Abstract
Enhanced biological phosphorus removal (EBPR) is one of the best-studied microbially mediated industrial processes because of its ecological and economic relevance. Despite this, it is not well understood at the metabolic level. Here we present a metagenomic analysis of two lab-scale EBPR sludges dominated by the uncultured bacterium, “Candidatus Accumulibacter phosphatis.” The analysis sheds light on several controversies in EBPR metabolic models and provides hypotheses explaining the dominance of A. phosphatis in this habitat, its lifestyle outside EBPR and probable cultivation requirements. Comparison of the same species from different EBPR sludges highlights recent evolutionary dynamics in the A. phosphatis genome that could be linked to mechanisms for environmental adaptation. In spite of an apparent lack of phylogenetic overlap in the flanking communities of the two sludges studied, common functional themes were found, at least one of them complementary to the inferred metabolism of the dominant organism. The present study provides a much needed blueprint for a systems-level understanding of EBPR and illustrates that metagenomics enables detailed, often novel, insights into even well-studied biological systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Harper, D. Eutrophication of Freshwaters (Chapman and Hall, London, 1991).
Farmer, A.M. (ed.) “Implementation of the 1991 EU Urban Waste Water Directive and its Role in Reducing Phosphate Discharges” (Summary of Report). Scope Newsletter No. 34 (1999).
US EPA/OW Clean Water Needs Survey (CWNS) for the United States and US Territories (US EPA/Office of Water, Washington, DC, 1996).
CEEP. in Second International Conference on the recovery of phosphorus from sewage and animal wastes. Noordwijkerhout, Netherlands, (2001).
Tchobanoglous, G. & Burton, F.L. . Wastewater Engineering: Treatment, Disposal, and Reuse. (McGraw-Hill, New York, 1991).
Blackall, L.L., Crocetti, G.R., Saunders, A.M. & Bond, P.L. A review and update of the microbiology of enhanced biological phosphorus removal in wastewater treatment plants. Antonie Van Leeuwenhoek 81, 681–691 (2002).
He, S., Gu, A.Z. & McMahon, K.D. Fine-scale differences between Accumulibacter-like bacteria in enhanced biological phosphorus activated sludge. Water Sci. Technol. 54, 111–117 (2006).
Fuhs, G.W. & Chen, M. Microbiological basis of phosphate removal in the activated sludge process for the treatment of wastewater. Microb. Ecol. 2, 119–138 (1975).
Streichan, M., Golecki, J.R. & Schon, G. Polyphosphate-accumulating bacteria from sewage treatment plants with different processes for biological phosphorus removal. FEMS Microbiol. Ecol. 73, 113–124 (1990).
Deinema, M.H., van Loosdrecht, M.C.M. & Scholten, A. Some physiological characteristics of Acinetobacter spp. accumulating large amounts of phosphate. Water Sci. Technol. 17, 119–125 (1985).
Wagner, M. et al. Development of an rRNA-targeted oligonucleotide probe specific for the genus Acinetobacter and its application for in situ monitoring in activated sludge. Appl. Environ. Microbiol. 60, 792–800 (1994).
Hesselmann, R.P., Werlen, C., Hahn, D., van der Meer, J.R. & Zehnder, A.J. Enrichment, phylogenetic analysis and detection of a bacterium that performs enhanced biological phosphate removal in activated sludge. Syst. Appl. Microbiol. 22, 454–465 (1999).
Crocetti, G.R. et al. Identification of Polyphosphate-Accumulating Organisms and Design of 16S rRNA-Directed Probes for Their Detection and Quantitation. Appl. Environ. Microbiol. 66, 1175–1182 (2000).
McMahon, K.D., Dojka, M.A., Pace, N.R., Jenkins, D. & Keasling, J.D. Polyphosphate kinase from activated sludge performing enhanced biological phosphorus removal. Appl. Environ. Microbiol. 68, 4971–4978 (2002).
Tyson, G.W. et al. Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428, 37–43 (2004).
Venter, J.C. et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004).
Tringe, S.G. et al. Comparative metagenomics of microbial communities. Science 308, 554–557 (2005).
Oehmen, A., Saunders, A.M., Vives, M.T., Yuan, Z. & Keller, J. Competition between polyphosphate and glycogen accumulating organisms in enhanced biological phosphorus removal systems with acetate and propionate as carbon sources. J. Biotechnol. 123, 22–32 (2005).
