Abstract
Schistosoma mansoni is responsible for the neglected tropical disease schistosomiasis that affects 210 million people in 76 countries. Here we present analysis of the 363 megabase nuclear genome of the blood fluke. It encodes at least 11,809 genes, with an unusual intron size distribution, and new families of micro-exon genes that undergo frequent alternative splicing. As the first sequenced flatworm, and a representative of the Lophotrochozoa, it offers insights into early events in the evolution of the animals, including the development of a body pattern with bilateral symmetry, and the development of tissues into organs. Our analysis has been informed by the need to find new drug targets. The deficits in lipid metabolism that make schistosomes dependent on the host are revealed, and the identification of membrane receptors, ion channels and more than 300 proteases provide new insights into the biology of the life cycle and new targets. Bioinformatics approaches have identified metabolic chokepoints, and a chemogenomic screen has pinpointed schistosome proteins for which existing drugs may be active. The information generated provides an invaluable resource for the research community to develop much needed new control tools for the treatment and eradication of this important and neglected disease.
Similar content being viewed by others
Main
Schistosomiasis is a neglected tropical disease that ranks with malaria and tuberculosis as a major source of morbidity affecting approximately 210 million people in 76 countries, despite strenuous control efforts1. It is caused by blood flukes of the genus Schistosoma (phylum Platyhelminthes), which exhibit dioecy and have complex life cycles comprising several morphologically distinct phenotypes in definitive human and intermediate snail hosts. Schistosoma mansoni, one of the three major human species, occurs across much of sub-Saharan Africa, parts of the Middle East, Brazil, Venezuela and some West Indian islands. The mature flukes dwell in the human portal vasculature, depositing eggs in the intestinal wall that either pass to the gut lumen and are voided in the faeces, or travel to the liver where they trigger immune-mediated granuloma formation and peri-portal fibrosis2. Approximately 280,000 deaths per annum are attributable to schistosomiasis in sub-Saharan Africa alone3. However, the disease is better known for its chronicity and debilitating morbidity4. A single drug, praziquantel, is almost exclusively used to treat the infection but this does not prevent reinfection, and with the large-scale control programmes in place, there is concern about the development of drug resistance. Indeed, resistance can be selected for in the laboratory and there are reports of increased drug tolerance in the field5.
In this study we present the sequence and analysis of the S. mansoni genome. Previous metazoan projects have been restricted to Deuterostomia (for example, Homo, Mus and Ciona) and the ecdysozoan clade of the Protostomia (for example, Drosophila, Caenorhabditis and Brugia). Together with the accompanying article on S. japonicum6, we present, to our knowledge, the first descriptions of metazoan genomes from the lophotrochozoan clade. The genome reveals features that aid our understanding of the evolution of complex body plans. We have mined the genome to predict new drug targets, on the basis of searches involving traditional areas for drug discovery, metabolic reconstruction, and bioinformatics screens that exploit shared pharmacology. It is hoped that these and other targets will accelerate drug discovery, generating the much needed new treatments for the control and eradication of schistosomiasis.
Genome structure and content
The nuclear genome sequence of S. mansoni was determined by whole-genome shotgun sequencing and assembled into 5,745 scaffolds greater than 2 kilobases (kb) (Supplementary Table 1), totalling 363 megabases (Mb). Although 40% of the genome is repetitive, 50% is assembled into scaffolds of at least 824.5 kb. Furthermore, 43% of the genome assembly (distributed over 153 scaffolds) was unambiguously assigned to chromosomes (seven autosomal, plus ZW sex-determination pairs) using fluorescence in situ hybridization (FISH; Fig. 1, Supplementary Fig. 1 and Supplementary Table 2).
a, b, Idiogram of S. mansoni chromosomes W, Z (a) and 3 (b). S. mansoni BAC clones were mapped to the karyotype of S. mansoni by FISH. The solid black areas are heterochromatin, the open areas are euchromatin. The BAC clones are identified by BAC numbers. c–f, Chromosome spreads with FISH-mapped BACS are shown. FISH-mapped BACS are identified by arrowheads on labelled chromosomes. Scale bar, 10 μm. See Supplementary Fig. 1 for idiograms of all S. mansoni chromosomes.
We identified 72 families of both long-terminal repeat (LTR) and non-LTR transposons, comprising 15% and 5% of the genome, respectively, and containing 63 and 60 new families each (Supplementary Table 3). The LTR transposons are from the Ty3/Gypsy and BEL clades, whereas the non-LTR transposons are restricted to the RTE, CR1 and R2 clades. Two previously described non-LTR retrotransposon families from the RTE clade (SR2 and Perere-3)7,8 seem to have undergone a burst of transposition events after divergence of S. mansoni and S. japonicum, and contribute to an overall higher representation of non-LTR retrotransposons in S. mansoni (15%, around 8% in S. japonicum). A new DNA transposon belonging to the Mu family was also found, which represents the first instance in a flatworm. The presence of target site duplications in some copies indicates recent transposition, and suggests that active copies may still exist in the genome. A lack of terminal inverted repeats, a feature of Mu family members, suggests a peculiar mechanism for recognition of this element by the transposition apparatus.
We identified 11,809 putative genes encoding 13,197 transcripts. Considering genes that do not span a gap, the average gene size is 4.7 kb, typically with large introns (the average is 1,692 base pairs (bp)) and much smaller exons (the average is 217 bp). Moreover, the introns show a markedly skewed size distribution that has not been observed in other eukaryotes, whereby 5′ introns are smaller than 3′ introns (Fig. 2, Supplementary Information and Supplementary Table 5). In multi-exon genes, the first few introns can be as small as 26 bp, whereas introns towards the 3′ end are typically kilobases in length (the largest is 33.8 kb). The reason for this is unclear but it suggests unusual transcriptional control. However, a survey of conserved transcription factor domains shows S. mansoni to be broadly similar to other eukaryotes (Supplementary Information, Supplementary Fig. 2 and Supplementary Table 6). It is noteworthy that 43% of transcription factor families with schistosome representatives also contained vertebrate sequences, nearly twice the number that matched nematode worms, emphasizing their evolutionary distance.
The length of introns varies according to their position in a transcript, counting from the 5′ end (solid circles) and the 3′ end (open circles). Mean lengths ± standard errors are shown. After about five introns, the length difference is no longer apparent owing to the variation in the number of introns per transcript (see Supplementary Information).
Micro-exon genes
At least 45 genes have an unusual micro-exon structure. Individual micro-exons have been described in other genomes, dispersed among several normal exons9. However, S. mansoni is notable in containing micro-exon genes (MEGs) that comprise 75% of the coding sequence, are flanked at the 5′ and 3′ extremes by conventional exons, and have lengths that are multiples of three bases (from 6 to 36).
