Abstract
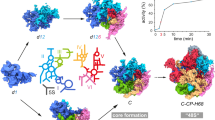
Rapidly growing cells produce thousands of new ribosomes each minute, in a tightly regulated process that is essential to cell growth1,2. How the Escherichia coli 16S ribosomal RNA and the 20 proteins that make up the 30S ribosomal subunit can assemble correctly in a few minutes remains a challenging problem, partly because of the lack of real-time data on the earliest stages of assembly. By providing snapshots of individual RNA and protein interactions as they emerge in real time, here we show that 30S assembly nucleates concurrently from different points along the rRNA. Time-resolved hydroxyl radical footprinting3 was used to map changes in the structure of the rRNA within 20 milliseconds after the addition of total 30S proteins. Helical junctions in each domain fold within 100 ms. In contrast, interactions surrounding the decoding site and between the 5′, the central and the 3′ domains require 2–200 seconds to form. Unexpectedly, nucleotides contacted by the same protein are protected at different rates, indicating that initial RNA–protein encounter complexes refold during assembly. Although early steps in assembly are linked to intrinsically stable rRNA structure, later steps correspond to regions of induced fit between the proteins and the rRNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Warner, J. R., Vilardell, J. & Sohn, J. H. Economics of ribosome biosynthesis. Cold Spring Harb. Symp. Quant. Biol. 66, 567–574 (2001)
Nierhaus, K. H. The assembly of prokaryotic ribosomes. Biochimie 73, 739–755 (1991)
Ralston, C. Y. et al. Time-resolved synchrotron X-ray footprinting and its application to RNA folding. Methods Enzymol. 317, 353–368 (2000)
Held, W. A., Mizushima, S. & Nomura, M. Reconstitution of Escherichia coli 30 S ribosomal subunits from purified molecular components. J. Biol. Chem. 248, 5720–5730 (1973)
Stern, S., Powers, T., Changchien, L. M. & Noller, H. F. RNA–protein interactions in 30S ribosomal subunits: folding and function of 16S rRNA. Science 244, 783–790 (1989)
Culver, G. M. Assembly of the 30S ribosomal subunit. Biopolymers 68, 234–249 (2003)
Williamson, J. R. After the ribosome structures: how are the subunits assembled? RNA 9, 165–167 (2003)
Noller, H. F. & Nomura, M. in Escherichia Coli and Salmonella Typhimurium, Cellular and Molecular Biology (ed. Neidhardt, F. C.) 104–125 (American Society for Microbiology, 1987)
Tullius, T. D. & Greenbaum, J. A. Mapping nucleic acid structure by hydroxyl radical cleavage. Curr. Opin. Chem. Biol. 9, 127–134 (2005)
Powers, T., Daubresse, G. & Noller, H. F. Dynamics of in vitro assembly of 16 S rRNA into 30 S ribosomal subunits. J. Mol. Biol. 232, 362–374 (1993)
Holmes, K. L. & Culver, G. M. Mapping structural differences between 30S ribosomal subunit assembly intermediates. Nature Struct. Mol. Biol. 11, 179–186 (2004)
Adilakshmi, T., Ramaswamy, P. & Woodson, S. A. Protein-independent folding pathway of the 16S rRNA 5′ domain. J. Mol. Biol. 351, 508–519 (2005)
Ramakrishnan, V. Distribution of protein and RNA in the 30S ribosomal subunit. Science 231, 1562–1564 (1986)
Wimberly, B. T. et al. Structure of the 30S ribosomal subunit. Nature 407, 327–339 (2000)
Schuwirth, B. S. et al. Structures of the bacterial ribosome at 3.5 Å resolution. Science 310, 827–834 (2005)
Traub, P. & Nomura, M. Structure and function of Escherichia coli ribosomes. VI. Mechanism of assembly of 30 s ribosomes studied in vitro . J. Mol. Biol. 40, 391–413 (1969)
Talkington, M. W., Siuzdak, G. & Williamson, J. R. An assembly landscape for the 30S ribosomal subunit. Nature 438, 628–632 (2005)
Pan, J., Thirumalai, D. & Woodson, S. A. Folding of RNA involves parallel pathways. J. Mol. Biol. 273, 7–13 (1997)
