Abstract
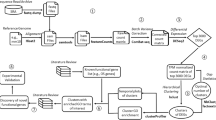
We report here the discovery of 137 previously unappreciated genes in yeast through a widely applicable and highly scalable approach integrating methods of gene-trapping, microarray-based expression analysis, and genome-wide homology searching. Our approach is a multistep process in which expressed sequences are first trapped using a modified transposon that produces protein fusions to β-galactosidase (β-gal); non-annotated open reading frames (ORFs) translated as β-gal chimeras are selected as a candidate pool of potential genes. To verify expression of these sequences, labeled RNA is hybridized against a microarray of oligonucleotides designed to detect gene transcripts in a strand-specific manner. In complement to this experimental method, novel genes are also identified in silico by homology to previously annotated proteins. As these methods are capable of identifying both short ORFs and antisense ORFs, our approach provides an effective supplement to current gene-finding schemes. In total, the genes discovered using this approach constitute 2% of the yeast genome and represent a wealth of overlooked biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Stein, L. Genome annotation: from sequence to biology. Nat. Rev. Genet. 2, 493–503 (2001).
Gopal, S. et al. Homology-based annotation yields 1,042 new candidate genes in the Drosophila melanogaster genome. Nat. Genet. 27, 337–340 (2001).
Adams, M.D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Reboul, J. et al. Open-reading frame sequence tags (OSTs) support the existence of at least 17,300 genes in C. elegans. Nat. Genet. 27, 332–336 (2001).
Shoemaker, D.D. et al. Experimental annotation of the human genome using microarray technology. Nature 409, 922–927 (2001).
Goffeau, A. et al. Life with 6000 genes. Science 274, 546, 563–567 (1996).
Mewes, H.W. et al. Overview of the yeast genome. Nature 387 (Suppl.), 7–8 (1997).
Philippsen, P. et al. The nucleotide sequence of Saccharomyces cerevisiae chromosome XIV and its evolutionary implications. Nature 387 (Suppl.), 93–98 (1997).
Velculescu, V.E. et al. Characterization of the yeast transcriptome. Cell 88, 243–251 (1997).
Blandin, G. et al. Genomic exploration of the hemiascomycetous yeasts: 4. the genome of Saccharomyces cerevisiae revisited. FEBS Lett. 487, 31–36 (2000).
Cliften, P.F. et al. Surveying Saccharomyces genomes to identify functional elements by comparative DNA sequence analysis. Genome Res. 11, 1175–1186 (2001).
Ross-Macdonald, P., Sheehan, A., Roeder, G.S. & Snyder, M. A multipurpose transposon system for analyzing protein production, localization, and function in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 94, 190–195 (1997).
Seifert, H.S., Chen, E.Y., So, M. & Heffron, F. Shuttle mutagenesis: a method of transposon mutagenesis for Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 83, 735–739 (1986).
Ross-MacDonald, P. et al. Large-scale analysis of the yeast genome by transposon tagging and gene disruption. Nature 402, 413–418 (1999).
Kumar, A., des Etages, S.A., Coelho, P.S.R., Roeder, G.S. & Snyder, M. High-throughput methods for the large-scale analysis of gene function by transposon tagging. Methods Enzymol. 328, 550–574 (2000).
Selinger, D.W. et al. RNA expression analysis using a 30 base pair resolution Escherichia coli genome array. Nat. Biotechnol. 18, 1262–1268 (2000).
Kane, M.D. et al. Assessment of the sensitivity and specificity of oligonucleotide (50mer) microarrays. Nucleic Acids Res. 28, 4552–4557 (2000).
Zhu, J. & Zhang, M.Q. SCPD: a promoter database of the yeast Saccharomyces cerevisiae. Bioinformatics 15, 607–611 (1999).
Sharp, P.M. & Li, W.H. The codon adaptation index—a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 15, 1281–1295 (1987).
Bairoch, A. & Apweiler, R. The SWISS–PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 28, 45–48 (2000).