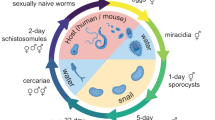
Other than having shared gene structure, no similarity could be detected between 14 MEG families (each with up to 23 members; Fig. 3 and Supplementary Table 7). Moreover, they showed no similarity with annotated genes from outside Schistosoma spp., nor any identifiable motifs or functional domains. Comparisons between MEG family members and related proteins from S. japonicum suggest that some gene duplication events preceded the divergence of the two species. Almost all encode a signal peptide at the 5′ end and three have membrane anchors, so most are probably secreted. Examination of the large expressed-sequence tag (EST) data set from across the life cycle shows that genes from all MEG families are transcribed in the intramammalian stages of the life cycle, and the germ balls of daughter sporocysts that develop into infective cercariae, but probably not in miracidia that infect the snail intermediate host (Fig. 3).
a, Structure of a representative member from each MEG family. Where several members were found, the total number detected is indicated in parentheses. Each box represents an exon drawn to scale, and the number above it indicates the exon size in nucleotides. For illustrative purposes, the introns are shown with fixed length. Black triangles and diamonds indicate exons encoding predicted signal peptides and transmembrane helices, respectively. Other characteristics associated with exons are indicated by colour and grouped as follow: micro-exons having lengths that either are multiples of 3 bp (red) or are indivisible by 3 bp (orange); exons longer than 36 bp and having lengths that either are multiples of 3 bp (blue) or are indivisible by 3 bp (green); putative initiation and termination exons (grey); untranslated region (UTR) (black). The asterisk indicates an exon deduced from transcript data, which did not match the sequenced genome. MEG-12 and MEG-13 structures were only partially predicted owing to the lack of transcripts containing the 5′ end of these genes. b, PCR with reverse transcription (RT–PCR) or EST-based evidence of transcription (black box) for each family across different life-cycle stages. C, cercaria; E, egg; G, germball; M, miracidium; 3s and 7s, 3- and 7-day schistosomula; 21li and 28li, 21- and 28-day liver worms; 45a, 45-day adult worm pairs.
Sequencing of transcripts from three MEG families revealed the occurrence of several alternative splice variants formed by exon skipping. In one of the families analysed, all internal exons except those coding for the signal peptide were missing in at least one transcript sampled, and a gene from a second family presented different transcripts with extended exons produced by the use of alternative splicing sites. These observations suggest that a ‘pick and mix’ strategy is used to create protein variation.
Evolution of triploblasty, parasitism and tissues
Schistosomes are the first Platyhelminthes to be fully sequenced, and provide insights into the evolution of ‘simple’ animals. Using Treefam to make comparisons with the sea anemone Nematostella vectensis, a representative of the Radiata, we sought gene families restricted to, or expanded in, the Bilateria (Supplementary Table 8). The advent of a third germ layer in flatworms is paralleled by the expansion of genes encoding cell adhesion molecules such as cadherins. Similarly, tissue-patterning developmental cues (for example, Notch/Delta) and histone-modifying enzymes (for example, histone acetyltransferases) have proliferated. Some genes, such as the tetraspanins that encode membrane structural proteins, have greatly proliferated in schistosomes, suggesting a critical role in worm physiology/parasitism. The large array of paralogues for fucosyl and xylosyltransferases involved in the generation of new glycans expressed at the host–parasite interface may be important for subverting the immune system. The expansion of proteases in schistosomes also seems to be directly related to parasitism, because it includes families involved in host invasion (invadolysins) and blood feeding (cathepsins). Furthermore, G-protein-coupled receptors (GPCRs) show varying levels of contraction in schistosomes, whereas several classes (for example, peropsins) are greatly expanded in Nematostella, indicating functions associated with the free-living lifestyle.
Although schistosomes are acoelomate, they possess tissues approaching the sophistication of organs—such as gut, nephridia, nerve and muscle—that are concerned with discrete physiological processes, such as feeding, excretion and locomotion. However, as lophotrochozoans they are evolutionarily distant from the previously sequenced parasitic nematodes Brugia10 and Meloidogyne11,12 (both ecdysozoans). Compartmentalisation of schistosome tissues and the formation of epithelial barriers are crucial for life in the hostile environment of the host bloodstream. Schistosomes possess the typical machinery of higher metazoa to interact with the cytoskeleton and control cell polarity (Supplementary Information and Supplementary Table 9), organize epithelia and denote tissue boundary lines.
S. mansoni possess a nervous system that includes an anterior brain and longitudinal nerve cords, which extend from the brain to run the length of the worm body. Furthermore, a variety of sensory structures (at least six types in the cercaria13) are able to transduce a wide range of stimuli that assist in host location, penetration and navigation through the vasculature. In common with more complex organisms, schistosomes possess the tools needed to mediate neurogenesis and control axon growth cones and migration of neural cells (Supplementary Information and Supplementary Table 9), supporting the ancient origins of neural complexity.
Insights into possible new drug targets
Historically, anti-schistosomiasis agents were identified by in vivo screening in animal models. The S. mansoni genome project makes a more target-based approach to drug discovery feasible, and some promising leads have already emerged. These include a family of nuclear receptors14 (Supplementary Information) and a redox enzyme, thioredoxin glutathione reductase, recently validated as a drug target15. The condensed redox biochemistry of S. mansoni, relative to its human host, may offer further drug development targets (Supplementary Information). In the context of drug discovery, we have explored other potential areas of vulnerability, including: lipid metabolism, GPCRs, ligand- and voltage-gated ion channels, kinases, proteases and neuropeptides. We also undertook two bioinformatics-led approaches: metabolic reconstruction to identify chokepoints, and sequence searches for structures related to known drug targets.
Lipid metabolism
S. mansoni contains a full complement of genes required for most core metabolic processes, such as glycolysis, tricarboxylic acid cycle and the pentose phosphate pathway. However, schistosomes are incapable of de novo synthesis of sterols or free fatty acids and must use complex precursors from the host16. An extensive lipid-carrying protein repertoire could be identified, but despite producing precursors for fatty acid synthesis, fatty acid synthase could not be identified. An inability to use isoprene products of the mevalonate pathway probably accounts for the lack of sterol biosynthesis (Supplementary Table 11 and Supplementary Information). The genes necessary for a complete β-oxidation pathway are present, and this usually inactive pathway might operate in reverse to perform syntheses17. Despite constituting 40% or more of the lipid content of adult worms16, triacylglycerol has an uncertain role in the schistosome’s life cycle—it is slow to turn over, does not contribute to the formation of other lipids16 and its use as an energy store is doubtful17. Nevertheless, S. mansoni possesses lipases capable of breaking down triacylglycerol, so it may have functions other than preventing too high concentrations of intracellular fatty acids16. Pathways responsible for synthesizing the phospholipid components of membranes are well represented, except that phosphatidylcholine must be derived from diacylglycerol18 and the parasite must depend on its host as a source of inositol.
