Abstract
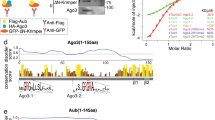
DNA methylation has important functions in stable, transcriptional gene silencing, immobilization of transposable elements and genome organization1. In Arabidopsis, DNA methylation can be induced by double-stranded RNA through the RNA interference (RNAi) pathway, a response known as RNA-directed DNA methylation2. This requires a specialized set of RNAi components, including ARGONAUTE4 (AGO4)3,4,5,6. Here we show that AGO4 binds to small RNAs including small interfering RNAs (siRNAs) originating from transposable and repetitive elements, and cleaves target RNA transcripts. Single mutations in the Asp-Asp-His catalytic motif of AGO4 do not affect siRNA-binding activity but abolish its catalytic potential. siRNA accumulation and non-CpG DNA methylation at some loci require the catalytic activity of AGO4, whereas others are less dependent on this activity. Our results are consistent with a model in which AGO4 can function at target loci through two distinct and separable mechanisms. First, AGO4 can recruit components that signal DNA methylation in a manner independent of its catalytic activity. Second, AGO4 catalytic activity can be crucial for the generation of secondary siRNAs that reinforce its repressive effects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chan, S. W., Henderson, I. R. & Jacobsen, S. E. Gardening the genome: DNA methylation in Arabidopsis thaliana. Nature Rev. Genet. 6, 351–360 (2005)
Matzke, M. A. & Birchler, J. A. RNAi-mediated pathways in the nucleus. Nature Rev. Genet. 6, 24–35 (2005)
Xie, Z. et al. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2, e104 (2004)
Zilberman, D., Cao, X. & Jacobsen, S. E. ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299, 716–719 (2003)
Chan, S. W. et al. RNA silencing genes control de novo DNA methylation. Science 303, 1336 (2004)
Zilberman, D. et al. Role of Arabidopsis ARGONAUTE4 in RNA-directed DNA methylation triggered by inverted repeats. Curr. Biol. 14, 1214–1220 (2004)
Herr, A. J., Jensen, M. B., Dalmay, T. & Baulcombe, D. C. RNA polymerase IV directs silencing of endogenous DNA. Science 308, 118–120 (2005)
Onodera, Y. et al. Plant nuclear RNA polymerase IV mediates siRNA and DNA methylation-dependent heterochromatin formation. Cell 120, 613–622 (2005)
Li, C. et al. An ARGONAUTE4-containing nuclear processing center colocalized with Cajal bodies in Arabidopsis thaliana. Cell 126, 93–106 (2006)
Lippman, Z., May, B., Yordan, C., Singer, T. & Martienssen, R. Distinct mechanisms determine transposon inheritance and methylation via small interfering RNA and histone modification. PLoS Biol. 1, e67 (2003)
Rohila, J. S., Chen, M., Cerny, R. & Fromm, M. E. Improved tandem affinity purification tag and methods for isolation of protein heterocomplexes from plants. Plant J. 38, 172–181 (2004)
Qi, Y., Denli, A. M. & Hannon, G. J. Biochemical specialization within Arabidopsis RNA silencing pathways. Mol. Cell 19, 421–428 (2005)
Baumberger, N. & Baulcombe, D. C. Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc. Natl Acad. Sci. USA 102, 11928–11933 (2005)
Hamilton, A., Voinnet, O., Chappell, L. & Baulcombe, D. Two classes of short interfering RNA in RNA silencing. EMBO J. 21, 4671–4679 (2002)
Singer, T., Yordan, C. & Martienssen, R. A. Robertson’s Mutator transposons in A. thaliana are regulated by the chromatin-remodeling gene Decrease in DNA Methylation (DDM1). Genes Dev. 15, 591–602 (2001)
Cao, X. & Jacobsen, S. E. Locus-specific control of asymmetric and CpNpG methylation by the DRM and CMT3 methyltransferase genes. Proc. Natl Acad. Sci. USA 99, (Suppl. 4)16491–16498 (2002)
Girard, A., Sachidanandam, R., Hannon, G. & Carmell, M. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006)
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005)
Vaucheret, H., Vazquez, F., Crete, P. & Bartel, D. P. The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes Dev. 18, 1187–1197 (2004)
Grewal, S. I. & Moazed, D. Heterochromatin and epigenetic control of gene expression. Science 301, 798–802 (2003)
Bao, N., Lye, K. W. & Barton, M. K. MicroRNA binding sites in Arabidopsis class III HD-ZIP mRNAs are required for methylation of the template chromosome. Dev. Cell 7, 653–662 (2004)
Qi, Y. & Hannon, G. J. Uncovering RNAi mechanisms in plants: biochemistry enters the foray. FEBS Lett. 579, 5899–5903 (2005)
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004)
Rivas, F. V. et al. Purified Argonaute2 and an siRNA form recombinant human RISC. Nature Struct. Mol. Biol. 12, 340–349 (2005)
Jacobsen, S. & Meyerowitz, E. Hypermethylated SUPERMAN epigenetic alleles in Arabidopsis. Science 277, 1100–1103 (1997)
Allen, E., Xie, Z., Gustafson, A. M. & Carrington, J. C. MicroRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121, 207–221 (2005)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Acknowledgements
We thank S. Ossowski and D. Weigel for help with the analysis of AGO1-associated RNAs; R. Sachidanandam for bioinformatics assistance; T. Mulligan for assistance with Arabidopsis culture; and members of the Hannon laboratory for discussion. This work was supported in part by grants from the NSF and the NIH (G.J.H.). G.J.H. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Sequences referred to in this study have been deposited in the GenBank database under accession numbers DQ927324–DQ972825. Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1–10. (PDF 1737 kb)
Supplementary Figure Legends
This file contains text to accompany the above Supplementary Figures (DOC 30 kb)
Supplementary Tables
This file contains Supplementary Tables 1–10. (PDF 181 kb)
Supplementary Methods
This file contains additional details of the methods used in this study. (PDF 88 kb)
Rights and permissions
About this article
Cite this article
Qi, Y., He, X., Wang, XJ. et al. Distinct catalytic and non-catalytic roles of ARGONAUTE4 in RNA-directed DNA methylation. Nature 443, 1008–1012 (2006). https://doi.org/10.1038/nature05198
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05198
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.