Burns, D.J. & Beever, R.E. Mechanisms controlling the two phosphate uptake systems in Neurospora crassa. J. Bacteriol. 139, 195–204 (1979).
Kortstee, G.J., Appeldoorn, K.J., Bonting, C.F., van Niel, E.W. & van Veen, H.W. Recent developments in the biochemistry and ecology of enhanced biological phosphorus removal. Biochemistry (Mosc.) 65, 332–340 (2000).
Seviour, R.J., Mino, T. & Onuki, M. The microbiology of biological phosphorus removal in activated sludge systems. FEMS Microbiol. Rev. 27, 99–127 (2003).
Schuler, A.J. & Jenkins, D. Enhanced biological phosphorus removal from wastewater by biomass with different phosphorus contents, Part III: Anaerobic sources of reducing equivalents. Water Environ. Res. 75, 512–522 (2003).
Pereira, H. et al. Model for carbon metabolism in biological phosphorus removal processes based on in vivo 13C-NMR labelling experiments. Water Res. 30, 2128–2138 (1996).
Louie, T.M., Mah, T.J., Oldham, W. & Ramey, W.D. Use of Metabolic Inhibitors and Gas Chromatography/Mass Spectrometry to Study Poly-β-Hydroxyalkanoates metabolism involving Cryptic Nutrients in Enhanced Biological Phosphorus Removal Systems. Water Res. 34, 1507–1514 (2000).
Lemos, P.C., Serafim, L.S., Santos, M.M., Reis, M.A. & Santos, H. Metabolic pathway for propionate utilization by phosphorus-accumulating organisms in activated sludge: 13C labeling and in vivo nuclear magnetic resonance. Appl. Environ. Microbiol. 69, 241–251 (2003).
Mino, T., Van Loosdrecht, M.C.M. & Heijnen, J.J. Microbiology and biochemistry of the enhanced biological phosphate removal process. Water Res. 32, 3193–3207 (1998).
Elbehti, A., Brasseur, G. & Lemesle-Meunier, D. First evidence for existence of an uphill electron transfer through the bc(1) and NADH-Q oxidoreductase complexes of the acidophilic obligate chemolithotrophic ferrous ion-oxidizing bacterium Thiobacillus ferrooxidans. J. Bacteriol. 182, 3602–3606 (2000).
Schuler, A.J. & Jenkins, D. Enhanced biological phosphorus removal from wastewater by biomass with different phosphorus contents, Part I: Experimental results and comparison with metabolic models. Water Environ. Res. 75, 485–498 (2003).
Maurer, M., Gujer, W., Hany, R. & Bachmann, S. Intracellular carbon flow in phosphorus accumulating organisms from activated sludge systems. Water Res. 31, 907–917 (1997).
Hesselmann, R.P.X., Von Rummell, R., Resnick, S.M., Hany, R. & Zehnder, A.J.B. Anaerobic metabolism of bacteria performing enhanced biological phosphate removal. Water Res. 34, 3487–3494 (2000).
Wilen, B.M., Jin, B. & Lant, P. Relationship between flocculation of activated sludge and composition of extracellular polymeric substances. Water Sci. Technol. 47, 95–103 (2003).
Zeng, R.J., Lemaire, R., Yuan, Z. & Keller, J. Simultaneous nitrification, denitrification, and phosphorus removal in a lab-scale sequencing batch reactor. Biotechnol. Bioeng. 84, 170–178 (2003).
White, D. The Physiology and Biochemistry of Prokaryotes. (Oxford University Press, New York, 1995).
Tyson, G.W. et al. Genome-directed isolation of the key nitrogen fixer Leptospirillum ferrodiazotrophum sp. nov. from an acidophilic microbial community. Appl. Environ. Microbiol. 71, 6319–6324 (2005).
Aparicio, S. et al. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297, 1301–1310 (2002).
Korotkova, N., Lidstrom, M.E. & Chistoserdova, L. Identification of genes involved in the glyoxylate regeneration cycle in Methylobacterium extorquens AM1, including two new genes, meaC and meaD. J. Bacteriol. 187, 1523–1526 (2005).