GPCRs, ligand-gated and voltage-gated ion channels
GPCRs, ligand-gated and voltage-gated ion channels are targets for 50% of all current pharmaceuticals19. At least 92 putative GPCR-encoding genes are present (Supplementary Table 12), the bulk (82) of which are from the rhodopsin family. The largest groups are the α-subfamily (30), which includes amine receptors, and the β-subfamily (24), which contains neuropeptide and hormone receptors. The diversity of the former subfamily underlines the wide range of potential amine/neurotransmitter reactivities of schistosomes, but the tentative identities assigned need to be confirmed by functional studies, as has already been performed for a histamine receptor20. Schistosomes detect chemosensory cues, but a large, unique clade of the mediating receptors was not found. However, the 26 ‘orphan’ rhodopsin family GPCRs may include proteins with this role. Outside the large rhodopsin family, representatives from each of the smaller families of GPCRs, glutamate family (2), frizzled family (3), and the secretin/adhesion family (4) are present.
Each of the three major ligand-gated ion channel families—the Cys-loop family, glutamate-activated cation channels, and ATP-gated ion channels—are represented in the schistosome genome. Of the 13 Cys-loop family ligand-gated ion channels, nine encode nicotinic acetylcholine receptor subunits (Supplementary Fig. 4 and Supplementary Table 13). The remaining four anion channel subunits group among GABA (γ-aminobutyric acid), glycine and glutamate receptors, but it is not possible to assign precise identities. The seven schistosome glutamate-activated cation channels comprise at least two sequences from each of the three common sub-groupings. The presence of a functional P2X receptor for ATP-mediated signalling in schistosomes was already known21, and the data here show at least four more.
Voltage-gated ion channels generate and control membrane potential in excitable cells, and are central to ionic homeostasis. There are examples of successful drugs targeting voltage-gated sodium, potassium and calcium channels22. Although voltage-gated sodium channels were not found, at least 41 members from each of the major six transmembrane (6TM) and four transmembrane (4TM) families of potassium channels (Supplementary Table 14) are present. The 6TM voltage-gated potassium channel family (20 members) is the largest, including the well-characterized Kv1.1 channel found in nerve and muscle of adult schsitosomes23. Other classes of 6TM potassium channels include the KQT channels, large calcium-activated channels, small calcium-activated channels, and cyclic-nucleotide-gated groups. This last group, comprising eight members, is most often associated with signal transduction in primary olfactory and visual sensory cells (Caenorhabditis elegans has only five; ref. 24). S. mansoni possesses six 4TM inward-rectifying TWIK-related potassium channels (about 46 in C. elegans). There are four α and two β subunits of voltage-gated calcium channels in schistosomes, and a β subunit is implicated as a molecular target of the anti-schistosomal praziquantel25.
The kinome
Protein kinases are important regulators of many different cellular functions. Both they and their inhibitors have entered the drug development pipeline in recent years26 but few schistosome kinases have been characterized to date. The S. mansoni genome encodes 249 kinases, including 22 genes with alternative splicing (Supplementary Information). This corresponds to 1.9% of the total coding proteins in the genome, a figure comparable to that found in other species27 (Supplementary Fig. 6). S. mansoni possesses representatives of all of the main kinase groups (Supplementary Fig. 7), the largest of which is the CMGC (cyclin-dependent kinases, mitogen-activated protein kinases, glycogen synthase kinase 3 and CK2-related kinases) group, in contrast to other analysed eukaryotic genomes. However, a single class (RCK) is absent from the CMGC family, a deficiency shared with yeast but not nematodes or mammals.
The least represented groups are the casein kinase (CK1) and receptor guanylate cyclase families with only seven and three members, respectively, contrasting with C. elegans, in which casein kinase is the largest group and receptor guanylate cyclase has 27 members. CK1 (and CMGC) group members that are expressed in sperm or during spermatogenesis in C. elegans are missing in S. mansoni.
The degradome
Proteolytic enzymes (proteases), making up an organism’s ‘degradome’28, operate in virtually every biological and pathological phenomenon29 and are proven drug targets in diverse biomedical contexts30,31. All five major classes of proteases (aspartic, cysteine, metallo-, serine and threonine) are represented as various clans (mechanistically related groups) in the parasite genome (Supplementary Table 17). The percentage distribution of the major clans is generally similar to that of the human host with some notable exceptions, mainly owing to the expansion of constituent protease families in humans. Of the 73 protease families, 61 are found in humans and in S. mansoni, and 60 families are shared. With 335 sequences, proteases comprise 2.5% of the putative proteome (Supplementary Table 18), consistent with the proportion in other organisms (1–5%), but this is only one-third of that in humans (945 sequences, if A2 family retrovirus and retrotransposon proteases are included).
The greatest difference between host and parasite is in the paucity of chymotrypsin-like S1 family enzymes in the latter (22 versus 135 human sequences). This reflects the evolution and diversification of family S1 for complex and highly regulated proteolysis cascades in vertebrates and some invertebrates, such as innate immunity, development, blood coagulation and complement activation32,33,34. From a therapeutic standpoint, the reduced complexity may prove valuable with fewer parasite proteases available for essential life-sustaining functions. For example, robust drug discovery programmes are in place for chymotrypsin-like S1 families35 and peptidase C14 (caspases)36, on which anti-schistosomal drug discovery could ‘piggy-back’37. It is also notable that a smaller number of schistosome protease families (for example, C1, M8 and M13) have more members than the respective families in humans. C1 proteases are involved in nutrient digestion by the parasite, which contrasts with the S1 enzymes used in the host. This disparity has already been exploited for a promising anti-schistosome therapy38. One protease family (C83) is apparently unique to S. mansoni.
Apart from the degradome, but involved in its modulation, 34 protease inhibitors were found (Supplementary Table 19). Most of these are serine protease inhibitors belonging to families I2 (Kunitz-type) and I4 (serpins). Two inhibitors of cysteine proteases (cystatins39,40) and two α-2-macroglobulin homologues (I39) were also identified, as were three inhibitor of apoptosis proteins (I32), one of which is highly expressed in adults, where it may function to regulate one or more of the four schistosome caspases.
Neuropeptides
Thirteen putative neuropeptides were identified (Supplementary Table 20), indicating that schistosomes may have much greater diversity than the two described previously. Apart from the neuropeptide Fs (NPFs), most are apparently restricted to the Platyhelminthes—their absence from humans making them a credible source of anthelmintic drug leads. The predicted product of npp-6 (the amidated heptapeptide AVRLMRLamide) resembles molluscan myomodulin, whereas the two NPP-13 peptides show 100% carboxy-terminal identity with vertebrate neuropeptide-FF-like peptides (peptides ending with a C-terminal sequence PQRFamide); neither of these has previously been reported in any non-vertebrate organism. The discovery of a second NPF (NPP-21b) as well as the known NPP-21a41 is reminiscent of the vertebrate neuropeptide Y (NPY) superfamily, and strengthens the argument that NPFs and NPYs have a common ancestry.
