Abstract
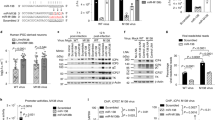
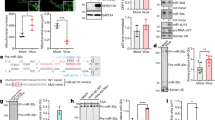
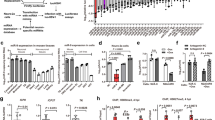
All metazoan eukaryotes express microRNAs (miRNAs), roughly 22-nucleotide regulatory RNAs that can repress the expression of messenger RNAs bearing complementary sequences1. Several DNA viruses also express miRNAs in infected cells, suggesting a role in viral replication and pathogenesis2. Although specific viral miRNAs have been shown to autoregulate viral mRNAs3,4 or downregulate cellular mRNAs5,6, the function of most viral miRNAs remains unknown. Here we report that the miR-K12-11 miRNA encoded by Kaposi’s-sarcoma-associated herpes virus (KSHV) shows significant homology to cellular miR-155, including the entire miRNA ‘seed’ region7. Using a range of assays, we show that expression of physiological levels of miR-K12-11 or miR-155 results in the downregulation of an extensive set of common mRNA targets, including genes with known roles in cell growth regulation. Our findings indicate that viral miR-K12-11 functions as an orthologue of cellular miR-155 and probably evolved to exploit a pre-existing gene regulatory pathway in B cells. Moreover, the known aetiological role of miR-155 in B-cell transformation8,9,10 suggests that miR-K12-11 may contribute to the induction of KSHV-positive B-cell tumours in infected patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004)
Cullen, B. R. Viruses and microRNAs. Nature Genet. 38 (Suppl.). S25–S30 (2006)
Pfeffer, S. et al. Identification of virus-encoded microRNAs. Science 304, 734–736 (2004)
Sullivan, C. S., Grundhoff, A. T., Tevethia, S., Pipas, J. M. & Ganem, D. SV40-encoded microRNAs regulate viral gene expression and reduce susceptibility to cytotoxic T cells. Nature 435, 682–686 (2005)
Stern-Ginossar, N. et al. Host immune system gene targeting by a viral miRNA. Science 317, 376–381 (2007)
Samols, M. A. et al. Identification of cellular genes targeted by KSHV-encoded microRNAs. PLoS Pathog. 3, e65 (2007)
Lewis, B. P., Burge, C. B. & Bartel, D. P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005)
Clurman, B. E. & Hayward, W. S. Multiple proto-oncogene activations in avian leukosis virus-induced lymphomas: evidence for stage-specific events. Mol. Cell. Biol. 9, 2657–2664 (1989)
Costinean, S. et al. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in Eμ-miR155 transgenic mice. Proc. Natl Acad. Sci. USA 103, 7024–7029 (2006)
Eis, P. S. et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc. Natl Acad. Sci. USA 102, 3627–3632 (2005)
van den Berg, A. et al. High expression of B-cell receptor inducible gene BIC in all subtypes of Hodgkin lymphoma. Genes Chromosom. Cancer 37, 20–28 (2003)
Haasch, D. et al. T cell activation induces a noncoding RNA transcript sensitive to inhibition by immunosuppressant drugs and encoded by the proto-oncogene, BIC . Cell. Immunol. 217, 78–86 (2002)
O’Connell, R. M., Taganov, K. D., Boldin, M. P., Cheng, G. & Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl Acad. Sci. USA 104, 1604–1609 (2007)
Thai, T. H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608 (2007)
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007)
Gottwein, E. & Cullen, B. R. Protocols for expression and functional analysis of viral microRNAs. Methods Enzymol. 427, 229–243 (2007)
Gottwein, E., Cai, X. & Cullen, B. R. A novel assay for viral microRNA function identifies a single nucleotide polymorphism that affects Drosha processing. J. Virol. 80, 5321–5326 (2006)
Cai, X. et al. Kaposi’s sarcoma-associated herpesvirus expresses an array of viral microRNAs in latently infected cells. Proc. Natl Acad. Sci. USA 102, 5570–5575 (2005)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Xie, X. et al. Systematic discovery of regulatory motifs in conserved regions of the human genome, including thousands of CTCF insulator sites. Proc. Natl Acad. Sci. USA 104, 7145–7150 (2007)
