Abstract
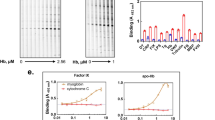
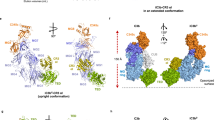
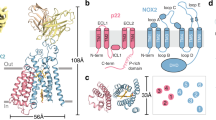
Red cell haemoglobin is the fundamental oxygen-transporting molecule in blood, but also a potentially tissue-damaging compound owing to its highly reactive haem groups. During intravascular haemolysis, such as in malaria and haemoglobinopathies1, haemoglobin is released into the plasma, where it is captured by the protective acute-phase protein haptoglobin. This leads to formation of the haptoglobin–haemoglobin complex, which represents a virtually irreversible non-covalent protein–protein interaction2. Here we present the crystal structure of the dimeric porcine haptoglobin–haemoglobin complex determined at 2.9 Å resolution. This structure reveals that haptoglobin molecules dimerize through an unexpected β-strand swap between two complement control protein (CCP) domains, defining a new fusion CCP domain structure. The haptoglobin serine protease domain forms extensive interactions with both the α- and β-subunits of haemoglobin, explaining the tight binding between haptoglobin and haemoglobin. The haemoglobin-interacting region in the αβ dimer is highly overlapping with the interface between the two αβ dimers that constitute the native haemoglobin tetramer. Several haemoglobin residues prone to oxidative modification after exposure to haem-induced reactive oxygen species are buried in the haptoglobin–haemoglobin interface, thus showing a direct protective role of haptoglobin. The haptoglobin loop previously shown to be essential for binding of haptoglobin–haemoglobin to the macrophage scavenger receptor CD163 (ref. 3) protrudes from the surface of the distal end of the complex, adjacent to the associated haemoglobin α-subunit. Small-angle X-ray scattering measurements of human haptoglobin–haemoglobin bound to the ligand-binding fragment of CD163 confirm receptor binding in this area, and show that the rigid dimeric complex can bind two receptors. Such receptor cross-linkage may facilitate scavenging and explain the increased functional affinity of multimeric haptoglobin–haemoglobin for CD163 (ref. 4).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bunn, H. F., Forget, B. G. & Ranney, H. M. Hemoglobinopathies 308 ( Saunders, W. B. Company, 1977)
Nagel, R. L. & Gibson, Q. H. The binding of hemoglobin to haptoglobin and its relation to subunit dissociation of hemoglobin. J. Biol. Chem. 246, 69–73 (1971)
Nielsen, M. J. et al. A unique loop extension in the serine protease domain of haptoglobin is essential for CD163 recognition of the haptoglobin–hemoglobin complex. J. Biol. Chem. 282, 1072–1079 (2007)
Kristiansen, M. et al. Identification of the haemoglobin scavenger receptor. Nature 409, 198–201 (2001)
Sadrzadeh, S. M., Graf, E., Panter, S. S., Hallaway, P. E. & Eaton, J. W. Hemoglobin. A biologic fenton reagent. J. Biol. Chem. 259, 14354–14356 (1984)
Shim, B. S., Lee, T. H. & Kang, Y. S. Immunological and biochemical investigations of human serum haptoglobin: composition of haptoglobin–haemoglobin intermediate, haemoglobin-binding sites and presence of additional alleles for β-chain. Nature 207, 1264–1267 (1965)
Lim, S. K. et al. Increased susceptibility in Hp knockout mice during acute hemolysis. Blood 92, 1870–1877 (1998)
Buehler, P. W. et al. Haptoglobin preserves the CD163 hemoglobin scavenger pathway by shielding hemoglobin from peroxidative modification. Blood 113, 2578–2586 (2009)
Fagoonee, S. et al. Plasma protein haptoglobin modulates renal iron loading. Am. J. Pathol. 166, 973–983 (2005)