Metabolic chokepoints
A chokepoint analysis of metabolic pathways reconstructed from the S. mansoni genome was used to identify further targets. A total of 607 enzymatic reactions could be placed in pathways, and 120 of these enzymes were identified as chokepoints (Supplementary Table 21). The list of chokepoints includes many that are drug targets in other organisms as well as target reactions already characterized in S. mansoni, validating the approach (Supplementary Information). The list also contains new candidate targets and comprises approximately 1% of the S. mansoni proteome.
Chemogenomics screening
In the context of neglected tropical diseases and with constrained investment in drug discovery, piggy-backing37 or ‘drug-repositioning’ strategies42 that re-use existing drugs offer potential time-saving and cost benefits. We adopted a twofold strategy to find significant matches between proteins from the parasite and known ‘druggable’ protein targets of the human host and human-infective pathogens. Using conservative parameters of >50% sequence identity over >80% of the target, we first performed a similarity search against a database of targets curated from medicinal chemistry literature. This revealed 240 distinct S. mansoni transcripts with matches to targets against which there are high quality compounds (Supplementary Table 22). Given the need for short-course, oral therapies against schistosomiasis, this list was further reduced to 94 S. mansoni targets by filtering for potency and predicted bioavailability. A second search, against a database of the targets for human-directed drugs, showed 66 significant matches with pharmaceuticals marketed at present (Supplementary Table 23), corresponding to 34 S. mansoni targets (26, after representing multicopy genes as a single instance; Table 1). For instance, disulfiram, for controlling substance abuse, was highlighted as a potential anti-schistosomal drug; its anti-parasite properties have already been investigated43. Manual inspection of the list for compounds with side effects and toxicity can further refine choices—for example, by eliminating the immunosuppressants, cyclosporin and rapamycin. The remaining known drugs could be directly tested in animal models, and either applied unmodified in anti-schistosomal therapy, or could serve as leads for further optimisation. Widening the search beyond the initial strict criteria would expand opportunities, for example, topoisomerase 1 is retrieved below our initial threshold, at 71% identity but only 58% overlap.
Conclusion
A century after Louis Sambon first named the species in 1907 (ref. 44), the sequencing of the S. mansoni genome is a landmark event. The sequence provides the scientific community with several avenues to study this under-researched human pathogen, and will drive future evolutionary, genetic and functional genomic research. Not least, given that just one drug is widely available to treat schistosomiasis at present, the genome sequence, including the genome-mining analysis presented, offers the possibility that new drug candidates will soon be identified.
Methods Summary
Mixed-sex cercariae from the Puerto Rico isolate of S. mansoni45, released from infected Biomphalaria glabrata snails, were placed in low-melting agarose plugs and genomic DNA was prepared by standard methods. Approximately sixfold coverage of the nuclear genome was obtained using a whole-genome shotgun sequencing approach, in which libraries of different cloned insert sizes (in plasmid, fosmid and bacterial artificial chromosomes (BAC) vectors) were randomly sequenced by Sanger technology from either end. Sequence reads were assembled, and scaffolds were FISH-mapped to individual chromosomes where possible (Supplementary Table 2). The output of several gene prediction algorithms, trained using 409 manually curated gene structures, were integrated into a single set of gene predictions (version 4), which were used for subsequent analyses. Data were accessed from GeneDB (http://www.genedb.org), and Artemis was used for manual annotation and curation of a further 958 genes during subsequent analyses (as described previously46).
Online Methods
Genome sequencing, assembly and mapping
The most commonly used Puerto Rican strain of S. mansoni45 was maintained in albino B. glabrata snail and NMRI mice and golden hamsters as laboratory hosts (Mesocricetus auratus). Cercariae released from infected snails were resuspended in PBS at a concentration of 5 × 105 cercariae ml-1. The parasites were transferred to a 42 °C water bath, incubated for 5 min, and mixed with an equal volume of 1.2% low-melting point agarose (Gibco-BRL) in PBS at 42 °C. The agarose/cell mixture was transferred to a disposable plug mould (Bio-Rad), placed on ice, treated twice for 24 h at 50 °C with 1% N-lauroyl sarcosine, 0.5 M EDTA, pH 8.0, 2 mg ml-1 proteinase K (Boehringer Mannheim). Proteinase K was then inhibited by a 30-min treatment with PMSF (40 μg ml-1), followed by three successive 30-min dialyses against 10 mM Tris-HCl, pH 8.0, 0.1 mM EDTA. Sequencing libraries were constructed using genomic DNA extracted from mixed-sex cercariae. Sequencing reads were produced from small insert plasmid clones containing a range of insert sizes. In addition, 12,305 BAC end sequences from S. mansoni BAC library Sm1 (DDBJ/EMBL/GenBank accession numbers BH199420–BH211620), 19,136 CHORI 103 BAC end sequences (DDBJ/EMBL/GenBank accession numbers DX983724–ED003998) and 16,628 fosmid end reads were included. After filtering out low quality reads, 85% of the remaining 3.19 million reads were assembled using Phusion47 into 381 Mb, or 363 Mb after filtering small (<2 kb) contigs (see Supplementary Table 1). The size is greater than the previously estimated size of 270 Mb48, although this size estimate can be revised to 300 Mb because the original measurements were made using the E. coli genome as a control, which has a length that is 10% greater than previously thought. From the assembly, a depth of coverage of ∼sixfold was calculated.
A physical map was generated using FISH to localize S. mansoni BACs to the seven autosomal and sex pairs of chromosomes using previously published methods49. Clones from two BAC libraries, Sm1 (ref. 50) and CHORI-103 (http://bacpac.chori.org/schis103.htm), each constructed from cercarial DNA were randomly picked and subjected to FISH analysis. Owing to the repetitive nature of the schistosome genome, BACs would often hybridize to more than one chromosome. This was in spite of using sheared genomic DNA to block the repetitive sequences. Of the 500 clones analysed, 334 showed unique hybridization patterns (Fig. 1, Supplementary Fig. 1 and Supplementary Table 2). A total of 118 BACs that were FISH-mapped were among those end-sequenced, and 153 scaffolds were assigned to a specific chromosome.
Retroelements analysis
We performed an iterative search of retroelements using the conserved reverse transcriptase domain as previously described51. Elements with higher than 80% nucleotide identity in the reverse-transcriptase region were considered as members of the same family. To obtain an unbiased estimate of abundance for each element in the genome, all the identified families were mapped to the shotgun reads using BLASTN52. The number of bases spanned by the alignment for each element was counted and compared with the total number of bases in the shotgun data to determine their representation in the S. mansoni genome.