Olsen, P. H. & Ambros, V. The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev. Biol. 216, 671–680 (1999)
Zeng, Y., Wagner, E. J. & Cullen, B. R. Both natural and designed micro RNAs can inhibit the expression of cognate mRNAs when expressed in human cells. Mol. Cell 9, 1327–1333 (2002)
Krutzfeldt, J. et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438, 685–689 (2005)
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007)
Cesarman, E. & Knowles, D. M. Kaposi’s sarcoma-associated herpesvirus: a lymphotropic human herpesvirus associated with Kaposi’s sarcoma, primary effusion lymphoma, and multicentric Castleman’s disease. Semin. Diagn. Pathol. 14, 54–66 (1997)
Jiang, J., Lee, E. J. & Schmittgen, T. D. Increased expression of microRNA-155 in Epstein–Barr virus transformed lymphoblastoid cell lines. Genes Chromosom. Cancer 45, 103–106 (2006)
Silva, J. M. et al. Second-generation shRNA libraries covering the mouse and human genomes. Nature Genet. 37, 1281–1288 (2005)
Buness, A., Huber, W., Steiner, K., Sultmann, H. & Poustka, A. arrayMagic: two-colour cDNA microarray quality control and preprocessing. Bioinformatics 21, 554–556 (2005)
Reich, M. et al. GenePattern 2.0. Nature Genet. 38, 500–501 (2006)
Majoros, W. H. & Ohler, U. Spatial preferences of microRNA targets in 3′ untranslated regions. BMC Genomics 8, 152 (2007)
Lee, M. T., Coburn, G. A., McClure, M. O. & Cullen, B. R. Inhibition of human immunodeficiency virus type 1 replication in primary macrophages by using Tat- or CCR5-specific small interfering RNAs expressed from a lentivirus vector. J. Virol. 77, 11964–11972 (2003)
Rubinson, D. A. et al. A lentivirus-based system to functionally silence genes in primary mammalian cells, stem cells and transgenic mice by RNA interference. Nature Genet. 33, 401–406 (2003)
Acknowledgements
We thank T. Curran for rabbit anti-Fos antiserum. Flow cytometry was performed by J. Whitesides in the Duke Center for AIDS Research BSL3 Flow Cytometry Core Facility. Microarrays were performed in the Duke Microarray Facility. This research was supported by a National Institutes of Health grant to B.R.C. and by a Feodor Lynen Fellowship to E.G. U.O. is an Alfred P. Sloan Fellow in Computational Molecular Biology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
J. Soutschek, R. Braich and M. Manoharan are employees of Alnylam Pharmaceuticals, Inc., which develops therapeutics based on siRNAs. C. Sachse and C. Frenzel are employees of Cenix BioScience GmbH, which is a specialist in RNAi-based discovery and functional characterization of therapeutic targets and drug candidates. The remaining authors declare no competing interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-7 and Legends, Supplementary Tables 1-3 and Legends and Supplementary Tables 4-5 of primers. The Supplementary Figures show control data and additional experimental data in support of the manuscript’s conclusions. (PDF 2298 kb)
Rights and permissions
About this article
Cite this article
Gottwein, E., Mukherjee, N., Sachse, C. et al. A viral microRNA functions as an orthologue of cellular miR-155. Nature 450, 1096–1099 (2007). https://doi.org/10.1038/nature05992
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature05992
This article is cited by
-
A structured RNA motif locks Argonaute2:miR-122 onto the 5’ end of the HCV genome
Nature Communications (2021)
-
Unveiling ancestral relations, host–pathogen interactions and comparative viral miRNA insights of dengue virus serotypes
Network Modeling Analysis in Health Informatics and Bioinformatics (2021)
-
Epigenetic control in Kaposi sarcoma-associated herpesvirus infection and associated disease
Seminars in Immunopathology (2020)
-
miR155 regulation of behavior, neuropathology, and cortical transcriptomics in Alzheimer's disease
Acta Neuropathologica (2020)
-
Towards Better Understanding of KSHV Life Cycle: from Transcription and Posttranscriptional Regulations to Pathogenesis
Virologica Sinica (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.