Abstract
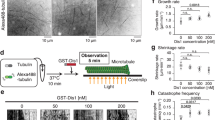
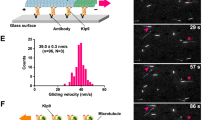
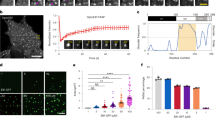
Chromosomes interact through their kinetochores with microtubule plus ends and they are segregated to the spindle poles as the kinetochore microtubules shorten during anaphase A of mitosis. The molecular natures and identities of coupling proteins that allow microtubule depolymerization to pull chromosomes to poles during anaphase have long remained elusive1. In budding yeast, the ten-protein Dam1 complex is a critical microtubule-binding component of the kinetochore2 that oligomerizes into a 50-nm ring around a microtubule in vitro3,4. Here we show, with the use of a real-time, two-colour fluorescence microscopy assay, that the ring complex moves processively for several micrometres at the ends of depolymerizing microtubules without detaching from the lattice. Electron microscopic analysis of ‘end-on views’ revealed a 16-fold symmetry of the kinetochore rings. This out-of-register arrangement with respect to the 13-fold microtubule symmetry is consistent with a sliding mechanism based on an electrostatically coupled ring–microtubule interface. The Dam1 ring complex is a molecular device that can translate the force generated by microtubule depolymerization into movement along the lattice to facilitate chromosome segregation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Maiato, H., Deluca, J., Salmon, E. D. & Earnshaw, W. C. The dynamic kinetochore–microtubule interface. J. Cell Sci. 117, 5461–5477 (2004)
Cheeseman, I. M. et al. Implication of a novel multiprotein Dam1p complex in outer kinetochore function. J. Cell Biol. 155, 1137–1145 (2001)
Westermann, S. et al. Formation of a dynamic kinetochore–microtubule interface through assembly of the Dam1 ring complex. Mol. Cell 17, 277–290 (2005)
Miranda, J. J., De Wulf, P., Sorger, P. K. & Harrison, S. C. The yeast DASH complex forms closed rings on microtubules. Nature Struct. Mol. Biol. 12, 138–143 (2005)
Inoue, S. & Salmon, E. D. Force generation by microtubule assembly/disassembly in mitosis and related movements. Mol. Biol. Cell 6, 1619–1640 (1995)
Koshland, D. E., Mitchison, T. J. & Kirschner, M. W. Polewards chromosome movement driven by microtubule depolymerization in vitro. Nature 331, 499–504 (1988)
Lombillo, V. A., Stewart, R. J. & McIntosh, J. R. Minus-end-directed motion of kinesin-coated microspheres driven by microtubule depolymerization. Nature 373, 161–164 (1995)
Coue, M., Lombillo, V. A. & McIntosh, J. R. Microtubule depolymerization promotes particle and chromosome movement in vitro. J. Cell Biol. 112, 1165–1175 (1991)
Hill, T. L. Theoretical problems related to the attachment of microtubules to kinetochores. Proc. Natl Acad. Sci. USA 82, 4404–4408 (1985)
Molodtsov, M. I., Grishchuk, E. L., Efremov, A. K., McIntosh, J. R. & Ataullakhanov, F. I. Force production by depolymerizing microtubules: a theoretical study. Proc. Natl Acad. Sci. USA 102, 4353–4358 (2005)
Straight, A. F., Marshall, W. F., Sedat, J. W. & Murray, A. W. Mitosis in living budding yeast: anaphase A but no metaphase plate. Science 277, 574–578 (1997)
Maddox, P. S., Bloom, K. S. & Salmon, E. D. The polarity and dynamics of microtubule assembly in the budding yeast Saccharomyces cerevisiae. Nature Cell Biol. 2, 36–41 (2000)
De Wulf, P., McAinsh, A. D. & Sorger, P. K. Hierarchical assembly of the budding yeast kinetochore from multiple subcomplexes. Genes Dev. 17, 2902–2921 (2003)
Westermann, S. et al. Architecture of the budding yeast kinetochore reveals a conserved molecular core. J. Cell Biol. 163, 215–222 (2003)
Cheeseman, I. M., Drubin, D. G. & Barnes, G. Simple centromere, complex kinetochore: linking spindle microtubules and centromeric DNA in budding yeast. J. Cell Biol. 157, 199–203 (2002)
Cheeseman, I. M., Enquist-Newman, M., Muller-Reichert, T., Drubin, D. G. & Barnes, G. Mitotic spindle integrity and kinetochore function linked by the Duo1p/Dam1p complex. J. Cell Biol. 152, 197–212 (2001)
Cheeseman, I. M. et al. Phospho-regulation of kinetochore-microtubule attachments by the Aurora kinase Ipl1p. Cell 111, 163–172 (2002)
Tanaka, K. et al. Molecular mechanisms of kinetochore capture by spindle microtubules. Nature 434, 987–994 (2005)
Kapitein, L. C. et al. The bipolar mitotic kinesin Eg5 moves on both microtubules that it crosslinks. Nature 435, 114–118 (2005)
