Abstract
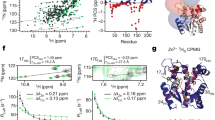
A long-standing challenge is to understand at the atomic level how protein dynamics contribute to enzyme catalysis. X-ray crystallography can provide snapshots of conformational substates sampled during enzymatic reactions1, while NMR relaxation methods reveal the rates of interconversion between substates and the corresponding relative populations1,2. However, these current methods cannot simultaneously reveal the detailed atomic structures of the rare states and rationalize the finding that intrinsic motions in the free enzyme occur on a timescale similar to the catalytic turnover rate. Here we introduce dual strategies of ambient-temperature X-ray crystallographic data collection and automated electron-density sampling to structurally unravel interconverting substates of the human proline isomerase, cyclophilin A (CYPA, also known as PPIA). A conservative mutation outside the active site was designed to stabilize features of the previously hidden minor conformation. This mutation not only inverts the equilibrium between the substates, but also causes large, parallel reductions in the conformational interconversion rates and the catalytic rate. These studies introduce crystallographic approaches to define functional minor protein conformations and, in combination with NMR analysis of the enzyme dynamics in solution, show how collective motions directly contribute to the catalytic power of an enzyme.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007)
Mittermaier, A. & Kay, L. E. Review—new tools provide new insights in NMR studies of protein dynamics. Science 312, 224–228 (2006)
Boehr, D. D., McElheny, D., Dyson, H. J. & Wright, P. E. The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313, 1638–1642 (2006)
Eisenmesser, E. Z. et al. Intrinsic dynamics of an enzyme underlies catalysis. Nature 438, 117–121 (2005)
Hammes-Schiffer, S. & Benkovic, S. J. Relating protein motion to catalysis. Annu. Rev. Biochem. 75, 519–541 (2006)
Schramm, V. L. & Shi, W. Atomic motion in enzymatic reaction coordinates. Curr. Opin. Struct. Biol. 11, 657–665 (2001)
Agarwal, P. K. Cis/trans isomerization in HIV-1 capsid protein catalyzed by cyclophilin A: insights from computational and theoretical studies. Proteins 56, 449–463 (2004)
Hamelberg, D. & McCammon, A. Mechanistic insight into the role of transition-state stabilization in cyclophilin A. J. Am. Chem. Soc. 131, 147–152 (2009)
Li, G. H. & Cui, Q. What is so special about Arg 55 in the catalysis of cyclophilin A? Insights from hybrid QM/MM simulations. J. Am. Chem. Soc. 125, 15028–15038 (2003)
Trzesniak, D. & Van Gunsteren, W. F. Catalytic mechanism of cyclophilin as observed in molecular dynamics simulations: pathway prediction and reconciliation of X-ray crystallographic and NMR solution data. Protein Sci. 15, 2544–2551 (2006)
Howard, B. R., Vajdos, F. F., Li, S., Sundquist, W. I. & Hill, C. P. Structural insights into the catalytic mechanism of cyclophilin A. Nature Struct. Biol. 10, 475–481 (2003)
Ke, H. M. & Huai, Q. Crystal structures of cyclophilin and its partners. Front. Biosci. 9, 2285–2296 (2004)
Lang, P. T. et al. Automated electron-density sampling reveals widespread conformational polymorphism in proteins. Protein Sci. (submitted)
Eisenmesser, E. Z., Bosco, D. A., Akke, M. & Kern, D. Enzyme dynamics during catalysis. Science 295, 1520–1523 (2002)
Halle, B. Biomolecular cryocrystallography: structural changes during flash-cooling. Proc. Natl Acad. Sci. USA 101, 4793–4798 (2004)
Rasmussen, B. F., Stock, A. M., Ringe, D. & Petsko, G. A. Crystalline ribonuclease A loses function below the dynamical transition at 220 K. Nature 357, 423–424 (1992)
Loria, J. P., Rance, M. & Palmer, A. G. A TROSY CPMG sequence for characterizing chemical exchange in large proteins. J. Biomol. NMR 15, 151–155 (1999)
Millet, O., Loria, J. P., Kroenke, C. D., Pons, M. & Palmer, A. G. The static magnetic field dependence of chemical exchange linebroadening defines the NMR chemical shift time scale. J. Am. Chem. Soc. 122, 2867–2877 (2000)
Kofron, J. L., Kuzmic, P., Kishore, V., Colonbonilla, E. & Rich, D. H. Determination of kinetic constants for peptidyl prolyl cis-trans isomerases by an improved spectrophotometric assay. Biochemistry 30, 6127–6134 (1991)
Farrow, N. A., Zhang, O. W., Forman-Kay, J. D. & Kay, L. E. A Heteronuclear correlation experiment for simultaneous determination of 15N longitudinal decay and chemical exchange rates of systems in slow equilibrium. J. Biomol. NMR 4, 727–734 (1994)
Zydowsky, L. D. et al. Active site mutants of human cyclophilin A separate peptidyl-prolyl isomerase activity from cyclosporine A binding and calcineurin inhibition. Protein Sci. 1, 1092–1099 (1992)
Frauenfelder, H. et al. Thermal expansion of a protein. Biochemistry 26, 254–261 (1987)
