Abstract
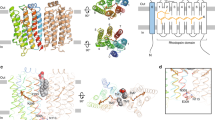
The ability to respond to light is crucial for most organisms. BLUF is a recently identified photoreceptor protein domain that senses blue light using a FAD chromophore1. BLUF domains are present in various proteins from the Bacteria, Euglenozoa and Fungi. Although structures of single-domain BLUF proteins have been determined2,3,4, none are available for a BLUF protein containing a functional output domain; the mechanism of light activation in this new class of photoreceptors has thus remained poorly understood. Here we report the biochemical, structural and mechanistic characterization of a full-length, active photoreceptor, BlrP1 (also known as KPN_01598), from Klebsiella pneumoniae5. BlrP1 consists of a BLUF sensor domain and a phosphodiesterase EAL output domain which hydrolyses cyclic dimeric GMP (c-di-GMP). This ubiquitous second messenger controls motility, biofilm formation, virulence and antibiotic resistance in the Bacteria6,7,8,9. Crystal structures of BlrP1 complexed with its substrate and metal ions involved in catalysis or in enzyme inhibition provide a detailed understanding of the mechanism of the EAL-domain c-di-GMP phosphodiesterases. These structures also sketch out a path of light activation of the phosphodiesterase output activity. Photon absorption by the BLUF domain of one subunit of the antiparallel BlrP1 homodimer activates the EAL domain of the second subunit through allosteric communication transmitted through conserved domain–domain interfaces.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The atomic coordinates and structure factor amplitudes of the following structures have been deposited in the Protein Data Bank: Ca2+ complex of BlrP1 at pH 4.5 (accession code 3GFX), metal-free complex of BlrP1 at pH 6.0 (3GFY), Mn2+ complex of BlrP1 at pH 6.0 (3GFZ), Mn2+ complex of BlrP1 at pH 9.0 (3GG0) and Ca2+ complex of BlrP1 at pH 8.0 (3GG1).
References
Gomelsky, M. & Klug, G. BLUF: a novel FAD-binding domain involved in sensory transduction in microorganisms. Trends Biochem. Sci. 27, 497–500 (2002)
Jung, A. et al. Structure of a bacterial BLUF photoreceptor: insights into blue light-mediated signal transduction. Proc. Natl Acad. Sci. USA 102, 12350–12355 (2005)
Kita, A., Okajima, K., Morimoto, Y., Ikeuchi, M. & Miki, K. Structure of a cyanobacterial BLUF protein, Tll0078, containing a novel FAD-binding blue light sensor domain. J. Mol. Biol. 349, 1–9 (2005)
Yuan, H. et al. Crystal structures of the Synechocystis photoreceptor Slr1694 reveal distinct structural states related to signaling. Biochemistry 45, 12687–12694 (2006)
Tyagi, A. et al. Photodynamics of the blue-light-regulated phosphodiesterase BlrP1 from Klebsiella pneumoniae and its photoreceptor BLUF domain. Chem. Phys. 354, 130–141 (2008)
Hisert, K. B. et al. A glutamate-alanine-leucine (EAL) domain protein of Salmonella controls bacterial survival in mice, antioxidant defence and killing of macrophages: role of cyclic diGMP. Mol. Microbiol. 56, 1234–1245 (2005)
Hoffman, L. R. et al. Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 436, 1171–1175 (2005)
Kulasakara, H. et al. Analysis of Pseudomonas aeruginosa diguanylate cyclases and phosphodiesterases reveals a role for bis-(3′-5′)-cyclic-GMP in virulence. Proc. Natl Acad. Sci. USA 103, 2839–2844 (2006)
Claret, L. et al. The flagellar sigma factor FliA regulates adhesion and invasion of Crohn disease-associated Escherichia coli via a cyclic dimeric GMP-dependent pathway. J. Biol. Chem. 282, 33275–33283 (2007)
Minasov, B. et al. Crystal structures of Ykul and its complex with second messenger c-di-GMP suggests catalytic mechanism of phosphodiester bond cleavage by EAL domains. J. Biol. Chem. 10.1074/jbc.M808221200 (in the press)
Chang, C. et al. Crystal structure of a putative diguanylate cyclase/phosphodiesterase from Thiobacillus denitrificans . RCSB PDB: Structure Explorer 〈http://www.rcsb.org/pdb/explore/explore.do?structureId=2R6O〉 (2007)
Gomelsky, M. & Kaplan, S. AppA, a redox regulator of photosystem formation in Rhodobacter sphaeroides 2.4.1, is a flavoprotein. Identification of a novel FAD binding domain. J. Biol. Chem. 273, 35319–35325 (1998)
Wu, Q. & Gardner, K. H. Structure and insight into blue light-induced changes in the BlrP1 BLUF domain. Biochemistry 48, 2620–2629 (2009)
Benach, J. et al. The structural basis of cyclic diguanylate signal transduction by PilZ domains. EMBO J. 26, 5153–5166 (2007)
Zhan, C. G. & Zheng, F. First computational evidence for a catalytic bridging hydroxide ion in a phosphodiesterase active site. J. Am. Chem. Soc. 123, 2835–2838 (2001)
Salter, E. A. & Wierzbicki, A. The mechanism of cyclic nucleotide hydrolysis in the phosphodiesterase catalytic site. J. Phys. Chem. B 111, 4547–4552 (2007)
Schmidt, A. J., Ryjenkov, D. A. & Gomelsky, M. The ubiquitous protein domain EAL is a cyclic diguanylate-specific phosphodiesterase: enzymatically active and inactive EAL domains. J. Bacteriol. 187, 4774–4781 (2005)
Rao, F., Yang, Y., Qi, Y. & Liang, Z.-X. Catalytic mechanism of c-di-GMP specific phosphodiesterase: a study of the EAL domain-containing RocR from Pseudomonas aeruginosa . J. Bacteriol. 190, 3622–3631 (2008)
Ross, P. et al. The cyclic diguanylic acid regulatory system of cellulose synthesis in Acetobacter xylinum. Chemical synthesis and biological activity of cyclic nucleotide dimer, trimer, and phosphothioate derivatives. J. Biol. Chem. 265, 18933–18943 (1990)