Genome annotation and repeat content analysis
A training set (for ab initio gene finding) of 409 genes was manually curated from S. mansoni sequences already within the Uniprot database and manual prediction of highly conserved genes. Further genome-wide gene predictions were made using both EVidenceModeller and PASA53. EVidenceModeller uses an evidence-combining strategy to compute an optimal set of protein-coding gene structures derived from several, often conflicting, sources of gene predictions. The sources of evidence for our annotation of the S. mansoni genome included the following: ab initio gene predictions derived from GlimmerHMM54, TWINSCAN55, and Augustus56; protein sequence homologies to a non-redundant protein database using AAT57; cross-genome sequence homologies between S. mansoni and S. japonicum using PROmer58; spliced genome alignments to ESTs using GMAP59; and repeat regions identified using RepeatScout60 and RepeatMasker (A. F. A. Smit, R. Hubley and P. Green, unpublished observations, http://www.repeatmasker.org). Consensus gene predictions generated by EVidenceModeller were further modified to include annotations of untranslated regions and alternative splicing isoforms for 1,038 genes by applying PASA leveraging the earlier GMAP aligned ESTs. A total of 13,197 transcripts were predicted for 11,809 genes. Of the 30,110 previously described EST clusters, 24,373 map to contigs >1 kb in the current genome assembly. The true number of genes could therefore be as high as 17,500. By parsing BLAST description lines, putative products were assigned to each gene. During the course of subsequent analyses, 958 of these were manually edited using the Artemis annotation tool.
For an unusually large gene, encoding a putative ryanodine receptor spanning ∼164 kb, 79 of its 93 intron–exon boundaries were confirmed by RT–PCR. Approximately 45% of the S. mansoni genome was found to be repetitive, computed by summing up genomic bases matching known S. mansoni mobile element sequences or repeat family consensus sequences derived from the RepeatScout de novo repeat library. The repeat content was also assessed on the basis of the distribution of random sequences 25 nucleotides in length, 104,028,213 out of 373,600,457 or 28% of bases were repetitive. Note, this value is significantly lower than that of RepeatMasker because the latter allows sequence divergence of up to 20%.
Analysis of putative transcription factors
Profile hidden Markov models (HMMs) of domains present in the proteins that constitute the TRANSFAC eukaryotic transcriptional factor database61 and the DBD DNA-binding domain database62 were used to search the genome of S. mansoni in conjunction with 63 other eukaryotic genomes. The score threshold was defined as the lowest pairwise score among all members of the Pfam family associated to the HMM. The putative transcriptional activator proteins were then clustered on the basis of sequence similarities (BLASTP E value ≤ 10-6 considered significant) using the TRIBE-MCL algorithm63 and an inflation value of 2.0 (ref. 64).
Micro-exon genes
MEGs were predicted as previously described9 with further manual refinement using available S. mansoni EST data (including both published data65 and unpublished data from GenBank/dbEST or ftp://ftp.sanger.ac.uk/pub/pathogens/Schistosoma/mansoni/ESTs/). Further family members were identified by similarity searches against the available supercontigs in the assembly with long flanking MEG exons as query sequences. Signal peptides and transmembrane domains were detected using SignalP66 and TMHMM 2.0 (ref. 67) programs, respectively.
Expression of MEG families at different stages throughout the life cycle was analysed by BLAST searching the sequences of all members of a family against the complete S. mansoni EST data set, which comprises ESTs from the following developmental stages: germball (28,497), cercaria (21,639), 3-day somule (6,122), 7-day somule (41,043), 21-day liver worm (6,044), 28-day liver worm (11,227), 45-day adult worm (59,552), egg (33,674) and miracidium (19,982).
Evolutionary analysis
To identify orthologues and paralogues of S. mansoni genes, we built a standalone version of the TreeFam database (version 7) of animal gene families68,69. For each S. mansoni predicted protein, we identified the top-matching TreeFam ‘clean’ family using HMMER70 (with E ≤ 10-10 as a cutoff). Similarly, the top-matching family was identified for each Nematostella vectensis (release version 1.0)71 and S. japonicum protein. Trees and alignments were built for the families as for the standard TreeFam pipeline. This resulted in trees for 5,829 families that contain S. mansoni, S. japonicum or N. vectensis genes. From these trees, we identified within-species paralogues in the three species, and identified the ancestral taxon in which the duplication that gave rise to each pair of paralogues occurred.
Kinome
A eukaryotic protein kinase domain HMM was built from a manually adjusted alignment of 68 diverse kinase domain sequences from yeast, worm, fly and human that share <50% sequence identity in the catalytic domain. To test the selectivity of the model, it was run against the Uniprot database. Using a P < 0.1 cutoff, the model detected 2,688 putative domains, all of which were annotated either as kinases or putative kinases in different description fields. Local and global HMM models were built with the HMMer package (http://hmmer.janelia.org/) from several sequence alignments generated by MAFFT software72 and were used for sensitive searches against the S. mansoni database.
Identified genes were annotated using Artemis, integrating data from Interproscan and Reverse PSI-BLAST searches73 and the size of the S. mansoni kinome was compared with those of: Plasmodium falciparum74, Homo sapiens27, Trypanosoma cruzi75, Trypanosoma brucei76, C. elegans77, Leishmania major78 and Mus musculus79. A dendogram was constructed using the kinase domains of the identified proteins with the CLC Main Workbench (CLC bio) using the neighbour-joining method with 1,000 replicates.
Identification of putative proteases and inhibitors
We used the MEROPS database80 (http://merops.sanger.ac.uk) to identify active S. mansoni proteases and protease inhibitor homologues, using BLASTP52,73 with E ≤ 10-9 as a cutoff. More distant relatives were identified through HMMER version 2.3.2 (ref. 70) searches of Pfam models81 that corresond to MEROPS families (Pfam version 22.0 (ref. 82), http://pfam.sanger.ac.uk/), using the same E-value cutoff. This initial data set contained 656 provisional homologues, having removed predicted proteases <80 residues in length as well as provisional inhibitors <50 residues long. A secondary screen against the NCBI non-redundant protein database retained a total of 369 S. mansoni sequences, which overlapped in at least 50% of their MEROPS hit or Pfam domain with an experimentally characterized protease or inhibitor homologue. False positives were removed by comparing nonspecific MEROPS description lines (for example, ‘non-peptidase homologues’) to the top non-redundant BLAST hits with an E-value at least 3 logs greater than the top MEROPS or Pfam hit but lacking associated experimental validation. This approach removed MEROPS proteins that are not functional proteases but are structurally related (such as hormone-sensitive lipases in the family S9; flagged as homologues of proteins that are inactive protease homologues in Supplementary Table 18). Similarly, the Pfam database models domains found not only in proteases and inhibitors but also in a wide range of other proteins (for example, PF00047, PF00059, PF00561, PF01476 and PF0764) were also removed as false positives in the absence of further evidence.