Gaboriaud, C., Rossi, V., Bally, I., Arlaud, G. J. & Fontecilla-Camps, J. C. Crystal structure of the catalytic domain of human complement C1s: a serine protease with a handle. EMBO J. 19, 1755–1765 (2000)
Budayova-Spano, M. et al. The crystal structure of the zymogen catalytic domain of complement protease C1r reveals that a disruptive mechanical stress is required to trigger activation of the C1 complex. EMBO J. 21, 231–239 (2002)
Perutz, M. F. et al. Structure of haemoglobin: a three-dimensional Fourier synthesis at 5.5-Å resolution, obtained by X-ray analysis. Nature 185, 416–422 (1960)
Wejman, J. C., Hovsepian, D., Wall, J. S., Hainfeld, J. F. & Greer, J. Structure of haptoglobin and the haptoglobin–hemoglobin complex by electron microscopy. J. Mol. Biol. 174, 319–341 (1984)
Perona, J. J. & Craik, C. S. Evolutionary divergence of substrate specificity within the chymotrypsin-like serine protease fold. J. Biol. Chem. 272, 29987–29990 (1997)
Smulevich, G., Possenti, M., D’Avino, R., di Prisco, G. & Coletta, M. Spectroscopic studies of the heme active site of hemoglobin from Chelidonichthys kumu. J. Raman Spectrosc. 29, 57–65 (1998)
Park, S.-Y., Yokoyama, T., Shibayama, N., Shiro, Y. & Tame, J. R. H. 1.25 Å resolution crystal structures of human haemoglobin in the oxy, deoxy and carbonmonoxy forms. J. Mol. Biol. 360, 690–701 (2006)
Kardos, J. et al. Revisiting the mechanism of the autoactivation of the complement protease C1r in the C1 complex: structure of the active catalytic region of C1r. Mol. Immunol. 45, 1752–1760 (2008)
Dickerson, R. E. & Geis, I. Hemoglobin: Structure, Function, Evolution and Pathology (The Benjamin/Cummings Publishing Company, 1983)
Nagel, R. L., Whittenberg, J. B. & Ranney, H. M. Oxygen Equilibria of the hemoglobin–haptoglobin complex. Biochim. Biophys. Acta 100, 286–289 (1965)
Venkatesh, B., Miyazaki, G., Imai, K., Morimoto, H. & Hori, H. Oxygen equilibrium and EPR studies on α1β1 hemoglobin dimer. J. Biochem. 136, 595–600 (2004)
Ip, S. H., Johnson, M. L. & Ackers, G. K. Kinetics of deoxyhemoglobin subunit dissociation determined by haptoglobin binding: estimation of the equilibrium constant from forward and reverse rates. Biochemistry 15, 654–660 (1976)
Azarov, I. et al. Rate of nitric oxide scavenging by hemoglobin bound to haptoglobin. Nitric Oxide 18, 296–302 (2008)
Miller, Y. I., Altamentova, S. M. & Shaklai, N. Oxidation of low-density lipoprotein by hemoglobin stems from a heme-initiated globin radical: antioxidant role of haptoglobin. Biochemistry 36, 12189–12198 (1997)
Pimenova, T. et al. Quantitative mass spectrometry defines an oxidative hotspot in hemoglobin that is specifically protected by haptoglobin. J. Proteome Res. 9, 4061–4070 (2010)
Jia, Y., Buehler, P. W., Boykins, R. A., Venable, R. M. & Alayash, A. I. Structural basis of peroxide-mediated changes in human hemoglobin: a novel oxidative pathway. J. Biol. Chem. 282, 4894–4907 (2007)
Smithies, O. & Walker, N. F. Notation for serum-protein groups and the genes controlling their inheritance. Nature 178, 694–695 (1956)
Maeda, N., Yang, F., Barnett, D. R., Bowman, B. H. & Smithies, O. Duplication within the haptoglobin Hp2 gene. Nature 309, 131–135 (1984)
Vanhollebeke, B. et al. A haptoglobin–hemoglobin receptor conveys innate immunity to Trypanosoma brucei in humans. Science 320, 677–681 (2008)
Nielsen, M. J. et al. Haptoglobin-related protein is a high-affinity hemoglobin-binding plasma protein. Blood 108, 2846–2849 (2006)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
Laskowski, R., MacArthur, M., Moss, D. & Thornton, J. Procheck – a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993)
The. PyMOL Molecular Graphics System v. 1.3r1 (Schrödinger, LLC, 2010)
