Abstract
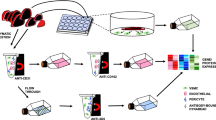
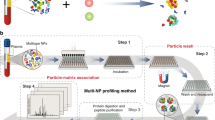
Endothelial cells can function differently in vitro and in vivo; however, the degree of microenvironmental modulation in vivo remains unknown at the molecular level largely because of analytical limitations. We use multidimensional protein identification technology (MudPIT) to identify 450 proteins (with three or more spectra) in luminal endothelial cell plasma membranes isolated from rat lungs and from cultured rat lung microvascular endothelial cells. Forty-one percent of proteins expressed in vivo are not detected in vitro. Statistical analysis measuring reproducibility reveals that seven to ten MudPIT measurements are necessary to achieve ≥95% confidence of analytical completeness with current ion trap equipment. Large-scale mapping of the proteome of vascular endothelial cell surface in vivo, as demonstrated here, is advisable because distinct protein expression is apparently regulated by the tissue microenvironment that cannot yet be duplicated in standard cell culture.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Schnitzer, J.E. Update on the cellular and molecular basis of capillary permeability. Trends Cardiovasc. Med. 3, 124–130 (1993).
Michiels, C. Endothelial cell functions. J. Cell. Physiol. 196, 430–443 (2003).
Madri, J.A. & Williams, S.K. Capillary endothelial cell culture: Phenotype modulation by matrix components. J. Cell Biol. 97, 153–165 (1983).
Goerdt, S. et al. Characterization and differential expression of an endothelial cell- specific surface antigen in continuous and sinusoidal endothelial, in skin vascular lesions and in vitro. Exp. Cell Biol. 57, 185–192 (1989).
Gumkowski, F., Kaminska, G., Kaminski, M., Morrissey, L.W. & Auerbach, R. Heterogeneity of mouse vascular endothelium. Blood Vessels 24, 11–23 (1987).
Aird, W.C. et al. Vascular bed-specific expression of an endothelial cell gene is programmed by the tissue microenvironment. J. Cell Biol. 138, 1117–1124 (1997).
Janzer, R.C. & Raff, M.C. Astrocytes induce blood-brain barrier properties in endothelial cells. Nature 325, 253–257 (1987).
Stewart, P.A. & Wiley, M.J. Developing nervous tissue induces formation of blood-brain barrier characteristics in invading endothelial cells: a study using quail-chick transplantation chimeras. Dev. Biol. 84, 183–192 (1981).
Auerbach, R., Alby, L., Morrissey, L.W., Tu, M. & Joseph, J. Expression of organ-specific antigens on capillary endothelial cells. Microvasc. Res. 29, 401–411 (1985).
St. Croix, B. et al. Genes expressed in human tumor endothelium. Science 289, 1197–1202 (2000).
Obermeyer, N., Janson, N., Bergmann, J., Buck, F. & Ito, W.D. Proteome analysis of migrating versus nonmigrating rat heart endothelial cells reveals distinct expression patterns. Endothelium 10, 167–178 (2003).
Bruneel, A. et al. Proteomic study of human umbilical vein endothelial cells in culture. Proteomics 3, 714–723 (2003).
Pasqualini, R. & Ruoslahti, E. Organ targeting in vivo using phage display peptide libraries. Nature 380, 364–366 (1996).
Rajotte, D. et al. Molecular heterogeneity of the vascular endothelium revealed by in vivo phage display. J. Clin. Invest. 102, 430–437 (1998).
Schnitzer, J.E., McIntosh, D.P., Dvorak, A.M., Liu, J. & Oh, P. Separation of caveolae from associated microdomains of GPI-anchored proteins. Science 269, 1435–1439 (1995).
Oh, P. & Schnitzer, J.E. Isolation and subfractionation of plasma membranes to purify caveolae seperately from glycosyl-phosphatidylinositol-anchored protein microdomain in Cell Biology: A Laboratory Handbook, vol. 2 (ed. Celis, J..) 34–45 (Academic Press, Orlando, 1998).
Rizzo, V., Morton, C., DePaola, N., Schnitzer, J.E. & Davies, P.F. Recruitment of endothelial caveolae into mechanotransduction pathways by flow conditioning in vitro. Am. J. Physiol. Heart Circ. Physiol. 285, H1720–H1729 (2003).
Schnitzer, J.E., Liu, J. & Oh, P. Endothelial caveolae have the molecular transport machinery for vesicle budding, docking, and fusion including VAMP, NSF, SNAP, annexins, and GTPases. J. Biol. Chem. 270, 14399–14404 (1995).
Schnitzer, J.E. & Oh, P. Aquaporin-1 in plasma membrane and caveolae provides mercury-sensitive water channels across lung endothelium. Am. J. Physiol. 270, H416–H422 (1996).
Schnitzer, J.E. & Oh, P. Antibodies to SPARC inhibit albumin binding to SPARC, gp60, and microvascular endothelium. Am. J. Physiol. 263, H1872–H1879 (1992).
Wolters, D.A., Washburn, M.P. & Yates, J.R., III. An automated multidimensional protein identification technology for shotgun proteomics. Anal. Chem. 73, 5683–5690 (2001).
Jeffries, W.A. et al. Transferrin receptor on endothelium of brain capillaries. Nature 312, 162–163 (1984).
