Abstract
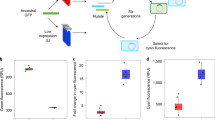
Green fluorescent protein (QFP) has rapidly become a widely used reporter of gene regulation. However, for many organisms, particularly eukaryotes, a stronger whole cell fluorescence signal is desirable. We constructed a synthetic GFP gene with improved codon usage and performed recursive cycles of DNA shuffling followed by screening for the brightest E. coli colonies. A visual screen using UV light, rather than FACS selection, was used to avoid red-shifting the excitation maximum. After 3 cycles of DNA shuffling, a mutant was obtained with a whole cell fluorescence signal that was 45-fold greater than a standard, the commercially available Clontech plasmid pGFP. The expression level in E. coli was unaltered at about 75% of total protein. The emission and excitation maxima were also unchanged. Whereas in E. coli most of the wildtype GFP ends up in inclusion bodies, unable to activate its chromophore, most of the mutant protein is soluble and active. Three amino acid mutations appear to guide the mutant protein into the native folding pathway rather than toward aggregation. Expressed in Chinese Hamster Ovary (CHO) cells, this shuffled GFP mutant showed a 42-fold improvement over wildtype GFP sequence, and is easily detected with UV light in a wide range of assays. The results demonstrate how molecular evolution can solve a complex practical problem without needing to first identify which process is limiting. DNA shuffling can be combined with screening of a moderate number of mutants. We envision that the combination of DNA shuffling and high throughput screening will be a powerful tool for the optimization of many commercially important enzymes for which selections do not exist.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Prasher, D.C., Eckenrode, V.K., Ward, W.W., Prendergast, F.G. and Cormier, M.J. 1992. Gene 111: 229–233.
Prasher, D.C. 1995. Using GFP to see the light. Trends in Genetics 11: 320–323.
Chalfie, M. Tu, Y., Euskirchen, G., Ward, W.W. and Prasher, D.C. 1994. Green fluorescent protein as a marker for gene expression. Science 263: 802–805.
Inouye, S. and Tsuji, F.I. 1994. FEBS Lett. 341: 277–280.
Wang, S. and Hazelrigg, T. 1994. Implications for bed mRNA localization from spatial distribution of exu protein in Drosophila oogenesis. Nature 369: 400–403.
Brand, A. 1995. GFP in Drosophila. Trends in Genetics 11: 324–325.
Pines, J. 1995. GFP in mammalian cells. Trends in Genetics 11: 326–327.
Hodgkinson, S. 1995. GFP in Dictyostelium. Trends in Genetics 11: 327–328.
Haseloff, J. and Amos, B. 1995. GFP in plants. Trends in Genetics 11: 328–329.
Heim, R., Prasher, D.C. and Tsien, R.Y. 1994. Wavelength mutations and post-translational autoxidation of green fluorescent protein. Proc. Natl. Acad. Sci. USA 91: 12501–12504.
Heim, R., Cubitt, A.B. and Tsien, R.Y. 1995. Improved green fluorescence. Nature 373: 663–664.
Cormack, B.P., Valdivia, R.H. and Falkow, S. 1996. FACS-optimized mutants of the green fluorescent protein (GFP). Gene. In press.
Delagrave, S., Hawtin, R.E., Silva, C.M., Yang, M.M. and Youvan, D.C. 1995. Red-shifted excitation mutants of the green fluorescent protein. Bio/Technology 13: 151–154.
Ward, W.W., Cody, C.W., Hart, R.C. and Cormier, M.J. 1982. Spectrophotometric identity of the energy transfer chromophores in Renilla and Aequorea green fluorescent proteins. Photochem. Photobiol. 35: 803–808.
Stemmer, W.P.C. 1994. DNA shuffling by random fragmentation and reassembly: In vitro recombination for molecular evolution. Proc. Natl. Acad. Sci. USA 91: 10747–10751.
Stemmer, W.P.C. 1994. Rapid evolution of a protein in vitro by DNA shuffling. Nature 370: 389–391.
Stemmer, W.P.C. 1995. Searching sequence space. Bio/Technology 13: 549–553.
Kyte, J. and Doolittle, R.F. 1982. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157: 105–132.
Whitehorn, E., Tate, E., Yanofsky, S.D., Kochersperger, L., Davis, A., Mortensen, R.B., Yonkovich, S., Bell, K., Dower, W. J. and Barrett, R.W. 1995. A generic method for expression and use of 'tagged' soluble versions of cell surface receptors. Bio/Technology 13: 1215–1219.
Guzman, L.-M., Belin, D., Carson, M.J. and Beckwith, J. 1995. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177: 4121–4130.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Crameri, A., Whitehorn, E., Tate, E. et al. Improved Green Fluorescent Protein by Molecular Evolution Using DNA Shuffling. Nat Biotechnol 14, 315–319 (1996). https://doi.org/10.1038/nbt0396-315
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0396-315
This article is cited by
-
Polyamine-mediated mechanisms contribute to oxidative stress tolerance in Pseudomonas syringae
Scientific Reports (2023)
-
Vibrio cholerae biofilms use modular adhesins with glycan-targeting and nonspecific surface binding domains for colonization
Nature Communications (2023)
-
Consequences of Genetic Recombination on Protein Folding Stability
Journal of Molecular Evolution (2023)
-
Low protein expression enhances phenotypic evolvability by intensifying selection on folding stability
Nature Ecology & Evolution (2022)
-
Performance assessment of a multi-epitope chimeric antigen for the serological diagnosis of acute Mayaro fever
Scientific Reports (2021)