Abstract
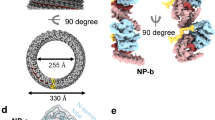
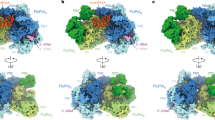
Influenza A viruses pose a serious threat to world public health, particularly the currently circulating avian H5N1 viruses. The influenza viral nucleoprotein forms the protein scaffold of the helical genomic ribonucleoprotein complexes, and has a critical role in viral RNA replication1. Here we report a 3.2 Å crystal structure of this nucleoprotein, the overall shape of which resembles a crescent with a head and a body domain, with a protein fold different compared with that of the rhabdovirus nucleoprotein2,3. Oligomerization of the influenza virus nucleoprotein is mediated by a flexible tail loop that is inserted inside a neighbouring molecule. This flexibility in the tail loop enables the nucleoprotein to form loose polymers as well as rigid helices, both of which are important for nucleoprotein functions. Single residue mutations in the tail loop result in the complete loss of nucleoprotein oligomerization. An RNA-binding groove, which is found between the head and body domains at the exterior of the nucleoprotein oligomer, is lined with highly conserved basic residues widely distributed in the primary sequence. The nucleoprotein structure shows that only one of two proposed nuclear localization signals are accessible, and suggests that the body domain of nucleoprotein contains the binding site for the viral polymerase. Our results identify the tail loop binding pocket as a potential target for antiviral development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lamb, R. A. & Krug, R. M. in Fields Virology (eds Knipe, D. M. & Howley, P. M.) 1487–1531 (Lippincott-Raven, Philadelphia, 2001)
Albertini, A. A. et al. Crystal structure of the rabies virus nucleoprotein-RNA complex. Science 313, 360–363 (2006)
Green, T. J., Zhang, X., Wertz, G. W. & Luo, M. Structure of the vesicular stomatitis virus nucleoprotein-RNA complex. Science 313, 357–360 (2006)
Elton, D., Medcalf, L., Bishop, K., Harrison, D. & Digard, P. Identification of amino acid residues of influenza virus nucleoprotein essential for RNA binding. J. Virol. 73, 7357–7367 (1999)
Albo, C., Valencia, A. & Portela, A. Identification of an RNA binding region within the N-terminal third of the influenza A virus nucleoprotein. J. Virol. 69, 3799–3806 (1995)
Kobayashi, M., Toyoda, T., Adyshev, D. M., Azuma, Y. & Ishihama, A. Molecular dissection of influenza virus nucleoprotein: deletion mapping of the RNA binding domain. J. Virol. 68, 8433–8436 (1994)
Ortega, J. et al. Ultrastructural and functional analyses of recombinant influenza virus ribonucleoproteins suggest dimerization of nucleoprotein during virus amplification. J. Virol. 74, 156–163 (2000)
Compans, R. W., Content, J. & Duesberg, P. H. Structure of the ribonucleoprotein of influenza virus. J. Virol. 10, 795–800 (1972)
Area, E. et al. 3D structure of the influenza virus polymerase complex: localization of subunit domains. Proc. Natl Acad. Sci. USA 101, 308–313 (2004)
Baudin, F., Bach, C., Cusack, S. & Ruigrok, R. W. Structure of influenza virus RNP. I. Influenza virus nucleoprotein melts secondary structure in panhandle RNA and exposes the bases to the solvent. EMBO J. 13, 3158–3165 (1994)
Rudolph, M. G. et al. Crystal structure of the borna disease virus nucleoprotein. Structure 11, 1219–1226 (2003)
Iseni, F., Barge, A., Baudin, F., Blondel, D. & Ruigrok, R. W. Characterization of rabies virus nucleocapsids and recombinant nucleocapsid-like structures. J. Gen. Virol. 79, 2909–2919 (1998)
Wang, P., Palese, P. & O’Neill, R. E. The NPI-1/NPI-3 (karyopherin α) binding site on the influenza A virus nucleoprotein NP is a nonconventional nuclear localization signal. J. Virol. 71, 1850–1856 (1997)
Neumann, G., Castrucci, M. R. & Kawaoka, Y. Nuclear import and export of influenza virus nucleoprotein. J. Virol. 71, 9690–9700 (1997)