We next predicted which of the putative protease homologues were likely to be active or inactive. BLAST alignments of proteins against putative homologues classified in MEROPS predicted active site positions and residues in the S. mansoni query sequence, followed by manual inspection of sequence alignments to refine active site residue predictions. In a few cases, in which an acceptable alignment was not produced by BLASTP of MEROPS, a non-redundant sequence was used. In more difficult cases, involving two closely related S. mansoni sequences, active site residues were identified from multiple alignments of S. mansoni sequences, a representative sequence for the corresponding MEROPS family, and the seed alignment sequences for the relevant Pfam model.
Metabolic chokepoint analysis
An S. mansoni metabolic pathways database, SchistoCyc (http://schistocyc.schistodb.net/ptools), was created using the Pathway Tools software83, which contains algorithms to predict an organism’s metabolic pathways from its genome by comparison to MetaCyc, a reference pathways database84. From the pathway database, potential chokepoint reactions85 were identified (those that uniquely consume a specific substrate or produce a specific product). Chokepoint reactions are probably critical to normal cellular physiology and therefore represent potential drug targets.
Chemogenomics
To identify, in S. mansoni, putative proteins for which therapeutic compound or high quality chemical tools may already be available, sequence similarity searching was performed using BLASTP against the Target Dictionary from Drugstore19 (database of Food and Drug Administration approved drugs) and StARlite (a database of Structure Activity Relationship data abstracted and indexed manually from the primary literature and at present containing 440,055 unique compounds, directed against approximately 3,500 distinct molecular targets from the primary literature). The results were stringently filtered for significance: ≥50% identity, ≥80% overlap of the target and a BLAST E ≥ 10-10. To prioritise 755 hits to StARlite, we applied filters for potency/affinity against the matched target, combined with an estimate of the likelihood that the compound could be orally absorbed. The potency cutoff was set at a half-maximal inhibitory concentration (IC50), inhibition constant (Ki), or dissociation constant (Kd) of 100 nM or better, and oral bioavailability was estimated using the ‘rule of five’ (molecular weight of no more than 500 Da, clogP less than five, no more than five hydrogen bond donors and no more than ten nitrogen or oxygen atoms)86. The drugs associated with matches in the DrugStore database were classified according to a broad range of current therapeutic categories: (1) direct and clear evidence that this interaction is primarily responsible for the therapeutic action of the drug; (2) direct and clear evidence that this interaction is one mechanism for the drug but other targets or mechanisms may exist; and (3) indirect or inferred evidence of the association of the drug, target and therapeutic action.
Accession codes
Primary accessions
EMBL/GenBank/DDBJ
Data deposits
The annotated genome sequence has been submitted to EMBL with the accession numbers FN357292–FN376313. All data are also available for browsing in the GeneDB database (http://www.genedb.org/genedb/smansoni/). The CHORI BAC clones used in this study are available from http://bacpac.chori.org/. This paper is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence, and is freely available to all readers at www.nature.com/nature.
References
Steinmann, P., Keiser, J., Bos, R., Tanner, M. & Utzinger, J. Schistosomiasis and water resources development: systematic review, meta-analysis, and estimates of people at risk. Lancet Infect. Dis. 6, 411–425 (2006)
Gryseels, B., Polman, K., Clerinx, J. & Kestens, L. Human schistosomiasis. Lancet 368, 1106–1118 (2006)
van der Werf, M. J. et al. Quantification of clinical morbidity associated with schistosome infection in sub-Saharan Africa. Acta Trop. 86, 125–139 (2003)
King, C. H., Dickman, K. & Tisch, D. J. Reassessment of the cost of chronic helmintic infection: a meta-analysis of disability-related outcomes in endemic schistosomiasis. Lancet 365, 1561–1569 (2005)
Doenhoff, M. J. & Pica-Mattoccia, L. Praziquantel for the treatment of schistosomiasis: its use for control in areas with endemic disease and prospects for drug resistance. Expert Rev. Anti Infect. Ther. 4, 199–210 (2006)
The Schistosoma japonicum Genome Sequencing and Functional Analysis Consortium The Schistosoma japonicum genome reveals features of host–parasite interplay. Nature 10.1038/nature08140 (this issue)
Drew, A. C., Minchella, D. J., King, L. T., Rollinson, D. & Brindley, P. J. SR2 elements, non-long terminal repeat retrotransposons of the RTE-1 lineage from the human blood fluke Schistosoma mansoni. Mol. Biol. Evol. 16, 1256–1269 (1999)
DeMarco, R., Machado, A. A., Bisson-Filho, A. W. & Verjovski-Almeida, S. Identification of 18 new transcribed retrotransposons in Schistosoma mansoni. Biochem. Biophys. Res. Commun. 333, 230–240 (2005)
Volfovsky, N., Haas, B. J. & Salzberg, S. L. Computational discovery of internal micro-exons. Genome Res. 13, 1216–1221 (2003)
Ghedin, E. et al. Draft genome of the filarial nematode parasite Brugia malayi. Science 317, 1756–1760 (2007)
Abad, P. et al. Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nature Biotechnol. 26, 909–915 (2008)
Opperman, C. H. et al. Sequence and genetic map of Meloidogyne hapla: a compact nematode genome for plant parasitism. Proc. Natl Acad. Sci. USA 105, 14802–14807 (2008)
Dorsey, C. H., Cousin, C. E., Lewis, F. A. & Stirewalt, M. A. Ultrastructure of the Schistosoma mansoni cercaria. Micron 33, 279–323 (2002)
Wu, W., Niles, E. G., Hirai, H. & LoVerde, P. T. Evolution of a novel subfamily of nuclear receptors with members that each contain two DNA binding domains. BMC Evol. Biol. 7, 27 (2007)
Sayed, A. A. et al. Identification of oxadiazoles as new drug leads for the control of schistosomiasis. Nature Med. 14, 407–412 (2008)
Brouwers, J. F., Smeenk, I. M., van Golde, L. M. & Tielens, A. G. The incorporation, modification and turnover of fatty acids in adult Schistosoma mansoni. Mol. Biochem. Parasitol. 88, 175–185 (1997)
Barrett, J. Biochemistry of Parasitic Helminths (Macmillan Publishers, 1981)