Vale, R. D., Soll, D. R. & Gibbons, I. R. One-dimensional diffusion of microtubules bound to flagellar dynein. Cell 59, 915–925 (1989)
Howard, J. & Hyman, A. A. Dynamics and mechanics of the microtubule plus end. Nature 422, 753–758 (2003)
Wang, H. W. & Nogales, E. Nucleotide-dependent bending flexibility of tubulin regulates microtubule assembly. Nature 435, 911–915 (2005)
Hyman, A. A., Chretien, D., Arnal, I. & Wade, R. H. Structural changes accompanying GTP hydrolysis in microtubules: information from a slowly hydrolyzable analogue guanylyl-(α,β)-methylene-diphosphonate. J. Cell Biol. 128, 117–125 (1995)
Mandelkow, E. M., Mandelkow, E. & Milligan, R. A. Microtubule dynamics and microtubule caps: a time-resolved cryo-electron microscopy study. J. Cell Biol. 114, 977–991 (1991)
Mahadevan, L. & Mitchison, T. J. Cell biology: powerful curves. Nature 435, 895–897 (2005)
Grishchuk, E. L., Molodtsov, M. I, Ataullakhanov, F. I. & McIntosh, J. R. Force production by disassembling microtubules. Nature 438, 384–388 (2005)
Pinsky, B. A. & Biggins, S. The spindle checkpoint: tension versus attachment. Trends Cell Biol. 15, 486–493 (2005)
Liu, X., McLeod, I., Anderson, S., Yates, J. R. & He, X. Molecular analysis of kinetochore architecture in fission yeast. EMBO J. 24, 2919–2930 (2005)
Sanchez-Perez, I. et al. The DASH complex and Klp5/Klp6 kinesin coordinate bipolar chromosome attachment in fission yeast. EMBO J. 24, 2931–2943 (2005)
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996)
Acknowledgements
We thank C. Walzcak for the gift of recombinant XMCAK1; M. Kaksonen and C. Toret for help with image analysis; and all members of the Barnes/Drubin laboratory for discussions. This work was supported by grants from the National Institute of General Medical Sciences to G.B. and E.N., and from the Office of Basic Energy Science of the US Department of Energy. E.N. is a Howard Hughes Medical Institute Investigator. Research in this study was in part supported by a grant to D.G.D. from Phillip Morris USA Inc. and Phillip Morris International and by a postdoctoral fellowship of the Deutsche Forschungsgemeinschaft to S.W.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Movie 1
Processive movement of the Dam1 ring complex on depolymerizing microtubule ends. (MOV 1666 kb)
Supplementary Movie 2
Microtubule is free in solution and depolymerizes from both ends. (MOV 830 kb)
Supplementary Movie 3
Force generation at the depolymerizing end. (MOV 714 kb)
Supplementary Movie 4
Dam1-coupled microbeads translocate with disassembling microtubule ends. (MOV 605 kb)
Supplementary Movie 5
Dam1 rings are “collected” at the depolymerizing end. (MOV 212 kb)
Supplementary Movie 6
Stabilization of microtubule ends as a function of the Dam1 ring concentration. (MOV 1288 kb)
Supplementary Movie 7
Alexa488 Dam1 complex shows undirected one-dimensional diffusion on taxol-stabilized microtubules. (MOV 10385 kb)
Supplementary Movie 8
Individual Dam1 signal on a single taxol stabilized microtubule shows back and forth movement. Time between frames is 1 second. (MOV 54 kb)
Supplementary Figure 9
Movements of single Dam1 signals on a microtubule versus Dam1 signal stuck to the coverslip surface. (MOV 424 kb)
Supplementary Figure 1
Sketch of the bead experiment and fluorescence micrograph of the bead experiment. (PDF 64 kb)
Rights and permissions
About this article
Cite this article
Westermann, S., Wang, HW., Avila-Sakar, A. et al. The Dam1 kinetochore ring complex moves processively on depolymerizing microtubule ends. Nature 440, 565–569 (2006). https://doi.org/10.1038/nature04409
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04409
This article is cited by
-
Measuring and modeling forces generated by microtubules
Biophysical Reviews (2023)
-
Regulation of microtubule dynamics, mechanics and function through the growing tip
Nature Reviews Molecular Cell Biology (2021)
-
Tubulin islands containing slowly hydrolyzable GTP analogs regulate the mechanism and kinetics of microtubule depolymerization
Scientific Reports (2020)
-
Killing two birds with one stone: how budding yeast Mps1 controls chromosome segregation and spindle assembly checkpoint through phosphorylation of a single kinetochore protein
Current Genetics (2020)
-
The phosphorylation of a kinetochore protein Dam1 by Aurora B/Ipl1 kinase promotes chromosome bipolar attachment in yeast
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.