Cros, J. F., Garcia-Sastre, A. & Palese, P. An unconventional NLS is critical for the nuclear import of the influenza A virus nucleoprotein and ribonucleoprotein. Traffic 6, 205–213 (2005)
Weber, F., Kochs, G., Gruber, S. & Haller, O. A classical bipartite nuclear localization signal on Thogoto and influenza A virus nucleoproteins. Virology 250, 9–18 (1998)
Fontes, M. R., Teh, T. & Kobe, B. Structural basis of recognition of monopartite and bipartite nuclear localization sequences by mammalian importin-α. J. Mol. Biol. 297, 1183–1194 (2000)
Digard, P. et al. Modulation of nuclear localization of the influenza virus nucleoprotein through interaction with actin filaments. J. Virol. 73, 2222–2231 (1999)
Medcalf, L., Poole, E., Elton, D. & Digard, P. Temperature-sensitive lesions in two influenza A viruses defective for replicative transcription disrupt RNA binding by the nucleoprotein. J. Virol. 73, 7349–7356 (1999)
Honda, A., Ueda, K., Nagata, K. & Ishihama, A. RNA polymerase of influenza virus: role of NP in RNA chain elongation. J. Biochem. 104, 1021–1026 (1988)
Shapiro, G. I. & Krug, R. M. Influenza virus RNA replication in vitro: synthesis of viral template RNAs and virion RNAs in the absence of an added primer. J. Virol. 62, 2285–2290 (1988)
Biswas, S. K., Boutz, P. L. & Nayak, D. P. Influenza virus nucleoprotein interacts with influenza virus polymerase proteins. J. Virol. 72, 5493–5501 (1998)
Gabriel, G. et al. The viral polymerase mediates adaptation of an avian influenza virus to a mammalian host. Proc. Natl Acad. Sci. USA 102, 18590–18595 (2005)
Otwinowski, Z. & Minor, W. Processing of X-ray Diffraction Data in Oscillation Mode (Academic Press, New York, 1997)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003)
Collaborative Computational Project, 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Acknowledgements
We thank V. Vikharia for providing the baculovirus for expressing nucleoprotein from influenza virus A/Hong Kong/1074/99 (H9N2); Z. Xie for electron microscopy analysis; P. Sliz for collecting the nucleoprotein native data; Y. Shamoo and G. Wu for providing the 24-nucleotide ssRNA; and Y. Shamoo, D. Mata and S. Harrison for discussions and critical reading of the manuscript. Nucleoprotein diffraction data were collected at the Brookhaven National Synchrotron Light Source and the Cornell High Energy Synchrotron Source. This work is supported by grants from the Welch Foundation (to Y.J.T and R.M.K), National Institutes of Health (to Y.J.T), and the Nanoscale Science and Engineering Initiative of the National Science Foundation. Author Contributions Q.Y. crystallized the nucleoprotein. Q.Y. and Y.J.T solved the nucleoprotein crystal structure. Y.J.T, Q.Y. and R.M.K. wrote the paper. All authors discussed the results and commented on the manuscript. The coordinates and the structure factor files have been deposited in the Protein Data Bank under accession code 2IQH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The coordinates and the structure factor files have been deposited in the Protein Data Bank under accession code 2IQH. Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Suplementary Information
The file contains Figures 1-5. Figure 1 shows experimental electron density maps of NP, Figure 2 shows a stereo diagram of NP, Figure 3 shows RNA binding activity of the influenza A/WSN/33 NP protein, Table 1 shows diffraction data statistics and Table 2 shows inter-subunit interactions of NP molecules in an NCS trimer.
Rights and permissions
About this article
Cite this article
Ye, Q., Krug, R. & Tao, Y. The mechanism by which influenza A virus nucleoprotein forms oligomers and binds RNA. Nature 444, 1078–1082 (2006). https://doi.org/10.1038/nature05379
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05379
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.