de Kroon, A. I. Metabolism of phosphatidylcholine and its implications for lipid acyl chain composition in Saccharomyces cerevisiae. Biochim. Biophys. Acta 1771, 343–352 (2007)
Overington, J. P., Al-Lazikani, B. & Hopkins, A. L. How many drug targets are there? Nature Rev. Drug Discov. 5, 993–996 (2006)
Hamdan, F. F. et al. A novel Schistosoma mansoni G protein-coupled receptor is responsive to histamine. Mol. Biochem. Parasitol. 119, 75–86 (2002)
Agboh, K. C., Webb, T. E., Evans, R. J. & Ennion, S. J. Functional characterization of a P2X receptor from Schistosoma mansoni. J. Biol. Chem. 279, 41650–41657 (2004)
Kaczorowski, G. J., McManus, O. B., Priest, B. T. & Garcia, M. L. Ion channels as drug targets: the next GPCRs. J. Gen. Physiol. 131, 399–405 (2008)
Kim, E., Day, T. A., Bennett, J. L. & Pax, R. A. Cloning and functional expression of a Shaker-related voltage-gated potassium channel gene from Schistosoma mansoni (Trematoda: Digenea). Parasitology 110, 171–180 (1995)
Salkoff, L. et al. Potassium channels in C. elegans. WormBook Dec 30, 1–15 (2005)
Jeziorski, M. C. & Greenberg, R. M. Voltage-gated calcium channel subunits from platyhelminths: potential role in praziquantel action. Int. J. Parasitol. 36, 625–632 (2006)
Boyle, S. N. & Koleske, A. J. Dissecting kinase signaling pathways. Drug Discov. Today 12, 717–724 (2007)
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002)
Lopez-Otin, C. & Overall, C. M. Protease degradomics: a new challenge for proteomics. Nature Rev. Mol. Cell Biol. 3, 509–519 (2002)
Rawlings, N. D. & Morton, F. R. The MEROPS batch BLAST: a tool to detect peptidases and their non-peptidase homologues in a genome. Biochimie 90, 243–259 (2008)
Abbenante, G. & Fairlie, D. P. Protease inhibitors in the clinic. Med. Chem. 1, 71–104 (2005)
Fear, G., Komarnytsky, S. & Raskin, I. Protease inhibitors and their peptidomimetic derivatives as potential drugs. Pharmacol. Ther. 113, 354–368 (2007)
Page, M. J. & Di Cera, E. Serine peptidases: classification, structure and function. Cell. Mol. Life Sci. 65, 1220–1236 (2008)
Krem, M. M. & Di Cera, E. Evolution of enzyme cascades from embryonic development to blood coagulation. Trends Biochem. Sci. 27, 67–74 (2002)
Zou, Z., Lopez, D. L., Kanost, M. R., Evans, J. D. & Jiang, H. Comparative analysis of serine protease-related genes in the honey bee genome: possible involvement in embryonic development and innate immunity. Insect Mol. Biol. 15, 603–614 (2006)
Ieko, M. et al. Factor Xa inhibitors: new anti-thrombotic agents and their characteristics. Front. Biosci. 11, 232–248 (2006)
Okun, I., Balakin, K. V., Tkachenko, S. E. & Ivachtchenko, A. V. Caspase activity modulators as anticancer agents. Anticancer Agents Med. Chem. 8, 322–341 (2008)
Caffrey, C. R. & Steverding, D. Recent initiatives and strategies to developing new drugs for tropical parasitic diseases. Expert Opin. Drug Discov. 3, 173–186 (2008)
Abdulla, M. H., Lim, K. C., Sajid, M., McKerrow, J. H. & Caffrey, C. R. Schistosomiasis mansoni: novel chemotherapy using a cysteine protease inhibitor. PLoS Med. 4, e14 (2007)
Cao, M., Chao, H. & Doughty, B. L. A cDNA from Schistosoma mansoni eggs sharing sequence features of mammalian cystatin. Mol. Biochem. Parasitol. 57, 175–176 (1993)
Morales, F. C., Furtado, D. R. & Rumjanek, F. D. The N-terminus moiety of the cystatin SmCys from Schistosoma mansoni regulates its inhibitory activity in vitro and in vivo. Mol. Biochem. Parasitol. 134, 65–73 (2004)
Humphries, J. E. et al. Structure and bioactivity of neuropeptide F from the human parasites Schistosoma mansoni and Schistosoma japonicum. J. Biol. Chem. 279, 39880–39885 (2004)
Ashburn, T. T. & Thor, K. B. Drug repositioning: identifying and developing new uses for existing drugs. Nature Rev. Drug Discov. 3, 673–683 (2004)
Nash, T. & Rice, W. G. Efficacies of zinc-finger-active drugs against Giardia lamblia. Antimicrob. Agents Chemother. 42, 1488–1492 (1998)
Sambon, L. W. New or little known African Entozoa. J. Trop. Med. Hyg. 10, 117 (1907)
Fletcher, M., LoVerde, P. T. & Woodruff, D. S. Genetic variation in Schistosoma mansoni: enzyme polymorphisms in populations from Africa, Southwest Asia, South America, and the West Indies. Am. J. Trop. Med. Hyg. 30, 406–421 (1981)
Berriman, M. & Harris, M. Annotation of parasite genomes. Methods Mol. Biol. 270, 17–44 (2004)
Mullikin, J. C. & Ning, Z. The Phusion assembler. Genome Res. 13, 81–90 (2003)
Simpson, A. J., Sher, A. & McCutchan, T. F. The genome of Schistosoma mansoni: isolation of DNA, its size, bases and repetitive sequences. Mol. Biochem. Parasitol. 6, 125–137 (1982)
Hirai, H. & Hirai, Y. FISH mapping for helminth genome. Methods Mol. Biol. 270, 379–394 (2004)
Le Paslier, M. C. et al. Construction and characterization of a Schistosoma mansoni bacterial artificial chromosome library. Genomics 65, 87–94 (2000)
Biedler, J. & Tu, Z. Non-LTR retrotransposons in the African malaria mosquito, Anopheles gambiae: unprecedented diversity and evidence of recent activity. Mol. Biol. Evol. 20, 1811–1825 (2003)
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990)
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7 (2008)
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004)
Korf, I., Flicek, P., Duan, D. & Brent, M. R. Integrating genomic homology into gene structure prediction. Bioinformatics 17 (suppl. 1). S140–S148 (2001)
Stanke, M., Schoffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinformatics 7, 62 (2006)
Huang, X., Adams, M. D., Zhou, H. & Kerlavage, A. R. A tool for analyzing and annotating genomic sequences. Genomics 46, 37–45 (1997)
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004)
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005)
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21 (suppl. 1). i351–i358 (2005)
Wingender, E., Dietze, P., Karas, H. & Knuppel, R. TRANSFAC: a database on transcription factors and their DNA binding sites. Nucleic Acids Res. 24, 238–241 (1996)
Kummerfeld, S. K. & Teichmann, S. A. DBD: a transcription factor prediction database. Nucleic Acids Res. 34, D74–D81 (2006)
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002)
Coulson, R. M. & Ouzounis, C. A. The phylogenetic diversity of eukaryotic transcription. Nucleic Acids Res. 31, 653–660 (2003)
Verjovski-Almeida, S. et al. Transcriptome analysis of the acoelomate human parasite Schistosoma mansoni. Nature Genet. 35, 148–157 (2003)
Emanuelsson, O., Brunak, S., von Heijne, G. & Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nature Protocols 2, 953–971 (2007)