Wood, V., Rutherford, K.M., Ivens, A., Rajandream, M.-A. & Barrell, B. A re-annotation of the Saccharomyces cerevisiae genome. Comp. Funct. Genom. 2, 143–154 (2001).
Ball, C. et al. Saccharomyces Genome Database provides tools to survey gene expression and functional analysis data. Nucleic Acids Res. 29, 80–81 (2001).
Basrai, M.A., Hieter, P. & Boeke, J.D. Small open reading frames: beautiful needles in the haystack. Genome Res. 7, 768–771 (1997).
Basrai, M.A., Velculescu, V.E., Kinzler, K.W. & Hieter, P. NORF5/HUG1 is a component of the MEC1-mediated checkpoint response to DNA damage and replication arrest in Saccharomyces cerevisiae. Mol. Cell. Biol. 19, 7041–7049 (1999).
Wagner, E.G. & Simons, R.W. Antisense RNA control in bacteria, phages, and plasmids. Annu. Rev. Microbiol. 48, 713–742 (1994).
Henikoff, S., Keene, M.A., Fechtel, K. & Fristrom, J.W. Gene within a gene: nested Drosophila genes encode unrelated proteins on opposite DNA strands. Cell 44, 33–42 (1986).
Spencer, C.A., Gietz, R.D. & Hodgetts, R.B. Overlapping transcription units in the dopa decarboxylase region of Drosophila. Nature 322, 279–281 (1986).
Vanhee-Brossollet, C. & Vaquero, C. Do natural antisense transcripts make sense in eukaryotes? Gene 211, 1–9 (1998).
Mackiewicz, P., Kowalczuk, M., Gierlik, A., Dudek, M.R. & Cebrat, S. Origin and properties of non-coding ORFs in the yeast genome. Nucleic Acids Res. 27, 3503–3509 (1999).
Zhang, C.-T. & Wang, J. Recognition of protein coding genes in the yeast genome at better than 95% accuracy based on the Z curve. Nucleic Acids Res. 28, 2804–2814 (2000).
Malpertuy, A. et al. Genomic exploration of the hemiascomycetous yeasts: 19. Ascomycetes-specific genes. FEBS Lett. 487, 113–121 (2000).
Ito, H., Fukuda, Y., Murata, K. & Kimura, A. Transformation of intact yeast cells treated with alkali cations. J. Bacteriol. 153, 163–168 (1983).
Adams, A., Gottschling, D.E., Kaiser, C.A. & Stearns, T. Methods in yeast genetics, 1997 Edn. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY; 1998).
Altschul, S.F., Gish, W., Miller, W., Meyers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Holstege, F.C.P. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998).
Pearson, W.R., Wood, T., Zhang, Z. & Miller, W. Comparison of DNA sequences with protein sequences. Genomics 46, 24–36 (1997).
Wootton, J.C. Non-globular domains in protein sequences: automated segmentation using complexity measures. Comput. Chem. 18, 269–285 (1994).
Acknowledgements
We thank Lara Umansky, Stacy Piccirillo, and Sandra Matson for technical assistance, and Metin Bilgin for helpful suggestions. This work was supported by NIH Grant R01-CA77808 to M.S. A.K. is supported by a post-doctoral fellowship from the American Cancer Society.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Kumar, A., Harrison, P., Cheung, KH. et al. An integrated approach for finding overlooked genes in yeast. Nat Biotechnol 20, 58–63 (2002). https://doi.org/10.1038/nbt0102-58
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0102-58
This article is cited by
-
Small open reading frames in plant research: from prediction to functional characterization
3 Biotech (2022)
-
Evaluation of the phytoremediation uptake model for predicting heavy metals (Pb, Cd, and Zn) from the soil using Nerium oleander L.
Environmental Science and Pollution Research (2020)
-
Microbial diversity in various types of paper mill sludge: identification of enzyme activities with potential industrial applications
SpringerPlus (2016)
-
Expression of the rDNA-encoded mitochondrial protein Tar1p is stringently controlled and responds differentially to mitochondrial respiratory demand and dysfunction
Current Genetics (2008)