Herschlag, D. & Jencks, W. P. Catalysis of the hydrolysis of phosphorylated pyridines by Mg(OH)+: a possible model for enzymatic phosphoryl transfer. Biochemistry 29, 5172–5179 (1990)
Jung, A., Reinstein, J., Domratcheva, T., Shoeman, R. L. & Schlichting, I. Crystal structures of the AppA BLUF domain photoreceptor provide insights into blue light-mediated signal transduction. J. Mol. Biol. 362, 717–732 (2006)
Anderson, S. et al. Structure of a novel photoreceptor, the BLUF domain of AppA from Rhodobacter sphaeroides . Biochemistry 44, 7998–8005 (2005)
Wu, Q., Ko, W. H. & Gardner, K. H. Structural requirements for key residues and auxiliary portions of a BLUF domain. Biochemistry 47, 10271–10280 (2008)
Domratcheva, T., Grigorenko, B. L., Schlichting, I. & Nemukhin, A. V. Molecular models predict light-induced glutamine tautomerization in BLUF photoreceptors. Biophys. J. 94, 3872–3879 (2008)
Gauden, M. et al. Hydrogen-bond switching through a radical pair mechanism in a flavin-binding photoreceptor. Proc. Natl Acad. Sci. USA 103, 10895–10900 (2006)
Sadeghian, K., Bocola, M. & Schütz, M. A conclusive mechanism of the photoinduced reaction cascade in blue light using flavin photoreceptors. J. Am. Chem. Soc. 130, 12501–12513 (2008)
Finn, R. D. et al. Pfam: clans, web tools and services. Nucleic Acids Res. 34, D247–D251 (2006)
Hur, S. & Bruice, T. C. Enzymes do what is expected (chalcone isomerase versus chorismate mutase). J. Am. Chem. Soc. 125, 1472–1473 (2003)
Schuck, P. & Rossmanith, P. Determination of the sedimentation coefficient distribution by least-squares boundary modeling. Biopolymers 54, 328–341 (2000)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–806 (1993)
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007)
Terwilliger, T. C. Automated structure solution, density modification and model building. Acta Crystallogr. D 58, 1937–1940 (2002)
Terwilliger, T. C. SOLVE and RESOLVE: automated structure solution and density modification. Methods Enzymol. 374, 22–37 (2003)
Terwilliger, T. SOLVE and RESOLVE: automated structure solution, density modification and model building. J. Synchrotron Radiat. 11, 49–52 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
McRee, D. E. XtalView/Xfit–A versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Brunger, A. T. Version 1.2 of the Crystallography and NMR system. Nature Protocols 2, 2728–2733 (2007)
DeLano, W. L. PyMOL Molecular Viewer 〈http://www.pymol.org〉 (2002)
Acknowledgements
We thank A. Jung and A. Schmidt for help during the initial stages of the project, C. Fang for c-di-GMP purification and Q. Wu and K. H. Gardner for discussions and for sharing unpublished data. We are grateful to I. Vetter for advice and support for the crystallographic software. Diffraction data were collected at the Swiss Light Source, beamline X10SA, Paul Scherrer Institute, Villigen, Switzerland. We thank the Dortmund–Heidelberg team for data collection and E. Pohl, A. Pauluhn and C. Schulze-Briese for their support in setting up the beamline. The K. pneumoniae genome was sequenced at the School of Medicine of the Washington University in St Louis. We acknowledge financial support from the Deutsche Forschungsgemeinschaft (Research Group FOR 526 ‘Sensory Blue Light Receptors’) to I.S., the US National Science Foundation (MCB 0645876) to M.G. and the Boehringer Ingelheim Fonds to J.J.G. I.S. and J.R. are members of the Cluster of Excellence CellNetworks, Heidelberg.
Author Contributions M.G. and I.S. designed the project; N.V.K. and M.G. identified and cloned blrP1; N.V.K. and D.A.R. purified BlrP1 and performed the initial biochemical analysis; J.J.G. constructed the expression clone and established the purification protocol of the protein used for crystallization and biochemical analysis; E.H. purified, crystallized and characterized wild-type and mutant BlrP1 by enzymatic and dynamic light scattering analysis; T.B. performed the spectroscopic analysis; T.R.M.B. determined the crystal structures; J.R. designed the kinetic experiments; R.L.S. performed the mass spectrometry and analytical ultracentrifugation; and I.S. did the practical crystallographic work. T.R.M.B., M.G. and I.S. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S8 with Legends, Supplementary Tables S1-S3 and Supplementary References. (PDF 3755 kb)
Rights and permissions
About this article
Cite this article
Barends, T., Hartmann, E., Griese, J. et al. Structure and mechanism of a bacterial light-regulated cyclic nucleotide phosphodiesterase. Nature 459, 1015–1018 (2009). https://doi.org/10.1038/nature07966
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature07966
This article is cited by
-
CdbA is a DNA-binding protein and c-di-GMP receptor important for nucleoid organization and segregation in Myxococcus xanthus
Nature Communications (2020)
-
A Cyclic-di-GMP signalling network regulates biofilm formation and surface associated motility of Acinetobacter baumannii 17978
Scientific Reports (2020)
-
Structure and dynamics of photoactivatable adenylyl cyclase
Russian Chemical Bulletin (2019)
-
Regulating exopolysaccharide gene wcaF allows control of Escherichia coli biofilm formation
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.