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001)
Li, H. et al. TreeFam: a curated database of phylogenetic trees of animal gene families. Nucleic Acids Res. 34, D572–D580 (2006)
Ruan, J. et al. TreeFam: 2008 update. Nucleic Acids Res. 36, D735–D740 (2008)
Eddy, S. R. Profile hidden Markov models. Bioinformatics 14, 755–763 (1998)
Putnam, N. H. et al. Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317, 86–94 (2007)
Katoh, K. & Toh, H. Improved accuracy of multiple ncRNA alignment by incorporating structural information into a MAFFT-based framework. BMC Bioinformatics 9, 212 (2008)
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997)
Anamika, K., Martin, J. & Srinivasan, N. Comparative kinomics of human and chimpanzee reveal unique kinship and functional diversity generated by new domain combinations. BMC Genomics 9, 625 (2008)
El-Sayed, N. M. et al. The genome sequence of Trypanosoma cruzi, etiologic agent of Chagas disease. Science 309, 409–415 (2005)
Berriman, M. et al. The genome of the African trypanosome Trypanosoma brucei. Science 309, 416–422 (2005)
Manning, G. Genomic overview of protein kinases. WormBook Dec 13, 1–19 (2005)
Parsons, M., Worthey, E. A., Ward, P. N. & Mottram, J. C. Comparative analysis of the kinomes of three pathogenic trypanosomatids: Leishmania major, Trypanosoma brucei and Trypanosoma cruzi. BMC Genomics 6, 127 (2005)
Caenepeel, S., Charydczak, G., Sudarsanam, S., Hunter, T. & Manning, G. The mouse kinome: discovery and comparative genomics of all mouse protein kinases. Proc. Natl Acad. Sci. USA 101, 11707–11712 (2004)
Rawlings, N. D., Morton, F. R., Kok, C. Y., Kong, J. & Barrett, A. J. MEROPS: the peptidase database. Nucleic Acids Res. 36, D320–D325 (2008)
Sonnhammer, E. L., Eddy, S. R. & Durbin, R. Pfam: a comprehensive database of protein domain families based on seed alignments. Proteins 28, 405–420 (1997)
Finn, R. D. et al. The Pfam protein families database. Nucleic Acids Res. 36, D281–D288 (2008)
Karp, P. D., Paley, S. & Romero, P. The Pathway Tools software. Bioinformatics 18 (suppl. 1). S225–S232 (2002)
Caspi, R. et al. MetaCyc: a multiorganism database of metabolic pathways and enzymes. Nucleic Acids Res. 34, D511–D516 (2006)
Yeh, I., Hanekamp, T., Tsoka, S., Karp, P. D. & Altman, R. B. Computational analysis of Plasmodium falciparum metabolism: organizing genomic information to facilitate drug discovery. Genome Res. 14, 917–924 (2004)
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001)
Acknowledgements
The genome sequencing and annotation work was funded by the Wellcome Trust (grant number WT085775/Z/08/Z) and the National Institutes of Health (NIH) National Institute of Allergy and Infectious Diseases (NIAID) grant AI48828 to N.M.E.-S. We thank N. D. Rawlings of the MEROPS database team at the Wellcome Trust Sanger Institute for his help, J. C. Illes for discussions on polarity complexes, and F. Prosdocimi and M. R. D. Sananes for early discussions and analyses in the project. FISH chromosome mappings were partially supported by Oyama Health Foundation (H.H.), Japan Society for the Promotion of Science (13557021) (H.H.), 21st century Centers of Exellence and global Centers of Excellence of Japan’s Ministry of Education, Culture, Sports, Science and Technology. Additional support was by The Sandler Foundation (C.R.C. and M.S.), NIH-Fogarty 5D43TW006580 (P.T.L.), NIH-Fogarty 5D43TW007012-03, NIH grant AI054711-01A2 (R.A.W. and G.P.D.), FAPEMIG REDE-281/05 (G.O.), the PhRMA Foundation (Postdoctoral Fellowship in Informatics to S.T.M.), The Burroughs Wellcome Fund (P.T.L.) and the United Nations Children’s Fund (UNICEF)/United Nations Development Program (UNDP)/World bank/World Health Organization (WHO) Special program for research and training in tropical diseases (TDR) (P.T.L.). R.D. was a recipient of CAPES and FAPESP fellowships.
Author Contributions A.I., R.A.W., C.M.F.-L., D.A.J., N.M.E.-S. and P.T.L. initiated the project; M.A.Q. constructed DNA libraries and J.P. and J.R. directed sequencing; Y.G. and Z.N. assembled the genome sequence data; H.H., P.T.L., R.J.P. and Y.H. produced the mapping data; A.De., A.Dj., A.R.T., B.J.H., D.C.B., D.L., G.C.C., J.W., M.A.A., M.-A.R., M.Sa., O.W., P.D.A., R.H., S.L.S. and T.E. provided computational and bioinformatic support; A.R.T., M.A.A. and R.H. set up and maintained the genome database; C.D.M., D.C.B., G.B., G.C.C. and J.A.G. produced the gene finding training set; B.J.H., M.P. and M.St. trained genefinding software; A.V.P., B.G.B. and B.J.H. annotated the genome data; A.C., A.Z., B.A.-L., C.R.C., D.L.W., G.O., G.P.D., J.P.O., L.F.A., M.Sa., M.Z., P.M., R.C., R.D., S.T.M., T.A.D. and W.W. contributed specific analysis topics presented in this manuscript; C.H.-F. and E.G. contributed to general project and sequencing management; B.G.B., C.H.-F., C.R.C. and J.P. commented on the manuscript drafts; G.C.C. performed data submission to GenBank; R.A.W., M.B., N.M.E.-S. and P.T.L. drafted and edited the paper; R.A.W., D.A.J. and P.T.L. provided DNA resources for the sequencing; M.B. and N.M.E.-S. directed the project and assembled the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Notes and accompanying Supplementary References, and Supplementary Figures 1-9. (PDF 2864 kb)
Supplementary Tables
This file contains Supplementary Tables 1-23. (XLS 2725 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Berriman, M., Haas, B., LoVerde, P. et al. The genome of the blood fluke Schistosoma mansoni. Nature 460, 352–358 (2009). https://doi.org/10.1038/nature08160
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08160
This article is cited by
-
An insight into the functional genomics and species classification of Eudiplozoon nipponicum (Monogenea, Diplozoidae), a haematophagous parasite of the common carp Cyprinus carpio
BMC Genomics (2023)
-
Neuronal gene expression in two generations of the marine parasitic worm, Cryptocotyle lingua
Communications Biology (2023)
-
The stage- and sex-specific transcriptome of the human parasite Schistosoma mansoni
Scientific Data (2023)
-
Understanding anthelmintic resistance in livestock using “omics” approaches
Environmental Science and Pollution Research (2023)
-
Metabolomics and lipidomics studies of parasitic helminths: molecular diversity and identification levels achieved by using different characterisation tools
Metabolomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.