Laederach, A., Shcherbakova, I., Liang, M. P., Brenowitz, M. & Altman, R. B. Local kinetic measures of macromolecular structure reveal partitioning among multiple parallel pathways from the earliest steps in the folding of a large RNA molecule. J. Mol. Biol. 358, 1179–1190 (2006)
Cate, J. H., Yusupov, M. M., Yusupova, G. Z., Earnest, T. N. & Noller, H. F. X-ray crystal structures of 70S ribosome functional complexes. Science 285, 2095–2104 (1999)
Brodersen, D. E., Clemons, W. M., Carter, A. P., Wimberly, B. T. & Ramakrishnan, V. Crystal structure of the 30 S ribosomal subunit from Thermus thermophilus: structure of the proteins and their interactions with 16 S RNA. J. Mol. Biol. 316, 725–768 (2002)
Dutca, L. M. & Culver, G. M. Assembly of the 5′ and 3′ minor domains of 16S ribosomal RNA as monitored by tethered probing from ribosomal protein S20. J. Mol. Biol. 376, 92–108 (2008)
Powers, T. & Noller, H. F. Hydroxyl radical footprinting of ribosomal proteins on 16S rRNA. RNA 1, 194–209 (1995)
Nowotny, V. & Nierhaus, K. H. Assembly of the 30S subunit from Escherichia coli ribosomes occurs via two assembly domains which are initiated by S4 and S7. Biochemistry 27, 7051–7055 (1988)
Powers, T. & Noller, H. F. A temperature-dependent conformational rearrangement in the ribosomal protein S4·16 S rRNA complex. J. Biol. Chem. 270, 1238–1242 (1995)
Sayers, E. W., Gerstner, R. B., Draper, D. E. & Torchia, D. A. Structural preordering in the N-terminal region of ribosomal protein S4 revealed by heteronuclear NMR spectroscopy. Biochemistry 39, 13602–13613 (2000)
Samaha, R. R., O’Brien, B., O’Brien, T. W. & Noller, H. F. Independent in vitro assembly of a ribonucleoprotein particle containing the 3′ domain of 16S rRNA. Proc. Natl Acad. Sci. USA 91, 7884–7888 (1994)
Dodd, J., Kolb, J. M. & Nomura, M. Lack of complete cooperativity of ribosome assembly in vitro and its possible relevance to in vivo ribosome assembly and the regulation of ribosomal gene expression. Biochimie 73, 757–767 (1991)
Acknowledgements
We thank R. Moss, A. Cukras, L. Cochella and R. Green for help with ribosome preparation and peptidyl transferase assays, P. Fleming for help with Calc-Surf software, and S. Gupta, M. Sullivan and M. Brenowitz for help with X-ray footprinting. This work was supported by the National Institutes of Health (NIH; GM60819). The NSLS X28C and the Center for Synchrotron Biosciences are supported by NIH P41-EB0001979.
Author Contributions T.A. performed the experiments, analysed the data and prepared the figures; D.L.B. analysed protections in the 3′ minor domain; and S.A.W. prepared the figures and wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Figures S1-S6 and Supplementary Tables S1-S2 and References (PDF 1427 kb)
Rights and permissions
About this article
Cite this article
Adilakshmi, T., Bellur, D. & Woodson, S. Concurrent nucleation of 16S folding and induced fit in 30S ribosome assembly. Nature 455, 1268–1272 (2008). https://doi.org/10.1038/nature07298
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07298
This article is cited by
-
Thermodynamic and Kinetic Analyses of Iron Response Element (IRE)-mRNA Binding to Iron Regulatory Protein, IRP1
Scientific Reports (2017)
-
Evolution of protein-coupled RNA dynamics during hierarchical assembly of ribosomal complexes
Nature Communications (2017)
-
Functional characterization of chloroplast-targeted RbgA GTPase in higher plants
Plant Molecular Biology (2017)
-
Sequential domain assembly of ribosomal protein S3 drives 40S subunit maturation
Nature Communications (2016)
-
Chemical modulators of ribosome biogenesis as biological probes
Nature Chemical Biology (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.