Acknowledgements
We thank Edward Berry for expert advice on cytochromes and Greg Crocetti, Daniel Noguera, Suzan Yilmaz, Paul Wilmes, Phil Bond, Jay Keasling and Eddy Rubin for helpful discussions. We also thank Aaron Saunders, Huabing Liu, Daniel Gall, Eugene Goltsman, Inna Dubchak, Matt Nolan, Steve Lowry, Alla Lapidus, Bryce Shepherd, Thomas Huber, Khrisna Palaniappan, Frank Korzeniewski and Sam Pitluck for technical assistance and additional analyses, and Chris Detter, Paul Richardson, Tijana Glavina del Rio, Susan Lucas, Alex Copeland, Dan Rokhsar, Igor Grigoriev and Victor Markowitz for facilitating the study. The sequencing for the project was provided by the DOE Community Sequencing Program at JGI (http://www.jgi.doe.gov/CSP/index.html). The National Science Foundation (BES 0332136) supported the efforts of K.D.M. and S.H. This work was performed under the auspices of the DOE's Office of Science, Biological and Environmental Research Program; the University of California, Lawrence Livermore National Laboratory, under contract no. W-7405-Eng-48; Lawrence Berkeley National Laboratory under contract no. DE-AC03-76SF00098; and Los Alamos National Laboratory under contract no. W-7405-ENG-36.
Author information
Authors and Affiliations
Contributions
H.G.M. contributed to metabolic reconstruction, binning and size estimation of A. phosphatis, comparison of the US and OZ genomes, gene-centric analysis and manuscript writing. N.I. contributed to the metabolic reconstruction of A. phosphatis, analysis of metabolism of the flanking community, gene-centric analysis and writing. V.K. contributed to the gene-centric analysis and A. phosphatis strain discrimination. F.W. contributed to community structure analysis. K.W.B. performed the Phrap assemblies and contributed to A. phosphatis binning. A.C.M. and I.R. contributed to the binning of the EBPR community. C.Y. and S.H. contributed to operation of and extraction of genomic DNA from the OZ and US EBPR reactors respectively. A.A.S. annotated the metagenomic assemblies. E.S. loaded the annotated assemblies into IMG/M and implemented tools specific for visualization of the data sets. E.D. constructed the shotgun libraries. N.H.P. and H.J.S. performed the JAZZ assemblies. J.L.P. contributed to the A. phosphatis genome size estimation. N.C.K. contributed to the genome completeness estimation. L.L.B., K.D.M. and P.H. contributed to the planning and design of the project, data analysis and manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Overview of US and OZ draft composite genomes. (DOC 202 kb)
Supplementary Fig. 2
Histogram of the number of reads with a given coverage. (DOC 252 kb)
Supplementary Fig. 3
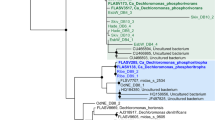
Expanded maximum likelihood 16SrRNA tree. (DOC 201 kb)
Supplementary Fig. 4
EBPR-relevant metabolism with gene OIDs included. (DOC 412 kb)
Supplementary Table 1
SBR operational differences between the US & OZ SBS sludges. (DOC 41 kb)
Supplementary Table 2
Estimate of completeness of the dominant US A. phosphatis genome. (DOC 209 kb)
Rights and permissions
About this article
Cite this article
Martín, H., Ivanova, N., Kunin, V. et al. Metagenomic analysis of two enhanced biological phosphorus removal (EBPR) sludge communities. Nat Biotechnol 24, 1263–1269 (2006). https://doi.org/10.1038/nbt1247
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1247
This article is cited by
-
Variation in the quantity and composition of phosphorus accumulating organisms in activated sludge driven by nitrate-nitrogen
Korean Journal of Chemical Engineering (2023)
-
Diversity and functional annotation of microorganisms in anaerobic chamber treating nitrate-rich wastewater
World Journal of Microbiology and Biotechnology (2023)
-
TbasCO: trait-based comparative ‘omics identifies ecosystem-level and niche-differentiating adaptations of an engineered microbiome
ISME Communications (2022)
-
Diluted Luria-Bertani medium vs. sewage sludge as growth media: comparison of community structure and diversity in the culturable bacteria
Applied Microbiology and Biotechnology (2021)
-
Diversity and functional annotation of microorganisms in French vertical flow constructed wetland treating greywater
World Journal of Microbiology and Biotechnology (2020)