Bond, C. S. & Schuettelkopf, A. W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta Crystallogr. D 65, 510–512 (2009)
Pedersen, J. A flux- and background-optimized version of the NanoSTAR small-angle X-ray scattering camera for solution scattering. J. Appl. Cryst. 37, 369–380 (2004)
Madsen, M. et al. Molecular characterization of the haptoglobin·hemoglobin receptor CD163. Ligand binding properties of the scavenger receptor cysteine-rich domain region. J. Biol. Chem. 279, 51561–51567 (2004)
Oliveira, C. L. P. et al. A SAXS study of glucagon fibrillation. J. Mol. Biol. 387, 147–161 (2009)
Pedersen, J. S., Hansen, S. & Bauer, R. The aggregation behavior of zinc-free insulin studied by small-angle neutron scattering. Eur. Biophys. J. 22, 379–389 (1994)
Glatter, O. New method for evaluation of small-angle scattering data. J. Appl. Crystallogr. 10, 415–421 (1977)
Svergun, D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 76, 2879–2886 (1999)
Svergun, D. I., Petoukhov, M. V. & Koch, M. H. Determination of domain structure of proteins from X-ray solution scattering. Biophys. J. 80, 2946–2953 (2001)
Svergun, D., Barberato, C. & Koch, M. CRYSOL — A program to evaluate x-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995)
Petoukhov, M. V. & Svergun, D. I. Global rigid body modeling of macromolecular complexes against small-angle scattering data. Biophys. J. 89, 1237–1250 (2005)
Volkov, V. & Svergun, D. Uniqueness of ab initio shape determination in small-angle scattering. J. Appl. Crystallogr. 36, 860–864 (2003)
Acknowledgements
We are grateful to G. Ratz for technical assistance, R. Kidmose, N. S. Laursen and the staff at the Swiss Light Source beamlines for help with data collection. We thank G. V. Jensen for performing SAXS measurements and R. E. Weber for scientific discussion. The research was supported by The Lundbeck Foundation, The Novo Nordisk Foundation, The Research Council of Norway, The European Research Council and The Danish Council for Independent Research.
Author information
Authors and Affiliations
Contributions
C.B.F.A.: purification, crystallization, data collection, structure determination and analysis, manuscript preparation and study design. M.T.-J.: purification, crystallization, data collection and structure determination. M.J.N.: cloning, expression and purification. C.L.P.d.O.: SAXS measurements and analysis. H.-P.H.: ultraviolet–visible spectroscopy and analysis. N.H.A.: Raman spectroscopy and analysis. J.S.P.: SAXS measurements and analysis. G.R.A.: study design. S.K.M.: manuscript preparation and study design.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures 1-13 and additional references. (PDF 1629 kb)
Rights and permissions
About this article
Cite this article
Andersen, C., Torvund-Jensen, M., Nielsen, M. et al. Structure of the haptoglobin–haemoglobin complex. Nature 489, 456–459 (2012). https://doi.org/10.1038/nature11369
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11369
This article is cited by
-
Oxidized hemoglobin triggers polyreactivity and autoreactivity of human IgG via transfer of heme
Communications Biology (2023)
-
Die Rolle des Hämoxygenase-1-CD163-Signalweges bei atherosklerotischen Gefäßerkrankungen
Gefässchirurgie (2022)
-
A Slam-dependent hemophore contributes to heme acquisition in the bacterial pathogen Acinetobacter baumannii
Nature Communications (2021)
-
βCysteine 93 in human hemoglobin: a gateway to oxidative stability in health and disease
Laboratory Investigation (2021)
-
The role of haptoglobin and hemopexin in the prevention of delayed cerebral ischaemia after aneurysmal subarachnoid haemorrhage: a review of current literature
Neurosurgical Review (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.