2005).
Roberts, C. et al. Signaling and circuitry of multiple MAPK pathways revealed by a matrix of global gene expression profiles. Science 287, 873–880 (2000).
Keles, S., Dudoit, S., van der Laan, M. & Cawley, S.E. Multiple testing methods for ChIP-Chip high density oligonucleotide array data. Berkeley Electronic Press (June, 2004). http://www.bepress.com/ucbbiostat/paper147
Kim, T.H. et al. A high-resolution map of active promoters in the human genome. Nature 436, 876–880 (2005).
Boyer, L.A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Bailey, T. & Elkan, C. The value of prior knowledge in discovering motifs with MEME. in Proceedings of the Third International Conference on Intelligent Systems for Molecular Biology, 21–29 (AAAI Press, Menlo Park, CA, 1995).
Wingender, E. et al. The TRANSFAC system on gene expression regulation. Nucleic Acids Res. 29, 281–283 (2001).
Lutfiyya, L. & Johnston, M. Two zinc-finger-containing repressors are responsible for glucose repression of SUC2 expression. Mol. Cell. Biol. 16, 4790–4797 (1996).
Neal, R.M. Probabilistic inference using Markov Chain Monte Carlo methods. Tech. Rep. CRG-TR-93–1, Dept. of Computer Science, University of Toronto (1993).
Brooks, S.P. Markov Chain Monte Carlo method and its application. Statistician 47, 69–100 (1998).
Minka, T.P. Expectation propagation for approximate Bayesian inference. in Proceedings of Uncertainty in Artificial Intelligence 362–369 (2001). http://research.microsoft.com/~minka/papers/ep/minka-ep-uai.pdf
Qi, Y. Extending expectation propagation for graphical models. Ph.D. thesis, MIT (2004). http://www.csail.mit.edu/~alanqi/papers/Qi-PhD-thesis-MIT-04.pdf
Gordon, D. B., Nekludova, L., McCallum, S. & Fraenkel, E. Tamo: a flexible, object-oriented framework for analyzing transcriptional regulation using DNA-sequence motifs. Bioinformatics 21, 3164–3165 (2005).
Acknowledgements
We would like to thank Duncan Odom and Laurie Boyer for providing DNA fragment length data for human transcription factors. This work was funded by the National Institutes of Health under grant number GM-069676.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
David Gifford consulted for Agilent Corporation during the course of this research.
Supplementary information
Supplementary Fig. 1
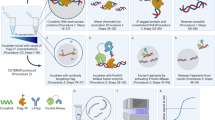
CHIP-chip experimental protocol. (PDF 54 kb)
Supplementary Fig. 2
Enrichment ratios for three replicates of an untagged control experiment and two replicates of Mig2. (PDF 109 kb)
Supplementary Fig. 3
Motif discovery output. (PDF 117 kb)
Supplementary Fig. 4
Influence function for yeast, human embryonic stem cells and human liver. (PDF 48 kb)
Supplementary Table 1
MIPS categories of Gen4 targets identified by JBD. (PDF 20 kb)
Supplementary Table 2
Known Gen4 and Mig2 log-odds weight matrix as discovered by motif discovery with positional priors from JBD and as previously published by Haribson et al. and Luftiya et al. (PDF 25 kb)
Supplementary Table 3
Performance (median distance in bp) of JBD on synthetic data. (PDF 35 kb)
Supplementary Table 4
Standard error of the mean spatial resolution of JBD on synthetic data. (PDF 34 kb)
Supplementary Table 5
Mean distance in nucleotides between binding calls and Gen4 motif in 573 promoter regions with a conserved Gen4 motif. (PDF 20 kb)
Rights and permissions
About this article
Cite this article
Qi, Y., Rolfe, A., MacIsaac, K. et al. High-resolution computational models of genome binding events. Nat Biotechnol 24, 963–970 (2006). https://doi.org/10.1038/nbt1233
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1233
This article is cited by
-
“Non-canonical protein-DNA interactions identified by ChIP are not artifacts”: response
BMC Genomics (2013)
-
ChIP-chip versus ChIP-seq: Lessons for experimental design and data analysis
BMC Genomics (2011)
-
Pinpointing transcription factor binding sites from ChIP-seq data with SeqSite
BMC Systems Biology (2011)
-
Deciphering transcription factor binding patterns from genome-wide high density ChIP-chip tiling array data
BMC Proceedings (2011)
-
The effect of prior assumptions over the weights in BayesPI with application to study protein-DNA interactions from ChIP-based high-throughput data
BMC Bioinformatics (2010)