Schnitzer, J.E. & Oh, P. Albondin-mediated capillary permeability to albumin. Differential role of receptors in endothelial transcytosis and endocytosis of native and modified albumins. J. Biol. Chem. 269, 6072–6082 (1994).
Schnitzer, J.E. gp60 is an albumin-binding glycoprotein expressed by continuous endothelium involved in albumin transcytosis. Am. J. Physiol. 262, H246–H254 (1992).
Christian, S. et al. Nucleolin expressed at the cell surface is a marker of endothelial cells in angiogenic blood vessels. J. Cell Biol. 163, 871–878 (2003).
Negrutskii, B.S. & El'skaya, A.V. Eukaryotic translation elongation factor 1 alpha: structure, expression, functions, and possible role in aminoacyl-tRNA channeling. Prog. Nucleic Acid Res. Mol. Biol. 60, 47–78 (1998).
Honscha, W., Ottallah, M., Kistner, A., Platte, H. & Petzinger, E. A membrane-bound form of protein disulfide isomerase (PDI) and the hepatic uptake of organic anions. Biochim. Biophys. Acta 1153, 175–183 (1993).
Hebert, C. et al. Cell surface colligin/Hsp47 associates with tetraspanin protein CD9 in epidermoid carcinoma cell lines. J. Cell. Biochem. 73, 248–258 (1999).
Schnitzer, J. The endothelial cell surface and caveolae in health and disease. in Vascular Endothelium: Physiology, Pathology and Therapeutic Opportunities (eds. Born, G.V.R. & Schwartz, C.J.) 77–95 (1997).
Washburn, M.P., Wolters, D. & Yates, J.R. III. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
McIntosh, D.P., Tan, X.Y., Oh, P. & Schnitzer, J.E. Targeting endothelium and its dynamic caveolae for tissue-specific transcytosis in vivo: a pathway to overcome cell barriers to drug and gene delivery. Proc. Natl. Acad. Sci. USA 99, 1996–2001 (2002).
Carver, L.A. & Schnitzer, J.E. Caveolae: mining little caves for new cancer targets. Nat. Rev. Cancer 3, 571–581 (2003).
Weinman, E.J., Steplock, D. & Shenolikar, S. Acute regulation of NHE3 by protein kinase A requires a multiprotein signal complex. Kidney Int. 60, 450–454 (2001).
Abe, J., Suzuki, H., Notoya, M., Yamamoto, T. & Hirose, S. Ig-hepta, a novel member of the G protein-coupled hepta-helical receptor (GPCR) family that has immunoglobulin-like repeats in a long N-terminal extracellular domain and defines a new subfamily of GPCRs. J. Biol. Chem. 274, 19957–19964 (1999).
van der Merwe, P.A. et al. The NH2-terminal domain of rat CD2 binds rat CD48 with a low affinity and binding does not require glycosylation of CD2. Eur. J. Immunol. 23, 1373–1377 (1993).
Magee, J.C., Stone, A.E., Oldham, K.T. & Guice, K.S. Isolation, culture, and characterization of rat lung microvascular endothelial cells. Am. J. Physiol. 267, L433–L441 (1994).
Link, A.J. et al. Direct analysis of protein complexes using mass spectrometry. Nat. Biotechnol. 17, 676–682 (1999).
Eng, J. & McCormac, A. & Yates, J.R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Tabb, D.L., McDonald, W.H. & Yates, J.R. III. DTASelect and Contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J. Proteome Res. 1, 21–26 (2002).
Acknowledgements
We thank Michelle Bourne, Lisa Randall and Traci Smith for technical assistance; David Tabb for help setting up DTASelect; and Yan Li for helpful discussion. This research was supported by grants to J.E.S from the National Institutes of Health (Heart, Lung and Blood nos. R01 HL52766, R01 HL58216), National Cancer Institute (no. R01 CA83989, R24 CA095893, R33 CA97528), Sidney Kimmel, Schutz Foundation, California Tobacco-related Disease Research Program (no. 11RT-0167) and California Breast Cancer Research Program (no. 8WB-00114).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Integral and lipid-anchored plasma membrane proteins (PDF 203 kb)
Supplementary Table 2
Endothelial cell associated marker proteins (PDF 173 kb)
Supplementary Notes
Additional findings and validation (PDF 338 kb)
Rights and permissions
About this article
Cite this article
Durr, E., Yu, J., Krasinska, K. et al. Direct proteomic mapping of the lung microvascular endothelial cell surface in vivo and in cell culture. Nat Biotechnol 22, 985–992 (2004). https://doi.org/10.1038/nbt993
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt993
This article is cited by
-
Mitigating the non-specific uptake of immunomagnetic microparticles enables the extraction of endothelium from human fat
Communications Biology (2021)
-
Establishment of a striped catfish skin explant model for studying the skin response in Aeromonas hydrophila infections
Scientific Reports (2021)
-
Tumor endothelial cell-derived cadherin-2 promotes angiogenesis and has prognostic significance for lung adenocarcinoma
Molecular Cancer (2019)
-
Proteomic atlas of organ vasculopathies triggered by Staphylococcus aureus sepsis
Nature Communications (2019)
-
PRL3-zumab as an immunotherapy to inhibit tumors expressing PRL3 oncoprotein
Nature Communications (2019)