Frauenfelder, H., Petsko, G. A. & Tsernoglou, D. Temperature-dependent X-ray diffraction as a probe of protein structural dynamics. Nature 280, 558–563 (1979)
Tilton, R. F., Dewan, J. C. & Petsko, G. A. Effects of temperature on protein structure and dynamics: X-ray crystallographic studies of the protein ribonuclease-A at nine different temperatures from 98 to 320K. Biochemistry 31, 2469–2481 (1992)
Beach, H., Cole, R., Gill, M. L. & Loria, J. P. Conservation of μs-ms enzyme motions in the apo- and substrate-mimicked state. J. Am. Chem. Soc. 127, 9167–9176 (2005)
Tokuriki, N. & Tawfik, D. S. Protein dynamism and evolvability. Science 324, 203–207 (2009)
Lee, G. M. & Craik, C. S. Trapping moving targets with small molecules. Science 324, 213–215 (2009)
Kiefersauer, R. et al. A novel free-mounting system for protein crystals: transformation and improvement of diffraction power by accurately controlled humidity changes. J. Appl. Crystallogr. 33, 1223–1230 (2000)
Davis, D. G., Perlman, M. E. & London, R. E. Direct measurements of the dissociation-rate constant for inhibitor-enzyme complexes via the T 1ρ and T 2(CPMG) methods. J. Magn. Reson. Ser. B 104, 266–275 (1994)
Hu, J. S., Grzesiek, S. & Bax, A. Two-dimensional NMR methods for determining χ1 angles of aromatic residues in proteins from three-bond JC'Cγ and JNCγ couplings. J. Am. Chem. Soc. 119, 1803–1804 (1997)
Southworth-Davies, R. J., Medina, M. A., Carmichael, I. & Garman, E. F. Observation of decreased radiation damage at higher dose rates in room temperature protein crystallography. Structure 15, 1531–1541 (2007)
Otwinowski, Z. & Minor, W. in Macromolecular Crystallography Part A 307–326 (Methods in Enzymology, Vol. 276, Academic, 1997)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Holton, J. & Alber, T. Automated protein crystal structure determination using ELVES. Proc. Natl Acad. Sci. USA 101, 1537–1542 (2004)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Laskowski, R. A., Macarthur, M. W., Moss, D. S. & Thornton, J. M. Procheck—a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993)
Delano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, Palo Alto, 2008); 〈http://www.pymol.org〉
Theobald, D. L. & Wuttke, D. S. THESEUS: maximum likelihood superpositioning and analysis of macromolecular structures. Bioinformatics 22, 2171–2172 (2006)
Mulder, F. A. A., Mittermaier, A., Hon, B., Dahlquist, F. W. & Kay, L. E. Studying excited states of proteins by NMR spectroscopy. Nature Struct. Biol. 8, 932–935 (2001)
Delaglio, F. et al. NMRPipe—a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995)
Johnson, B. A. & Blevins, R. A. NMR View—a computer program for the visualization and analysis of NMR data. J. Biomol. NMR 4, 603–614 (1994)
Jeener, J., Meier, B. H., Bachmann, P. & Ernst, R. R. Investigation of exchange processes by 2-dimensional NMR-spectroscopy. J. Chem. Phys. 71, 4546–4553 (1979)
Acknowledgements
We thank S. Marqusee and B. Krantz for discussions; S. Classen, G. Meigs, J. Holton, A. Samelson, N. Echols, P. Afonine, and the Phenix team for technical support; J. Tainer for access to Rigaku free-mounting device at ALS Beamline 12.3.1; J. Pelton and D. Wemmer for providing essential help and access to NMR facilities. J.S.F. was supported by US NSF and Canadian NSERC fellowships. This work was funded by the US National Institutes of Health (to T.A.) and the US National Institutes of Health, the US Department of Energy Office of Basic Energy Sciences, and the Howard Hughes Medical Institute (to D.K.).
Author Contributions J.S.F., S.C.D. and R.E. performed the X-ray experiments, M.W.C. performed the NMR experiments, and M.W.C. and J.S.F. performed the activity and binding assays. J.S.F., M.W.C., D.K. and T.A. analysed data and wrote the paper. All authors contributed to data interpretation and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-11 with Legends and Supplementary Tables 1-4. (PDF 7505 kb)
Rights and permissions
About this article
Cite this article
Fraser, J., Clarkson, M., Degnan, S. et al. Hidden alternative structures of proline isomerase essential for catalysis . Nature 462, 669–673 (2009). https://doi.org/10.1038/nature08615
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08615
This article is cited by
-
Pushed to extremes: distinct effects of high temperature versus pressure on the structure of STEP
Communications Biology (2024)
-
Computational remodeling of an enzyme conformational landscape for altered substrate selectivity
Nature Communications (2023)
-
Ratcheting synthesis
Nature Reviews Chemistry (2023)
-
A Minireview on Temperature Dependent Protein Conformational Sampling
The Protein Journal (2021)
-
DNA mismatches reveal conformational penalties in protein–DNA recognition
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.